Gene
KWMTBOMO16799
Annotation
phosphatidylinositol-4-phosphate_5-kinase_type-1_alpha_[Lasius_niger]
Location in the cell
Cytoplasmic Reliability : 1.844 Nuclear Reliability : 1.645
Sequence
CDS
ATGGAGAGCATACAGGCGGAAAGCGAGCCCATAGATGAGGAGGAAGATGTCCCGTCCGGCGGCATCCCGGCCCGGAACGCGCGTGGCGAGCGTCTGCTGCTGTTCCTGGGCATCATCGACATACTCCAGTCGTATCGTCTCCGGAAGAAGCTCGAGCACACTTGGAAGAGCATGATACACGATGGCGACACTGTATCGGTCCACAGACCCAGCTTCTACGCTCAGAGATTCTTAGATTTCATGGCTAAAACCGTTTTCAAGAAGATACCTTCGTATCCAGAGACATGA
Protein
MESIQAESEPIDEEEDVPSGGIPARNARGERLLLFLGIIDILQSYRLRKKLEHTWKSMIHDGDTVSVHRPSFYAQRFLDFMAKTVFKKIPSYPET
Summary
Uniprot
A0A222AJ73
A0A0J7KTF7
A0A2A4JJ66
A0A3S2LXW5
A0A2H1VA20
A0A084VMW5
+ More
A0A151X330 E2A2X3 A0A2P8Z7B0 A0A151JRA7 A0A195ERZ8 E2BHP5 A0A151I2H8 A0A195D3Y7 A0A158NSF4 F4W5I7 A0A026VWW3 A0A088AUC9 A0A2A3EJ11 A0A310SBV2 A0A0C9RPZ2 A0A0C9RBR0 A0A1B6HKL2 A0A182SH93 A0A0L7RBR5 A0A154PG84 A0A2J7Q593 A0A2J7Q595 A0A2J7Q590 A0A2J7Q587 A0A2J7Q592 A0A2J7Q599 A0A0T6B6K4 A0A3L8DB04 A0A067QM30 Q16ED7 A0A182MFP5 A0A023ET03 A0A1S4G417 A0A1B6HGF5 A0A2M3Z6Y0 A0A1Q3EUA4 A0A1Y1MVB0 A0A182PFR2 A0A2M4AIV6 A0A1B6ITM7 A0A131YJ98 A0A1B6JKY6 A0A1W7R6X7 A0A1B6IAT8 A0A2M4AJ23 A0A1Q3FDR1 Q16TH1 A0A182KC06 A0A182Y9R8 U5EVS4 B0WJ81 A0A2M4BF98 A0A2M4CK77 A0A182KUG7 A0A2M3Z358 T1D6B5 T1E1Y7 A0A2M4CIM2 A0A2M4AEI7 A0A2M4CKB8 A0A139WFZ9 A0A2M4AES6 A0A182WH63 A0A139WFM3 W5JFQ6 A0A139WFP1 A0A1B6E678 A0A1B6CKI5 A0A1B6CWM0 A0A1B6C7D3 A0A1B6LKU7 A0A1B6KNH3 A0A1B6EY88 A0A1B6KUV2 K7IVE5 A0A1B6KF32 A0A182NF42 E0VBM0 A0A182R6J4 A0A182J2D4 A0A1J1IIV5 A0A453Z004 A0A1J1IM69 A0A182I559 A0A194QZN8 A0A224XQJ1 N6UKR1 T1J4I1 T1HHL0 A0A423SC62 A0A023FMB7 A0A293MA48 A0A131Y2B5 A0A0P4VL11
A0A151X330 E2A2X3 A0A2P8Z7B0 A0A151JRA7 A0A195ERZ8 E2BHP5 A0A151I2H8 A0A195D3Y7 A0A158NSF4 F4W5I7 A0A026VWW3 A0A088AUC9 A0A2A3EJ11 A0A310SBV2 A0A0C9RPZ2 A0A0C9RBR0 A0A1B6HKL2 A0A182SH93 A0A0L7RBR5 A0A154PG84 A0A2J7Q593 A0A2J7Q595 A0A2J7Q590 A0A2J7Q587 A0A2J7Q592 A0A2J7Q599 A0A0T6B6K4 A0A3L8DB04 A0A067QM30 Q16ED7 A0A182MFP5 A0A023ET03 A0A1S4G417 A0A1B6HGF5 A0A2M3Z6Y0 A0A1Q3EUA4 A0A1Y1MVB0 A0A182PFR2 A0A2M4AIV6 A0A1B6ITM7 A0A131YJ98 A0A1B6JKY6 A0A1W7R6X7 A0A1B6IAT8 A0A2M4AJ23 A0A1Q3FDR1 Q16TH1 A0A182KC06 A0A182Y9R8 U5EVS4 B0WJ81 A0A2M4BF98 A0A2M4CK77 A0A182KUG7 A0A2M3Z358 T1D6B5 T1E1Y7 A0A2M4CIM2 A0A2M4AEI7 A0A2M4CKB8 A0A139WFZ9 A0A2M4AES6 A0A182WH63 A0A139WFM3 W5JFQ6 A0A139WFP1 A0A1B6E678 A0A1B6CKI5 A0A1B6CWM0 A0A1B6C7D3 A0A1B6LKU7 A0A1B6KNH3 A0A1B6EY88 A0A1B6KUV2 K7IVE5 A0A1B6KF32 A0A182NF42 E0VBM0 A0A182R6J4 A0A182J2D4 A0A1J1IIV5 A0A453Z004 A0A1J1IM69 A0A182I559 A0A194QZN8 A0A224XQJ1 N6UKR1 T1J4I1 T1HHL0 A0A423SC62 A0A023FMB7 A0A293MA48 A0A131Y2B5 A0A0P4VL11
Pubmed
EMBL
KY609557
ASO76421.1
LBMM01003447
KMQ93529.1
NWSH01001293
PCG71816.1
+ More
RSAL01000127 RVE46519.1 ODYU01001460 SOQ37685.1 ATLV01014632 KE524975 KFB39309.1 KQ982562 KYQ54831.1 GL436229 EFN72215.1 PYGN01000163 PSN52385.1 KQ978642 KYN29517.1 KQ981993 KYN31003.1 GL448322 EFN84793.1 KQ976543 KYM81004.1 KQ976881 KYN07602.1 ADTU01024787 ADTU01024788 GL887655 EGI70484.1 KK107874 EZA47349.1 KZ288247 PBC31176.1 KQ791045 OAD47036.1 GBYB01010570 JAG80337.1 GBYB01010572 JAG80339.1 GECU01032498 JAS75208.1 KQ414617 KOC68367.1 KQ434899 KZC10869.1 NEVH01017587 PNF23746.1 PNF23750.1 PNF23751.1 PNF23748.1 PNF23753.1 PNF23752.1 LJIG01009489 KRT82978.1 QOIP01000010 RLU17516.1 KK853162 KDR10437.1 CH478832 EAT32594.1 AXCM01001968 GAPW01001171 JAC12427.1 GECU01033960 JAS73746.1 GGFM01003531 MBW24282.1 GFDL01016150 JAV18895.1 GEZM01019595 JAV89601.1 GGFK01007395 MBW40716.1 GECU01017421 JAS90285.1 GEDV01009919 JAP78638.1 GECU01007856 JAS99850.1 GEHC01000741 JAV46904.1 GECU01023736 JAS83970.1 GGFK01007450 MBW40771.1 GFDL01009417 JAV25628.1 CH477649 EAT37821.1 GANO01003279 JAB56592.1 DS231957 EDS29006.1 GGFJ01002584 MBW51725.1 GGFL01001481 MBW65659.1 GGFM01002200 MBW22951.1 GALA01000138 JAA94714.1 GALA01001239 JAA93613.1 GGFL01000975 MBW65153.1 GGFK01005846 MBW39167.1 GGFL01001170 MBW65348.1 KQ971351 KYB26781.1 GGFK01005963 MBW39284.1 KYB26780.1 ADMH02001355 ETN62891.1 KYB26783.1 GEDC01003899 JAS33399.1 GEDC01023417 JAS13881.1 GEDC01019396 JAS17902.1 GEDC01027902 JAS09396.1 GEBQ01015681 JAT24296.1 GEBQ01026969 JAT13008.1 GECZ01026895 JAS42874.1 GEBQ01024748 JAT15229.1 AAZX01001175 GEBQ01029952 JAT10025.1 DS235033 EEB10776.1 CVRI01000054 CRL00175.1 AAAB01008984 CRL00174.1 APCN01003786 KQ460890 KPJ11008.1 GFTR01006165 JAW10261.1 APGK01019954 KB740150 KB632173 ENN81251.1 ERL89576.1 JH431845 ACPB03000431 ACPB03000432 QCYY01003988 ROT61799.1 GBBK01002522 JAC21960.1 GFWV01013104 MAA37833.1 GEFM01003286 JAP72510.1 GDKW01003558 JAI53037.1
RSAL01000127 RVE46519.1 ODYU01001460 SOQ37685.1 ATLV01014632 KE524975 KFB39309.1 KQ982562 KYQ54831.1 GL436229 EFN72215.1 PYGN01000163 PSN52385.1 KQ978642 KYN29517.1 KQ981993 KYN31003.1 GL448322 EFN84793.1 KQ976543 KYM81004.1 KQ976881 KYN07602.1 ADTU01024787 ADTU01024788 GL887655 EGI70484.1 KK107874 EZA47349.1 KZ288247 PBC31176.1 KQ791045 OAD47036.1 GBYB01010570 JAG80337.1 GBYB01010572 JAG80339.1 GECU01032498 JAS75208.1 KQ414617 KOC68367.1 KQ434899 KZC10869.1 NEVH01017587 PNF23746.1 PNF23750.1 PNF23751.1 PNF23748.1 PNF23753.1 PNF23752.1 LJIG01009489 KRT82978.1 QOIP01000010 RLU17516.1 KK853162 KDR10437.1 CH478832 EAT32594.1 AXCM01001968 GAPW01001171 JAC12427.1 GECU01033960 JAS73746.1 GGFM01003531 MBW24282.1 GFDL01016150 JAV18895.1 GEZM01019595 JAV89601.1 GGFK01007395 MBW40716.1 GECU01017421 JAS90285.1 GEDV01009919 JAP78638.1 GECU01007856 JAS99850.1 GEHC01000741 JAV46904.1 GECU01023736 JAS83970.1 GGFK01007450 MBW40771.1 GFDL01009417 JAV25628.1 CH477649 EAT37821.1 GANO01003279 JAB56592.1 DS231957 EDS29006.1 GGFJ01002584 MBW51725.1 GGFL01001481 MBW65659.1 GGFM01002200 MBW22951.1 GALA01000138 JAA94714.1 GALA01001239 JAA93613.1 GGFL01000975 MBW65153.1 GGFK01005846 MBW39167.1 GGFL01001170 MBW65348.1 KQ971351 KYB26781.1 GGFK01005963 MBW39284.1 KYB26780.1 ADMH02001355 ETN62891.1 KYB26783.1 GEDC01003899 JAS33399.1 GEDC01023417 JAS13881.1 GEDC01019396 JAS17902.1 GEDC01027902 JAS09396.1 GEBQ01015681 JAT24296.1 GEBQ01026969 JAT13008.1 GECZ01026895 JAS42874.1 GEBQ01024748 JAT15229.1 AAZX01001175 GEBQ01029952 JAT10025.1 DS235033 EEB10776.1 CVRI01000054 CRL00175.1 AAAB01008984 CRL00174.1 APCN01003786 KQ460890 KPJ11008.1 GFTR01006165 JAW10261.1 APGK01019954 KB740150 KB632173 ENN81251.1 ERL89576.1 JH431845 ACPB03000431 ACPB03000432 QCYY01003988 ROT61799.1 GBBK01002522 JAC21960.1 GFWV01013104 MAA37833.1 GEFM01003286 JAP72510.1 GDKW01003558 JAI53037.1
Proteomes
UP000036403
UP000218220
UP000283053
UP000030765
UP000075809
UP000000311
+ More
UP000245037 UP000078492 UP000078541 UP000008237 UP000078540 UP000078542 UP000005205 UP000007755 UP000053097 UP000005203 UP000242457 UP000075901 UP000053825 UP000076502 UP000235965 UP000279307 UP000027135 UP000008820 UP000075883 UP000075885 UP000075881 UP000076408 UP000002320 UP000075882 UP000007266 UP000075920 UP000000673 UP000002358 UP000075884 UP000009046 UP000075900 UP000075880 UP000183832 UP000075840 UP000053240 UP000019118 UP000030742 UP000015103 UP000283509
UP000245037 UP000078492 UP000078541 UP000008237 UP000078540 UP000078542 UP000005205 UP000007755 UP000053097 UP000005203 UP000242457 UP000075901 UP000053825 UP000076502 UP000235965 UP000279307 UP000027135 UP000008820 UP000075883 UP000075885 UP000075881 UP000076408 UP000002320 UP000075882 UP000007266 UP000075920 UP000000673 UP000002358 UP000075884 UP000009046 UP000075900 UP000075880 UP000183832 UP000075840 UP000053240 UP000019118 UP000030742 UP000015103 UP000283509
Pfam
PF01504 PIP5K
Interpro
Gene 3D
ProteinModelPortal
A0A222AJ73
A0A0J7KTF7
A0A2A4JJ66
A0A3S2LXW5
A0A2H1VA20
A0A084VMW5
+ More
A0A151X330 E2A2X3 A0A2P8Z7B0 A0A151JRA7 A0A195ERZ8 E2BHP5 A0A151I2H8 A0A195D3Y7 A0A158NSF4 F4W5I7 A0A026VWW3 A0A088AUC9 A0A2A3EJ11 A0A310SBV2 A0A0C9RPZ2 A0A0C9RBR0 A0A1B6HKL2 A0A182SH93 A0A0L7RBR5 A0A154PG84 A0A2J7Q593 A0A2J7Q595 A0A2J7Q590 A0A2J7Q587 A0A2J7Q592 A0A2J7Q599 A0A0T6B6K4 A0A3L8DB04 A0A067QM30 Q16ED7 A0A182MFP5 A0A023ET03 A0A1S4G417 A0A1B6HGF5 A0A2M3Z6Y0 A0A1Q3EUA4 A0A1Y1MVB0 A0A182PFR2 A0A2M4AIV6 A0A1B6ITM7 A0A131YJ98 A0A1B6JKY6 A0A1W7R6X7 A0A1B6IAT8 A0A2M4AJ23 A0A1Q3FDR1 Q16TH1 A0A182KC06 A0A182Y9R8 U5EVS4 B0WJ81 A0A2M4BF98 A0A2M4CK77 A0A182KUG7 A0A2M3Z358 T1D6B5 T1E1Y7 A0A2M4CIM2 A0A2M4AEI7 A0A2M4CKB8 A0A139WFZ9 A0A2M4AES6 A0A182WH63 A0A139WFM3 W5JFQ6 A0A139WFP1 A0A1B6E678 A0A1B6CKI5 A0A1B6CWM0 A0A1B6C7D3 A0A1B6LKU7 A0A1B6KNH3 A0A1B6EY88 A0A1B6KUV2 K7IVE5 A0A1B6KF32 A0A182NF42 E0VBM0 A0A182R6J4 A0A182J2D4 A0A1J1IIV5 A0A453Z004 A0A1J1IM69 A0A182I559 A0A194QZN8 A0A224XQJ1 N6UKR1 T1J4I1 T1HHL0 A0A423SC62 A0A023FMB7 A0A293MA48 A0A131Y2B5 A0A0P4VL11
A0A151X330 E2A2X3 A0A2P8Z7B0 A0A151JRA7 A0A195ERZ8 E2BHP5 A0A151I2H8 A0A195D3Y7 A0A158NSF4 F4W5I7 A0A026VWW3 A0A088AUC9 A0A2A3EJ11 A0A310SBV2 A0A0C9RPZ2 A0A0C9RBR0 A0A1B6HKL2 A0A182SH93 A0A0L7RBR5 A0A154PG84 A0A2J7Q593 A0A2J7Q595 A0A2J7Q590 A0A2J7Q587 A0A2J7Q592 A0A2J7Q599 A0A0T6B6K4 A0A3L8DB04 A0A067QM30 Q16ED7 A0A182MFP5 A0A023ET03 A0A1S4G417 A0A1B6HGF5 A0A2M3Z6Y0 A0A1Q3EUA4 A0A1Y1MVB0 A0A182PFR2 A0A2M4AIV6 A0A1B6ITM7 A0A131YJ98 A0A1B6JKY6 A0A1W7R6X7 A0A1B6IAT8 A0A2M4AJ23 A0A1Q3FDR1 Q16TH1 A0A182KC06 A0A182Y9R8 U5EVS4 B0WJ81 A0A2M4BF98 A0A2M4CK77 A0A182KUG7 A0A2M3Z358 T1D6B5 T1E1Y7 A0A2M4CIM2 A0A2M4AEI7 A0A2M4CKB8 A0A139WFZ9 A0A2M4AES6 A0A182WH63 A0A139WFM3 W5JFQ6 A0A139WFP1 A0A1B6E678 A0A1B6CKI5 A0A1B6CWM0 A0A1B6C7D3 A0A1B6LKU7 A0A1B6KNH3 A0A1B6EY88 A0A1B6KUV2 K7IVE5 A0A1B6KF32 A0A182NF42 E0VBM0 A0A182R6J4 A0A182J2D4 A0A1J1IIV5 A0A453Z004 A0A1J1IM69 A0A182I559 A0A194QZN8 A0A224XQJ1 N6UKR1 T1J4I1 T1HHL0 A0A423SC62 A0A023FMB7 A0A293MA48 A0A131Y2B5 A0A0P4VL11
PDB
6CN2
E-value=1.47913e-28,
Score=307
Ontologies
GO
PANTHER
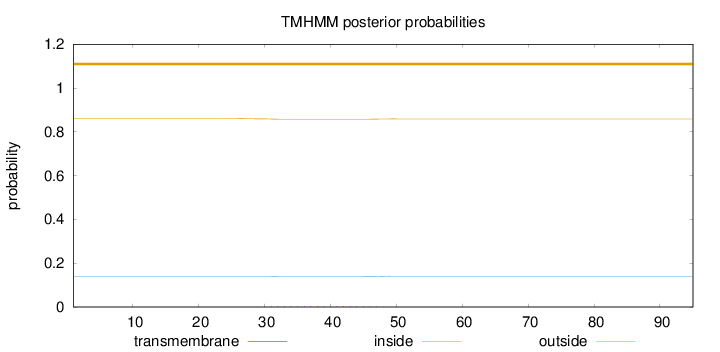
Topology
Length:
95
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.06752
Exp number, first 60 AAs:
0.06751
Total prob of N-in:
0.13953
outside
1 - 95
Population Genetic Test Statistics
Pi
0
Theta
0
Tajima's D
0
CLR
0
CSRT
0
Interpretation
Uncertain
