Gene
KWMTBOMO16786
Annotation
Putative_115_kDa_protein_in_type-1_retrotransposable_element_R1DM-like_Protein_[Tribolium_castaneum]
Location in the cell
Cytoplasmic Reliability : 1.621 Mitochondrial Reliability : 1.857
Sequence
CDS
ATGGGCATGGCACAAATTCAAAGCATAGCTGATGGCCTCTTCCCGGAACACATAGATTTCGAACGGAAAACGAAACCGTCACAGGAGTTCCCAGAGGTGACCGAGGAGGAGATCATCCAACTAAAAGCTCGCATAAAGCTCAGGAAAGCCCCAGGGCCCGACCGGATTCCGCCGGAGATAGTGACTCTAGCAGTAGCAGTAATTCCAGACAGGATAAGGCTTGTGTTGCAGAAGGTGCTGGAGTCTGGCGTCTTCCCCAACAGCTGGAAAACAGCCAGACTAGTCCTGCTAAGAAAACCAGGCAAACCCGCTGAAGATCCTGCGGGGCACAGGCCGCTATGTCTACTGGACTGCTACGGAAAGCTGCTGGAGTCGCTGATCCTAGGCAGAATAGAAAAACAACTAGTACAATCGGACTGTTTACAAGATTCGCAGTACGGGTTCAGAAAGGGGAGGTCCACAGTGGATGCAGCCAACAAGCTACTTGAGATAGCGGAAGACGCGAGACGAGGAACCCACCGCACCAGAAGAATCTGCGCAGTAGTGGCCCTCGACGTCCGTAATGCTTTCAATTCTGCATCGTGGGAGCTCATCCTGGAGGAAATGGCTGCAATAGGTATGCCTCAATACATCTATAATATCATAAACAGCTACCTCCAGGACAGATGCATCATACTGACAGACGCTGAGGGAGGGGTTGTAGTGAAAAAGGTTACCGGTGGTGTTCCGCAGGGGTCGATACTGGGCCCCACGCTGTGGAACATACTGTATGATGGGGTACTGCGCCTCGAACTGGGGGATGGGGCACAATTAATAGGCTTCGCGGACGATTTAGCTCTGGTGGTGTCAGCAAAGAAGGAAGCAGAACTAATGGCAATAACGAATGTAGCCCTACAAAAGATCAGCGCTTGGATGAAGCAAAAGCGGCTACACTTGGCCCCAGAAAAGACCGAGGCAATTGTGCTAAGTGGGCGAAGAAAGCTTAGCGAGATTTGCTTTACCCTCCAGGGCGTCAGGGTGGTTCCAACCGACAAACTGAAATACCTCGGGTTTTGGTTTGACAAAAGTCTGAAGTTCGGCAAACACATTGAAGAAACAACTGGTAAAGCCGAAAGACTGACGACCGCCCTTGGCCAAATCATGCCCAGGAAAGGAGGGCCGAGGTCGAGCAAGCGCAGACTATTGGCGTCCGCAATACAATCTGTGATGTTATATGGAGCACCGGTGTGGGCAAAGGTAATGAATGTAGCGAAGTACAGGAACATGTATAGCAAAGTGCAACGACAGCTGGCAATACGCATCTGTGGAGCTTACCGTACCATATCGCTAGACGCCGTCTTGGTGATAGCGGGCCTGATACCGATAGAACGGCTAGCAAAGGAACGGCAACGCTACTACCACGAGGGGACAAGAACGAGAGAAGAGGAAAGAGCACATACAATAAAAGAATGGGAACGAGACTGGCAGAAGGAGGGAAAAGGTAAGTGGACTCGTACAATCATACCCAACCTTAGGCTCTGGATGGAACGGAAACATGGCGAAACCGACAGACTGCTGACACAGGCCCTAAGTGGGCATGGAGTTTTCAACACTTACCTGCATTCAATAGGGAAAGCAGAAAGCGCAAGTTGCCCATACTGCGAACATGTTGATACACCGGAACATGCCCTATTTCAATGCATAAAGTTCACAGAACTAAGAGGGAAAACGGAGGGAAGTGACGGAAAGGTGACTGTGGAAAACCTGATGGTGAGAATGATCCATAGTAAAGAAGAATGGGATAAATATGCGGGAATGCTCAAAGAGATAATGAGAGTGCGGGAAAAGGACGAAAGACTAAGGCAAGAATCTATCTATAATAATATATAA
Protein
MGMAQIQSIADGLFPEHIDFERKTKPSQEFPEVTEEEIIQLKARIKLRKAPGPDRIPPEIVTLAVAVIPDRIRLVLQKVLESGVFPNSWKTARLVLLRKPGKPAEDPAGHRPLCLLDCYGKLLESLILGRIEKQLVQSDCLQDSQYGFRKGRSTVDAANKLLEIAEDARRGTHRTRRICAVVALDVRNAFNSASWELILEEMAAIGMPQYIYNIINSYLQDRCIILTDAEGGVVVKKVTGGVPQGSILGPTLWNILYDGVLRLELGDGAQLIGFADDLALVVSAKKEAELMAITNVALQKISAWMKQKRLHLAPEKTEAIVLSGRRKLSEICFTLQGVRVVPTDKLKYLGFWFDKSLKFGKHIEETTGKAERLTTALGQIMPRKGGPRSSKRRLLASAIQSVMLYGAPVWAKVMNVAKYRNMYSKVQRQLAIRICGAYRTISLDAVLVIAGLIPIERLAKERQRYYHEGTRTREEERAHTIKEWERDWQKEGKGKWTRTIIPNLRLWMERKHGETDRLLTQALSGHGVFNTYLHSIGKAESASCPYCEHVDTPEHALFQCIKFTELRGKTEGSDGKVTVENLMVRMIHSKEEWDKYAGMLKEIMRVREKDERLRQESIYNNI
Summary
Uniprot
D7GXX9
D6WC19
A0A1W7R6C5
J9KJG8
F7IYV2
F7IYU8
+ More
F7IYV5 A0A142LX45 T1DG89 T1DG59 T1DCM0 A0A2H8TR36 A0A139WAH4 V5I881 A0A1W7R6I9 A0A139W9J0 A0A023EW96 J9KUM6 V5I8F3 F7IYV3 A0A139W940 A0A139W9A6 A0A139W9B3 A0A139WNC9 A0A139WN93 A0A139W939 A0A2M4ADW9 A0A2M4CS98 J9L6Z9 A0A224XIZ2 N6TYN9 W8ADE4 W8ANK7 A0A2H8TIE5 A0A2M4BDI3 A0A034WR48 A0A2M4BBV9 A0A2M4CSA9 A0A142LX37 A0A2M4BC01 A0A2S2P488 X1X6F1 A0A2M4CKQ7 J9KV04 X1WZ64 J9LVU7 K7JP42 J9KM84 A0A2M4CSI6 F7IYV6 A0A2M4BC28 A0A2M4BC24 A0A2M4BC18 A0A142LX43 A0A0E3W2C5 Q868S0 A0A0K8VPH1 J9KT70 Q868T0 F7IYV7 Q868S4 N6UIP2 J9KPP8 D7EIB0 A0A2S2N8I2 A0A1B6KDF3 A0A224XA82 V5G2Q2 A0A146MAT2 Q868R6 A0A139W8J5 A0A2S2NJ82 V9GZD9 J9JWJ3 A0A2M3Z1C1 J9KT61 A0A2M4BB78 A0A2M4BB89 A0A139W942 A0A2S2NQQ6 Q868S6 A0A139W9I2 A0A2M4CJ33 N6UHH2 A0A2M4BBA0 A0A139WA31 Q868S8 J9LJ80 A0A2M4AK35 J9KNG8 Q868R2 A0A2S2PRL8 Q868Q8 A0A1S4EMG7 Q868R4 A0A3Q0JIB3 J9LM37 A0A2M4CKD9 Q868S2 A0A2M4CJ59
F7IYV5 A0A142LX45 T1DG89 T1DG59 T1DCM0 A0A2H8TR36 A0A139WAH4 V5I881 A0A1W7R6I9 A0A139W9J0 A0A023EW96 J9KUM6 V5I8F3 F7IYV3 A0A139W940 A0A139W9A6 A0A139W9B3 A0A139WNC9 A0A139WN93 A0A139W939 A0A2M4ADW9 A0A2M4CS98 J9L6Z9 A0A224XIZ2 N6TYN9 W8ADE4 W8ANK7 A0A2H8TIE5 A0A2M4BDI3 A0A034WR48 A0A2M4BBV9 A0A2M4CSA9 A0A142LX37 A0A2M4BC01 A0A2S2P488 X1X6F1 A0A2M4CKQ7 J9KV04 X1WZ64 J9LVU7 K7JP42 J9KM84 A0A2M4CSI6 F7IYV6 A0A2M4BC28 A0A2M4BC24 A0A2M4BC18 A0A142LX43 A0A0E3W2C5 Q868S0 A0A0K8VPH1 J9KT70 Q868T0 F7IYV7 Q868S4 N6UIP2 J9KPP8 D7EIB0 A0A2S2N8I2 A0A1B6KDF3 A0A224XA82 V5G2Q2 A0A146MAT2 Q868R6 A0A139W8J5 A0A2S2NJ82 V9GZD9 J9JWJ3 A0A2M3Z1C1 J9KT61 A0A2M4BB78 A0A2M4BB89 A0A139W942 A0A2S2NQQ6 Q868S6 A0A139W9I2 A0A2M4CJ33 N6UHH2 A0A2M4BBA0 A0A139WA31 Q868S8 J9LJ80 A0A2M4AK35 J9KNG8 Q868R2 A0A2S2PRL8 Q868Q8 A0A1S4EMG7 Q868R4 A0A3Q0JIB3 J9LM37 A0A2M4CKD9 Q868S2 A0A2M4CJ59
Pubmed
EMBL
KQ972160
EFA13619.2
KQ971309
EEZ99146.1
GEHC01000964
JAV46681.1
+ More
ABLF02041886 AB593323 BAK38643.1 AB593321 BAK38639.1 AB593325 BAK38646.1 KU543683 AMS38371.1 GALA01001776 JAA93076.1 GALA01000302 JAA94550.1 GALA01001777 JAA93075.1 GFXV01003903 MBW15708.1 KQ971388 KYB24913.1 GALX01005145 JAB63321.1 GEHC01000915 JAV46730.1 KQ972226 KYB24577.1 GAPW01000192 JAC13406.1 ABLF02041557 GALX01004785 JAB63681.1 AB593324 BAK38644.1 KQ972691 KXZ75806.1 KQ972544 KXZ75870.1 KXZ75869.1 KQ971310 KYB29500.1 KYB29499.1 KXZ75807.1 GGFK01005660 MBW38981.1 GGFL01004044 MBW68222.1 ABLF02021100 GFTR01008427 JAW07999.1 APGK01005465 KB736249 ENN83233.1 GAMC01020519 GAMC01020515 JAB86040.1 GAMC01020517 JAB86038.1 GFXV01002090 MBW13895.1 GGFJ01001727 MBW50868.1 GAKP01002292 JAC56660.1 GGFJ01001369 MBW50510.1 GGFL01004046 MBW68224.1 KU543679 AMS38363.1 GGFJ01001370 MBW50511.1 GGMR01011682 MBY24301.1 ABLF02016340 GGFL01001583 MBW65761.1 ABLF02004674 ABLF02004675 ABLF02004676 ABLF02011637 ABLF02011183 ABLF02041884 ABLF02034925 AAZX01012143 AAZX01012194 ABLF02018942 ABLF02041885 GGFL01004047 MBW68225.1 AB593326 BAK38647.1 GGFJ01001439 MBW50580.1 GGFJ01001454 MBW50595.1 GGFJ01001453 MBW50594.1 KU543682 AMS38369.1 HACL01000281 CFW94575.1 AB090817 BAC57910.1 GDHF01029467 GDHF01011533 JAI22847.1 JAI40781.1 ABLF02010223 AB090812 BAC57900.1 AB593327 BAK38648.1 AB090815 BAC57906.1 APGK01033685 KB740885 ENN78512.1 ABLF02009991 ABLF02009992 ABLF02054952 AB593322 BAK38641.1 GGMR01000779 MBY13398.1 GEBQ01030504 JAT09473.1 GFTR01007160 JAW09266.1 GALX01004171 JAB64295.1 GDHC01002080 JAQ16549.1 AB090819 BAC57914.1 KQ973256 KXZ75595.1 GGMR01004600 MBY17219.1 M93691 AAA29365.1 ABLF02013358 ABLF02013361 ABLF02054869 GGFM01001554 MBW22305.1 ABLF02009073 ABLF02030702 ABLF02042963 GGFJ01001164 MBW50305.1 GGFJ01001165 MBW50306.1 KXZ75805.1 GGMR01006838 MBY19457.1 AB090814 BAC57904.1 KQ972255 KYB24568.1 GGFL01001192 MBW65370.1 APGK01020183 KB740172 ENN81190.1 GGFJ01001166 MBW50307.1 KQ971675 KYB24771.1 AB090813 BAC57902.1 ABLF02015311 ABLF02015320 ABLF02034009 ABLF02043484 GGFK01007771 MBW41092.1 ABLF02005203 ABLF02041316 AB090821 BAC57918.1 GGMR01019493 MBY32112.1 AB090823 BAC57922.1 AB090820 BAC57916.1 ABLF02014650 GGFL01001190 MBW65368.1 AB090816 BAC57908.1 GGFL01001189 MBW65367.1
ABLF02041886 AB593323 BAK38643.1 AB593321 BAK38639.1 AB593325 BAK38646.1 KU543683 AMS38371.1 GALA01001776 JAA93076.1 GALA01000302 JAA94550.1 GALA01001777 JAA93075.1 GFXV01003903 MBW15708.1 KQ971388 KYB24913.1 GALX01005145 JAB63321.1 GEHC01000915 JAV46730.1 KQ972226 KYB24577.1 GAPW01000192 JAC13406.1 ABLF02041557 GALX01004785 JAB63681.1 AB593324 BAK38644.1 KQ972691 KXZ75806.1 KQ972544 KXZ75870.1 KXZ75869.1 KQ971310 KYB29500.1 KYB29499.1 KXZ75807.1 GGFK01005660 MBW38981.1 GGFL01004044 MBW68222.1 ABLF02021100 GFTR01008427 JAW07999.1 APGK01005465 KB736249 ENN83233.1 GAMC01020519 GAMC01020515 JAB86040.1 GAMC01020517 JAB86038.1 GFXV01002090 MBW13895.1 GGFJ01001727 MBW50868.1 GAKP01002292 JAC56660.1 GGFJ01001369 MBW50510.1 GGFL01004046 MBW68224.1 KU543679 AMS38363.1 GGFJ01001370 MBW50511.1 GGMR01011682 MBY24301.1 ABLF02016340 GGFL01001583 MBW65761.1 ABLF02004674 ABLF02004675 ABLF02004676 ABLF02011637 ABLF02011183 ABLF02041884 ABLF02034925 AAZX01012143 AAZX01012194 ABLF02018942 ABLF02041885 GGFL01004047 MBW68225.1 AB593326 BAK38647.1 GGFJ01001439 MBW50580.1 GGFJ01001454 MBW50595.1 GGFJ01001453 MBW50594.1 KU543682 AMS38369.1 HACL01000281 CFW94575.1 AB090817 BAC57910.1 GDHF01029467 GDHF01011533 JAI22847.1 JAI40781.1 ABLF02010223 AB090812 BAC57900.1 AB593327 BAK38648.1 AB090815 BAC57906.1 APGK01033685 KB740885 ENN78512.1 ABLF02009991 ABLF02009992 ABLF02054952 AB593322 BAK38641.1 GGMR01000779 MBY13398.1 GEBQ01030504 JAT09473.1 GFTR01007160 JAW09266.1 GALX01004171 JAB64295.1 GDHC01002080 JAQ16549.1 AB090819 BAC57914.1 KQ973256 KXZ75595.1 GGMR01004600 MBY17219.1 M93691 AAA29365.1 ABLF02013358 ABLF02013361 ABLF02054869 GGFM01001554 MBW22305.1 ABLF02009073 ABLF02030702 ABLF02042963 GGFJ01001164 MBW50305.1 GGFJ01001165 MBW50306.1 KXZ75805.1 GGMR01006838 MBY19457.1 AB090814 BAC57904.1 KQ972255 KYB24568.1 GGFL01001192 MBW65370.1 APGK01020183 KB740172 ENN81190.1 GGFJ01001166 MBW50307.1 KQ971675 KYB24771.1 AB090813 BAC57902.1 ABLF02015311 ABLF02015320 ABLF02034009 ABLF02043484 GGFK01007771 MBW41092.1 ABLF02005203 ABLF02041316 AB090821 BAC57918.1 GGMR01019493 MBY32112.1 AB090823 BAC57922.1 AB090820 BAC57916.1 ABLF02014650 GGFL01001190 MBW65368.1 AB090816 BAC57908.1 GGFL01001189 MBW65367.1
Proteomes
Pfam
Interpro
Gene 3D
ProteinModelPortal
D7GXX9
D6WC19
A0A1W7R6C5
J9KJG8
F7IYV2
F7IYU8
+ More
F7IYV5 A0A142LX45 T1DG89 T1DG59 T1DCM0 A0A2H8TR36 A0A139WAH4 V5I881 A0A1W7R6I9 A0A139W9J0 A0A023EW96 J9KUM6 V5I8F3 F7IYV3 A0A139W940 A0A139W9A6 A0A139W9B3 A0A139WNC9 A0A139WN93 A0A139W939 A0A2M4ADW9 A0A2M4CS98 J9L6Z9 A0A224XIZ2 N6TYN9 W8ADE4 W8ANK7 A0A2H8TIE5 A0A2M4BDI3 A0A034WR48 A0A2M4BBV9 A0A2M4CSA9 A0A142LX37 A0A2M4BC01 A0A2S2P488 X1X6F1 A0A2M4CKQ7 J9KV04 X1WZ64 J9LVU7 K7JP42 J9KM84 A0A2M4CSI6 F7IYV6 A0A2M4BC28 A0A2M4BC24 A0A2M4BC18 A0A142LX43 A0A0E3W2C5 Q868S0 A0A0K8VPH1 J9KT70 Q868T0 F7IYV7 Q868S4 N6UIP2 J9KPP8 D7EIB0 A0A2S2N8I2 A0A1B6KDF3 A0A224XA82 V5G2Q2 A0A146MAT2 Q868R6 A0A139W8J5 A0A2S2NJ82 V9GZD9 J9JWJ3 A0A2M3Z1C1 J9KT61 A0A2M4BB78 A0A2M4BB89 A0A139W942 A0A2S2NQQ6 Q868S6 A0A139W9I2 A0A2M4CJ33 N6UHH2 A0A2M4BBA0 A0A139WA31 Q868S8 J9LJ80 A0A2M4AK35 J9KNG8 Q868R2 A0A2S2PRL8 Q868Q8 A0A1S4EMG7 Q868R4 A0A3Q0JIB3 J9LM37 A0A2M4CKD9 Q868S2 A0A2M4CJ59
F7IYV5 A0A142LX45 T1DG89 T1DG59 T1DCM0 A0A2H8TR36 A0A139WAH4 V5I881 A0A1W7R6I9 A0A139W9J0 A0A023EW96 J9KUM6 V5I8F3 F7IYV3 A0A139W940 A0A139W9A6 A0A139W9B3 A0A139WNC9 A0A139WN93 A0A139W939 A0A2M4ADW9 A0A2M4CS98 J9L6Z9 A0A224XIZ2 N6TYN9 W8ADE4 W8ANK7 A0A2H8TIE5 A0A2M4BDI3 A0A034WR48 A0A2M4BBV9 A0A2M4CSA9 A0A142LX37 A0A2M4BC01 A0A2S2P488 X1X6F1 A0A2M4CKQ7 J9KV04 X1WZ64 J9LVU7 K7JP42 J9KM84 A0A2M4CSI6 F7IYV6 A0A2M4BC28 A0A2M4BC24 A0A2M4BC18 A0A142LX43 A0A0E3W2C5 Q868S0 A0A0K8VPH1 J9KT70 Q868T0 F7IYV7 Q868S4 N6UIP2 J9KPP8 D7EIB0 A0A2S2N8I2 A0A1B6KDF3 A0A224XA82 V5G2Q2 A0A146MAT2 Q868R6 A0A139W8J5 A0A2S2NJ82 V9GZD9 J9JWJ3 A0A2M3Z1C1 J9KT61 A0A2M4BB78 A0A2M4BB89 A0A139W942 A0A2S2NQQ6 Q868S6 A0A139W9I2 A0A2M4CJ33 N6UHH2 A0A2M4BBA0 A0A139WA31 Q868S8 J9LJ80 A0A2M4AK35 J9KNG8 Q868R2 A0A2S2PRL8 Q868Q8 A0A1S4EMG7 Q868R4 A0A3Q0JIB3 J9LM37 A0A2M4CKD9 Q868S2 A0A2M4CJ59
PDB
6AR3
E-value=0.00874827,
Score=94
Ontologies
GO
Topology
Length:
622
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
14.22548
Exp number, first 60 AAs:
0.00021
Total prob of N-in:
0.00567
outside
1 - 622
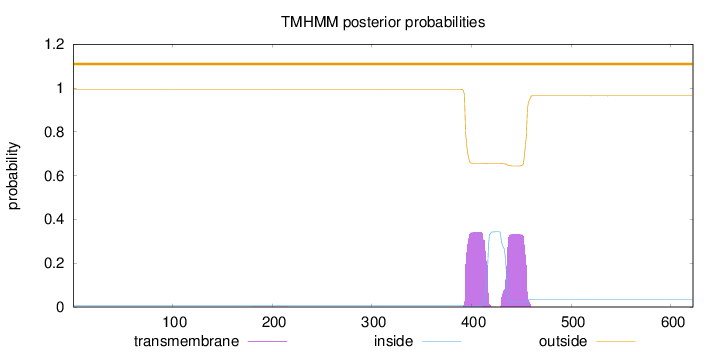
Population Genetic Test Statistics
Pi
0.241085
Theta
0.245856
Tajima's D
-0.167928
CLR
0
CSRT
0.502924853757312
Interpretation
Uncertain
