Gene
KWMTBOMO03649
Pre Gene Modal
BGIBMGA014550
Annotation
gag-like_protein_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 2.569
Sequence
CDS
ATGGCTTTCTTCTGCAGCAGTGGCCAGAATTCAATATACAACTGGGTGAGGGATCACAGACTCCATCACAAGTTCGCCGACACCGATGCCGACCCTCACAACGTGAACCGAGGCTTCTTCTTCTCCCACATAGGATGGCTGATGATGAAGAAAAACGAAAGCGTCCTTACGAAAGGAAAGCTGATTGACATGAGTGACATAGAGAGCGATTCCCATTTGATGTTTCAACATAAGTATGATGTTACTTTTACCTTCGACCTTAGATTTAAGTTGTATCTTGGATGTCCGAGAAGTTGGGGCCTCGCCGCAAGGCGAGTGTTTTCAAGACCCGGGGCCGCCCCACCTGGGCACTCAGCGATCATGGACGCTGTATTCGCGGAATTCCTCCGACTTCGCCACCCACAGCTCGCCTCAGAGTTTCTGGCCTTCAAGGCCGATCACACCGCGAGCCCTCTCGTGGACTCCGCCGCGCTCGCTGCTCCTGCGTCGCCTGTACCTGCGTGCAAAGCTTCTGCATCGAGCACCGCAGCCTCCGTCGTGCCTGCCGTTCCCGCGTCGCCTATACTGGCGAGTAAAACTGCTGCGTCGTCCGCCGTGGCCACCGTTCCAGCAGCGCGATCATCCGCAGCCTCCGTCGTGCCCTCCAAAACACCTGCTCGTAGGTCACCTGCGCCCGCCTCCTCGTCCTCCGACTCTGACTCGGAGATGGAGGTCGACCTCGCTCCCGCCTCATCGACGGATGGATTCACCCTGGTACAAAAGGACCCCTCCCCGGTTATCCTTCAAGAGAAGGCAGCTTGGGATCGAGTTTCTCTGGCCCTTAAGGCCAAAAATATCAACTTCACGAATGCCCGTAACCTCGCGAACGGCATTCAAATTAAGGTTCGAACACCCGACGACCATAGGGCCCTCTCTTCTTACCTCCGTAAGGAACGTATAAGTTTCCACACGTATACGCTCCAGGAGGAGCGCGAACTCCGTGCCGTCATACGTGGCATCCCTAAAGAGTTGAATGCCGAGCTAGTCAAGGCCGACCTTCTCGAACAAGGCCTACCGGTTAACTCCGTACACCGCATGCACACAGGTCGCGGAAGGGAGCCATATAACATGGTTCTCGTCGCCCTCCAGCCTACCCCCGAGGGTAAGCAAATATTCAACATCCGAACGGTCTGTAGCCTTTCCGGAATCGCCGTCGAAACCCCACACAAGAAAGGCACACCTAGCCAGTGCCATAACTGCCAATTATACGGGCATTCTTCCCGTAACTGTCACGCGCGCCCCCGATGCGTCAAGTGTTTAGGAGATCACGCCACGGCCCTCTGCACTCGCGATCAAAAAACCGCGACGGAACCGCCTAGCTGCGTCCTGTGTCGTACACAGGGTCACCCCGCAAATTACCGCGGATGCCCCCGAGCCCCGAAAATAAATCGCCGCGTCGCCCGCCAAAACCGCCTCCGAGCTTCCGGCCCCGACATCAAAGCCTCGGCACCCTCTGTGTCGCAGGCTAAGCCAGCGTTCGATCCGGCGCCGGTGCCCAGTGTCTCGGCTTGGGCGAAACCGCTGCCGTACACGAACACGGTTACAACTCCCTCCTCCGCGATTCGTCCCGCCCCCGCAACTCGTCCCTCTCCCGCGACTTGCCCTCCGACCGCGTCCGACAATCTCGCTTTAGCGATCGACTTCTTTCAGTCGATCAACTTTGAGCGCGTTAACGCTTTGGGCGACGCCATTCGCGCTGCCTCCACTGCACAACACTTTATCGCCGTTGTGCAAGAATACGCCGACGTATACGCGTCATTAAATACGTACGTCCTCCCCTCACTCCGCCGGTAA
Protein
MAFFCSSGQNSIYNWVRDHRLHHKFADTDADPHNVNRGFFFSHIGWLMMKKNESVLTKGKLIDMSDIESDSHLMFQHKYDVTFTFDLRFKLYLGCPRSWGLAARRVFSRPGAAPPGHSAIMDAVFAEFLRLRHPQLASEFLAFKADHTASPLVDSAALAAPASPVPACKASASSTAASVVPAVPASPILASKTAASSAVATVPAARSSAASVVPSKTPARRSPAPASSSSDSDSEMEVDLAPASSTDGFTLVQKDPSPVILQEKAAWDRVSLALKAKNINFTNARNLANGIQIKVRTPDDHRALSSYLRKERISFHTYTLQEERELRAVIRGIPKELNAELVKADLLEQGLPVNSVHRMHTGRGREPYNMVLVALQPTPEGKQIFNIRTVCSLSGIAVETPHKKGTPSQCHNCQLYGHSSRNCHARPRCVKCLGDHATALCTRDQKTATEPPSCVLCRTQGHPANYRGCPRAPKINRRVARQNRLRASGPDIKASAPSVSQAKPAFDPAPVPSVSAWAKPLPYTNTVTTPSSAIRPAPATRPSPATCPPTASDNLALAIDFFQSINFERVNALGDAIRAASTAQHFIAVVQEYADVYASLNTYVLPSLRR
Summary
Uniprot
EMBL
PRIDE
Pfam
PF07530 PRE_C2HC
Interpro
IPR006579
Pre_C2HC_dom
ProteinModelPortal
PDB
4ZYO
E-value=1.41275e-22,
Score=265
Ontologies
PATHWAY
GO
Topology
Length:
610
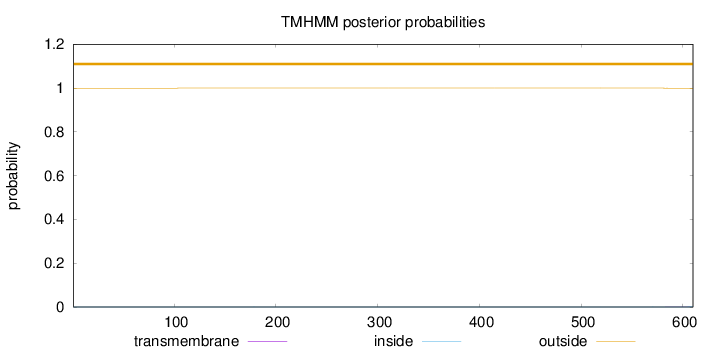
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.01591
Exp number, first 60 AAs:
0.00066
Total prob of N-in:
0.00043
outside
1 - 610
Population Genetic Test Statistics
Pi
1.030513
Theta
1.198003
Tajima's D
-0.899541
CLR
149.059725
CSRT
0.162241887905605
Interpretation
Uncertain
