Gene
KWMTBOMO02987
Pre Gene Modal
BGIBMGA003622
Annotation
PREDICTED:_LOW_QUALITY_PROTEIN:_dynein_beta_chain?_ciliary-like_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 3.417
Sequence
CDS
ATGGGAGGTGGTCCTGCTGGGCCTGCTGGCACGGGAAAAACAGAGACTACCAAAGATCTGGGAAGGGCTTTGGGAATCATGGTGTACGTCTTCAATTGTTCTGAACAAATGGACTATCAGTCTTGCGGGAATATCTACAAAGGTCTGGCGCAATCTGGTGCTTGGGGTTGTTTTGATGAATTCAACAGAATTTCCGTAGAAGTATTATCCGTGGTGGCGGTTCAAGTTAAATCTGTACAAGACGCTATAAGAGACAAAAAGGAAAAATTTAATTTCATGGGCGAAATTATTAGTTTGGTGCCAACTGTCGGGATATTTATAACGATGAACCCCGGTTATGCCGGTAGAACAGAGCTTCCGGAGAACCTCAAAGCGCTATTCAGACCATGCGCCATGGTCGTACCAGATTTCGAATTGATTTGTGAGATCATGTTAGTCGCCGAAGGCTTCCAAGAGGCAAAAATTTTGGCCAGAAAATTCATAACACTTTATACTTTGTGCAAAGAACTTCTTTCGAAACAAGATCACTATGATTGGGGTTTAAGGGCTATAAAGTCTGTGTTGGTAGTAGCCGGGTCACTCAAGCGAGGGGATCCGGGCAGACCGGAGGAAGAAGTATTGATGAGGACACTTCGCGATTTCAATATACCGAAAATAATCACTGACGACATGCCAGTTTTTATGGGACTGATTGGAGATCTATTTCCCGCCCTTGAAGTGCCTCGTAAACGTGATTTAGAATTCGAAAGAACTGTCAAGCAAGCAGCCATGGATCTCACGTTACAGCCCGAAGATAATTTCATACTGAAGGTTGTACAGTTAGAAGAATTACTAGAGGTCCGCCATTCGGTATTCATAGTTGGAAATGCTGGTACTGGTAAAACGCAAGTCTGGAAAACTTTGTTTAAAACTTATCAAAACTTGAAAAAGAAACCATTGTTTAATGACTTGAATCCGAAAGCAGTGACAAACGATGAGTTGTTCGGAATTATAAATCCGGCGACCAGAGAGTGGAAAGATGGCTTGTTTTCAGTTATAATGAGAGATCAAGCTAACATTGTCGGCGAGAATCCGAAGTGGATTATTTTAGACGGAGATATAGATCCTATGTGGATTGAATCTTTAAATACTGTTATGGATGACAATAAAATTCTCACTCTAGCTAGCAACGAAAGGATTGCTCTAACGCCTACCATGAGATTGATGTTTGAAATATCAAATCTTCGTACAGCTACCCCGGCAACAGTGTCGAGAGCTGGCATCCTGTATATTAATCCACAAGATTTAGGATGGAATCCGTAA
Protein
MGGGPAGPAGTGKTETTKDLGRALGIMVYVFNCSEQMDYQSCGNIYKGLAQSGAWGCFDEFNRISVEVLSVVAVQVKSVQDAIRDKKEKFNFMGEIISLVPTVGIFITMNPGYAGRTELPENLKALFRPCAMVVPDFELICEIMLVAEGFQEAKILARKFITLYTLCKELLSKQDHYDWGLRAIKSVLVVAGSLKRGDPGRPEEEVLMRTLRDFNIPKIITDDMPVFMGLIGDLFPALEVPRKRDLEFERTVKQAAMDLTLQPEDNFILKVVQLEELLEVRHSVFIVGNAGTGKTQVWKTLFKTYQNLKKKPLFNDLNPKAVTNDELFGIINPATREWKDGLFSVIMRDQANIVGENPKWIILDGDIDPMWIESLNTVMDDNKILTLASNERIALTPTMRLMFEISNLRTATPATVSRAGILYINPQDLGWNP
Summary
Uniprot
H9J285
A0A212FLC8
A0A0L7LUC4
A0A2A4JJP5
A0A2H1X2P9
A0A182IVR3
+ More
A0A182S5M4 A0A182MNY4 A0A182RBC6 A0A182NDL4 A0A182WR89 A0A182VFZ9 A0A182YIP1 A0A182JRE6 A0A182W3T1 A0A182TS04 A0A182HX97 Q5TUD8 A0A1S4H4T4 A0A084VZI7 A0A182PVS5 A0A182QB97 B0WUU8 A0A0P8Y0J0 A0A2M4A0B6 A0A182FM39 W5J456 B3M2X7 B4KE24 B4GM15 B4M4V0 B4JSY1 Q293J4 A0A0R3NKH1 A0A1W4VZS9 A0A1W4VLP2 Q9VDG0 A0A0R1E5P0 B3P2Z8 A0A0B4KHJ4 A0A0R1E127 B4PQ99 A0A0R1E125 A0A0R1E1G1 B4IKM6 A0A0M4ELU1 A0A0Q9X5F3 E0VLA6 A0A182GMZ9 B4R1N9 B4N969 A0A1S4FKM6 A0A1A9XQ49 A0A3B0JSI1 A0A232F8J8 Q16XH8 A0A1B0BD95 A0A1A9WGW2 A0A1B0G9S0 A0A1I8M6W0 A0A1I8M6W5 A0A1A9ZDG9 A0A3L8DCZ9 A0A1A9UNH8 A0A1I8PWQ2 A0A0L0BTW6 A0A0L7QYM1 A0A0M9AA00 A0A310S6L6 E9IZF0 A0A154PPG4 E2AYZ0 D6W7L1 A0A139WPL9 E2BE63 A0A0J7P079 A0A151JXC9 A0A195AUI3 A0A158P169 F4WD53 A0A151WVZ6 A0A195CXR3 A0A2A3EQV0 A0A1J1IHV8 A0A088AF70 A0A151JQJ9 Q71JD2 J9K4Q1 A0A1W4X3T6 A0A182H543 Q17G87 U4UP43 N6U0M0 A0A026WFI7 B0W485 A0A2M4CMP8 A0A182NRZ1 A0A084WBP9 A0A182QZC2 A0A182JYB8
A0A182S5M4 A0A182MNY4 A0A182RBC6 A0A182NDL4 A0A182WR89 A0A182VFZ9 A0A182YIP1 A0A182JRE6 A0A182W3T1 A0A182TS04 A0A182HX97 Q5TUD8 A0A1S4H4T4 A0A084VZI7 A0A182PVS5 A0A182QB97 B0WUU8 A0A0P8Y0J0 A0A2M4A0B6 A0A182FM39 W5J456 B3M2X7 B4KE24 B4GM15 B4M4V0 B4JSY1 Q293J4 A0A0R3NKH1 A0A1W4VZS9 A0A1W4VLP2 Q9VDG0 A0A0R1E5P0 B3P2Z8 A0A0B4KHJ4 A0A0R1E127 B4PQ99 A0A0R1E125 A0A0R1E1G1 B4IKM6 A0A0M4ELU1 A0A0Q9X5F3 E0VLA6 A0A182GMZ9 B4R1N9 B4N969 A0A1S4FKM6 A0A1A9XQ49 A0A3B0JSI1 A0A232F8J8 Q16XH8 A0A1B0BD95 A0A1A9WGW2 A0A1B0G9S0 A0A1I8M6W0 A0A1I8M6W5 A0A1A9ZDG9 A0A3L8DCZ9 A0A1A9UNH8 A0A1I8PWQ2 A0A0L0BTW6 A0A0L7QYM1 A0A0M9AA00 A0A310S6L6 E9IZF0 A0A154PPG4 E2AYZ0 D6W7L1 A0A139WPL9 E2BE63 A0A0J7P079 A0A151JXC9 A0A195AUI3 A0A158P169 F4WD53 A0A151WVZ6 A0A195CXR3 A0A2A3EQV0 A0A1J1IHV8 A0A088AF70 A0A151JQJ9 Q71JD2 J9K4Q1 A0A1W4X3T6 A0A182H543 Q17G87 U4UP43 N6U0M0 A0A026WFI7 B0W485 A0A2M4CMP8 A0A182NRZ1 A0A084WBP9 A0A182QZC2 A0A182JYB8
Pubmed
19121390
22118469
26227816
25244985
12364791
24438588
+ More
17994087 20920257 23761445 15632085 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 17550304 20566863 26483478 17510324 28648823 25315136 30249741 26108605 21282665 20798317 18362917 19820115 21347285 21719571 23537049 24508170
17994087 20920257 23761445 15632085 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 17550304 20566863 26483478 17510324 28648823 25315136 30249741 26108605 21282665 20798317 18362917 19820115 21347285 21719571 23537049 24508170
EMBL
BABH01007454
BABH01007455
BABH01007456
BABH01007457
AGBW02007827
OWR54500.1
+ More
JTDY01000081 KOB78984.1 NWSH01001294 PCG71814.1 ODYU01012953 SOQ59508.1 AXCM01000636 APCN01005160 AAAB01008849 EAL41019.3 ATLV01018849 KE525251 KFB43381.1 AXCN02001753 DS232111 EDS35091.1 CH902617 KPU80296.1 GGFK01000870 MBW34191.1 ADMH02002126 ETN58641.1 EDV43507.1 CH933806 EDW16045.2 CH479185 EDW38589.1 CH940652 EDW59661.2 CH916373 EDV94871.1 CM000070 EAL29221.2 KRT01415.1 AE014297 AAF55834.3 CM000160 KRK03002.1 CH954181 EDV48312.1 AGB96173.1 KRK03003.1 EDW96208.2 KRK03004.1 KRK03001.1 CH480853 EDW51630.1 CP012526 ALC46327.1 CH964232 KRF99495.1 AAZO01003313 DS235271 EEB14162.1 JXUM01075435 KQ562886 KXJ74924.1 CM000364 EDX12148.1 EDW81616.1 OUUW01000007 SPP83382.1 NNAY01000654 OXU27164.1 CH477538 EAT39332.1 JXJN01012383 CCAG010011423 QOIP01000009 RLU18201.1 JRES01001345 KNC23525.1 KQ414688 KOC63714.1 KQ435706 KOX80265.1 KQ767877 OAD53287.1 GL767121 EFZ14108.1 KQ434980 KZC13060.1 GL444028 EFN61360.1 KQ971307 EFA11309.2 KYB29781.1 GL447755 EFN86050.1 LBMM01000571 KMQ98045.1 KQ981572 KYN39778.1 KQ976738 KYM75717.1 ADTU01005848 ADTU01005849 GL888084 EGI67861.1 KQ982691 KYQ52109.1 KQ977141 KYN05431.1 KZ288199 PBC33642.1 CVRI01000047 CRK98041.1 KQ978644 KYN29472.1 AF494076 AAQ06635.1 ABLF02030722 ABLF02030734 ABLF02030735 ABLF02030737 ABLF02030738 JXUM01026200 JXUM01026201 JXUM01026202 JXUM01026203 KQ560749 KXJ81025.1 CH477265 EAT45561.1 KB632384 ERL94273.1 APGK01047112 APGK01047113 APGK01047114 APGK01047115 APGK01047116 APGK01047117 APGK01047118 KB741077 ENN74141.1 KK107235 EZA54880.1 DS231835 EDS32929.1 GGFL01002293 MBW66471.1 ATLV01022415 KE525332 KFB47643.1 AXCN02000255
JTDY01000081 KOB78984.1 NWSH01001294 PCG71814.1 ODYU01012953 SOQ59508.1 AXCM01000636 APCN01005160 AAAB01008849 EAL41019.3 ATLV01018849 KE525251 KFB43381.1 AXCN02001753 DS232111 EDS35091.1 CH902617 KPU80296.1 GGFK01000870 MBW34191.1 ADMH02002126 ETN58641.1 EDV43507.1 CH933806 EDW16045.2 CH479185 EDW38589.1 CH940652 EDW59661.2 CH916373 EDV94871.1 CM000070 EAL29221.2 KRT01415.1 AE014297 AAF55834.3 CM000160 KRK03002.1 CH954181 EDV48312.1 AGB96173.1 KRK03003.1 EDW96208.2 KRK03004.1 KRK03001.1 CH480853 EDW51630.1 CP012526 ALC46327.1 CH964232 KRF99495.1 AAZO01003313 DS235271 EEB14162.1 JXUM01075435 KQ562886 KXJ74924.1 CM000364 EDX12148.1 EDW81616.1 OUUW01000007 SPP83382.1 NNAY01000654 OXU27164.1 CH477538 EAT39332.1 JXJN01012383 CCAG010011423 QOIP01000009 RLU18201.1 JRES01001345 KNC23525.1 KQ414688 KOC63714.1 KQ435706 KOX80265.1 KQ767877 OAD53287.1 GL767121 EFZ14108.1 KQ434980 KZC13060.1 GL444028 EFN61360.1 KQ971307 EFA11309.2 KYB29781.1 GL447755 EFN86050.1 LBMM01000571 KMQ98045.1 KQ981572 KYN39778.1 KQ976738 KYM75717.1 ADTU01005848 ADTU01005849 GL888084 EGI67861.1 KQ982691 KYQ52109.1 KQ977141 KYN05431.1 KZ288199 PBC33642.1 CVRI01000047 CRK98041.1 KQ978644 KYN29472.1 AF494076 AAQ06635.1 ABLF02030722 ABLF02030734 ABLF02030735 ABLF02030737 ABLF02030738 JXUM01026200 JXUM01026201 JXUM01026202 JXUM01026203 KQ560749 KXJ81025.1 CH477265 EAT45561.1 KB632384 ERL94273.1 APGK01047112 APGK01047113 APGK01047114 APGK01047115 APGK01047116 APGK01047117 APGK01047118 KB741077 ENN74141.1 KK107235 EZA54880.1 DS231835 EDS32929.1 GGFL01002293 MBW66471.1 ATLV01022415 KE525332 KFB47643.1 AXCN02000255
Proteomes
UP000005204
UP000007151
UP000037510
UP000218220
UP000075880
UP000075901
+ More
UP000075883 UP000075900 UP000075884 UP000076407 UP000075903 UP000076408 UP000075881 UP000075920 UP000075902 UP000075840 UP000007062 UP000030765 UP000075885 UP000075886 UP000002320 UP000007801 UP000069272 UP000000673 UP000009192 UP000008744 UP000008792 UP000001070 UP000001819 UP000192221 UP000000803 UP000002282 UP000008711 UP000001292 UP000092553 UP000007798 UP000009046 UP000069940 UP000249989 UP000000304 UP000092443 UP000268350 UP000215335 UP000008820 UP000092460 UP000091820 UP000092444 UP000095301 UP000092445 UP000279307 UP000078200 UP000095300 UP000037069 UP000053825 UP000053105 UP000076502 UP000000311 UP000007266 UP000008237 UP000036403 UP000078541 UP000078540 UP000005205 UP000007755 UP000075809 UP000078542 UP000242457 UP000183832 UP000005203 UP000078492 UP000007819 UP000192223 UP000030742 UP000019118 UP000053097
UP000075883 UP000075900 UP000075884 UP000076407 UP000075903 UP000076408 UP000075881 UP000075920 UP000075902 UP000075840 UP000007062 UP000030765 UP000075885 UP000075886 UP000002320 UP000007801 UP000069272 UP000000673 UP000009192 UP000008744 UP000008792 UP000001070 UP000001819 UP000192221 UP000000803 UP000002282 UP000008711 UP000001292 UP000092553 UP000007798 UP000009046 UP000069940 UP000249989 UP000000304 UP000092443 UP000268350 UP000215335 UP000008820 UP000092460 UP000091820 UP000092444 UP000095301 UP000092445 UP000279307 UP000078200 UP000095300 UP000037069 UP000053825 UP000053105 UP000076502 UP000000311 UP000007266 UP000008237 UP000036403 UP000078541 UP000078540 UP000005205 UP000007755 UP000075809 UP000078542 UP000242457 UP000183832 UP000005203 UP000078492 UP000007819 UP000192223 UP000030742 UP000019118 UP000053097
Pfam
Interpro
IPR042228
Dynein_2_C
+ More
IPR041228 Dynein_C
IPR041658 AAA_lid_11
IPR035699 AAA_6
IPR013594 Dynein_heavy_dom-1
IPR026983 DHC_fam
IPR042219 AAA_lid_11_sf
IPR013602 Dynein_heavy_dom-2
IPR004273 Dynein_heavy_D6_P-loop
IPR003593 AAA+_ATPase
IPR041466 Dynein_AAA5_ext
IPR027417 P-loop_NTPase
IPR024743 Dynein_HC_stalk
IPR024317 Dynein_heavy_chain_D4_dom
IPR041589 DNAH3_AAA_lid_1
IPR035706 AAA_9
IPR042222 Dynein_2_N
IPR009069 Cys_alpha_HP_mot_SF
IPR036084 Ser_inhib-like_sf
IPR011704 ATPase_dyneun-rel_AAA
IPR041228 Dynein_C
IPR041658 AAA_lid_11
IPR035699 AAA_6
IPR013594 Dynein_heavy_dom-1
IPR026983 DHC_fam
IPR042219 AAA_lid_11_sf
IPR013602 Dynein_heavy_dom-2
IPR004273 Dynein_heavy_D6_P-loop
IPR003593 AAA+_ATPase
IPR041466 Dynein_AAA5_ext
IPR027417 P-loop_NTPase
IPR024743 Dynein_HC_stalk
IPR024317 Dynein_heavy_chain_D4_dom
IPR041589 DNAH3_AAA_lid_1
IPR035706 AAA_9
IPR042222 Dynein_2_N
IPR009069 Cys_alpha_HP_mot_SF
IPR036084 Ser_inhib-like_sf
IPR011704 ATPase_dyneun-rel_AAA
Gene 3D
ProteinModelPortal
H9J285
A0A212FLC8
A0A0L7LUC4
A0A2A4JJP5
A0A2H1X2P9
A0A182IVR3
+ More
A0A182S5M4 A0A182MNY4 A0A182RBC6 A0A182NDL4 A0A182WR89 A0A182VFZ9 A0A182YIP1 A0A182JRE6 A0A182W3T1 A0A182TS04 A0A182HX97 Q5TUD8 A0A1S4H4T4 A0A084VZI7 A0A182PVS5 A0A182QB97 B0WUU8 A0A0P8Y0J0 A0A2M4A0B6 A0A182FM39 W5J456 B3M2X7 B4KE24 B4GM15 B4M4V0 B4JSY1 Q293J4 A0A0R3NKH1 A0A1W4VZS9 A0A1W4VLP2 Q9VDG0 A0A0R1E5P0 B3P2Z8 A0A0B4KHJ4 A0A0R1E127 B4PQ99 A0A0R1E125 A0A0R1E1G1 B4IKM6 A0A0M4ELU1 A0A0Q9X5F3 E0VLA6 A0A182GMZ9 B4R1N9 B4N969 A0A1S4FKM6 A0A1A9XQ49 A0A3B0JSI1 A0A232F8J8 Q16XH8 A0A1B0BD95 A0A1A9WGW2 A0A1B0G9S0 A0A1I8M6W0 A0A1I8M6W5 A0A1A9ZDG9 A0A3L8DCZ9 A0A1A9UNH8 A0A1I8PWQ2 A0A0L0BTW6 A0A0L7QYM1 A0A0M9AA00 A0A310S6L6 E9IZF0 A0A154PPG4 E2AYZ0 D6W7L1 A0A139WPL9 E2BE63 A0A0J7P079 A0A151JXC9 A0A195AUI3 A0A158P169 F4WD53 A0A151WVZ6 A0A195CXR3 A0A2A3EQV0 A0A1J1IHV8 A0A088AF70 A0A151JQJ9 Q71JD2 J9K4Q1 A0A1W4X3T6 A0A182H543 Q17G87 U4UP43 N6U0M0 A0A026WFI7 B0W485 A0A2M4CMP8 A0A182NRZ1 A0A084WBP9 A0A182QZC2 A0A182JYB8
A0A182S5M4 A0A182MNY4 A0A182RBC6 A0A182NDL4 A0A182WR89 A0A182VFZ9 A0A182YIP1 A0A182JRE6 A0A182W3T1 A0A182TS04 A0A182HX97 Q5TUD8 A0A1S4H4T4 A0A084VZI7 A0A182PVS5 A0A182QB97 B0WUU8 A0A0P8Y0J0 A0A2M4A0B6 A0A182FM39 W5J456 B3M2X7 B4KE24 B4GM15 B4M4V0 B4JSY1 Q293J4 A0A0R3NKH1 A0A1W4VZS9 A0A1W4VLP2 Q9VDG0 A0A0R1E5P0 B3P2Z8 A0A0B4KHJ4 A0A0R1E127 B4PQ99 A0A0R1E125 A0A0R1E1G1 B4IKM6 A0A0M4ELU1 A0A0Q9X5F3 E0VLA6 A0A182GMZ9 B4R1N9 B4N969 A0A1S4FKM6 A0A1A9XQ49 A0A3B0JSI1 A0A232F8J8 Q16XH8 A0A1B0BD95 A0A1A9WGW2 A0A1B0G9S0 A0A1I8M6W0 A0A1I8M6W5 A0A1A9ZDG9 A0A3L8DCZ9 A0A1A9UNH8 A0A1I8PWQ2 A0A0L0BTW6 A0A0L7QYM1 A0A0M9AA00 A0A310S6L6 E9IZF0 A0A154PPG4 E2AYZ0 D6W7L1 A0A139WPL9 E2BE63 A0A0J7P079 A0A151JXC9 A0A195AUI3 A0A158P169 F4WD53 A0A151WVZ6 A0A195CXR3 A0A2A3EQV0 A0A1J1IHV8 A0A088AF70 A0A151JQJ9 Q71JD2 J9K4Q1 A0A1W4X3T6 A0A182H543 Q17G87 U4UP43 N6U0M0 A0A026WFI7 B0W485 A0A2M4CMP8 A0A182NRZ1 A0A084WBP9 A0A182QZC2 A0A182JYB8
PDB
5NUG
E-value=7.83735e-93,
Score=869
Ontologies
GO
PANTHER
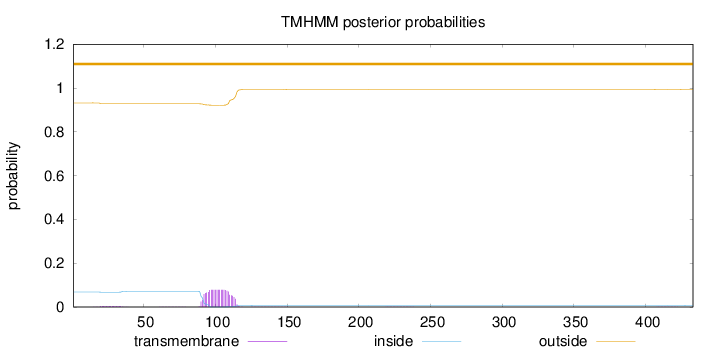
Topology
Length:
433
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
1.75714
Exp number, first 60 AAs:
0.06792
Total prob of N-in:
0.06794
outside
1 - 433
Population Genetic Test Statistics
Pi
28.863878
Theta
22.597351
Tajima's D
0.810795
CLR
0.2959
CSRT
0.619569021548923
Interpretation
Uncertain
