Gene
KWMTBOMO16761
Pre Gene Modal
BGIBMGA005642
Annotation
PREDICTED:_LOW_QUALITY_PROTEIN:_cytokine_receptor_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 2.236
Sequence
CDS
ATGAAATATGAAATCAGGATCTACATCAAATCTAAAGCTGCAGTGAAAGAAGAGTTTTGGTCAGACTTCACATATACCATGGTATATACACTTAGCGAGAGACCTAGAAGACCACCAGACACCGCAGTTGGAGCCTTTGATCAATCTTTATATCAAACAAATAGAATGATATACATTTATTGGAAACAACTTGAAGAATATGAAGAGGCAGGCCGAATGAACTTCATTGTTTTAAGTGCCAATAAAACAATGCAAGTCATAGAACTACCTAGAGAAAACAGGTATCAGTTTGCCATATCGGCTAATAATGGGACGAAAACTAGTGGCATGGTATGGGCTATCTGTGATATATCTAAAGATGCAATTCCAATGTATGGTTTTCCGGTCAAAATGGACCATGGTATTCCAGGGAAAACTTTTGTAAAGATTAAATGGGCAATGACTTGTACACTTCAAGAAGGAATCATTACAGGATATTTGATCGCATATTGTCCTGCTCTAGATACCACTAATGTTTGTGCAACAAAAAATAAAACTAACTTATACATATCTAATCCTAAGCAAATGGAGATCAACATTACAAATCTTGAACCATACACTACATATATATTTACACTGGCATTGAACACCACTTATGGATTTAAAACAGTAGAAAATGCTTTCACGAGTATGACTACAACTGAAGATACTCCAACTAGTCCAGTTAATATTAATATAACCGATGTGCTTAGTGATTCATTGACGATTTCATGGGACCCACCTATTCATAAAAACGGAGTTATTGGAAAATATGTCATAAATAACTATGGTCAAGAAGTTTATGTAGAAAAAGTTTCAGATAAAGATGAAGATTCAAAAAGAAGGCATATTCGTTTGACTGGCTTACAGGGGTTTACTAATTATTCATTGAGTGTACAAGCATGTAGTACAGCTATAAGCAGTTGTTCGGTTTTGAATCCACATGATGCCCTTTTTGTCAGGACGAGAATAGGTCCTCCTAGTAAAATGAGGGCTCCAACTGTTAAAAATAATCCAGATTTTATTAAGTGGGTTCCACCAATAATACCAGGGGGGACGATTGACCTTTACCAAATTAAAAGGATTAAAGATGAGAGTGAAGCTGAAATAATAAATACTACAAACCTTTCATTTTCATTGATACATTGTGAAGGTGGAGTTGCCGGACAAACTTACCAAGTTAGAGCTGTTAATTTTGATGAAGATCCTTACCACGGTGCTTTAACTGACAATAAGGATGTATACCTACATAATGCTAACTATAAAGGTATACTAGAATATCCTGGAGAATGGAGTGAGCCAAGTTTTGTAAATTGTACAAGCAAAGAACATGTGACACTAACTCTTATTTTAATGGGAATATTTATGATAATTGGTATTATGTATGGTTTTATTAAATGTTACAAGAAATACCGAAAAATGGAAGATATTAAACCAGTTTTACCAAGTGGACTAGGAGTCCCCGAAAAGGATATTTCGAAATATGCTATTGGTGGTTGGAATCCTACAAACAAGGATGAGAAACCATCTTCAGACGAAATGTTGCTATTACCAAATAGTAGAACCACTGTATCTTCAGAATTGAAACAGAAAGAAAACAATAACAGTGCTTCTAGTGATCACACTGATAGTACAGCTTTATCAGATACATCACATGGGCCTGTAGAAAGACAAATCTCCTCATCTGATGATGGATCTAATTCATCACTACATCTTGAAGTAGAACCTACGAGAACAGCTGACAGTAACATCGATGGCGAATCATGCAATTCCGAAACTGAAACATCACAAATACCGTCACCATTCTTTAATGATAATACTTTCAAGAAAAATCCATGTGGCTATGTGCAACAATCTGTAGTGAGTCCTAATACAGGATATGTTCAAAGTGTTCAAGCAGGCGTTACAAACCCACCACAACCCTCATCACTCAGTGCCAGCTCTAGTTATGTAATGGCAGCTCTGTCTCCACCTATATTTACGACACGTGTTGCTCAACCTTGCGTAACGAGTTCACAATCAGCAGGAGGCTATGTGCACCCTGAAGGTGCTCAAACTATATCAGCAATGAATTTCCCGAAATTGGGGCAGACAACGAACAAACTGTTTGGTCCTGAAAGCTTGCCAACAATGCAAACCTTGCCACCAGCGAAACACACTGCAGACAGCAGCTACATTCAACTGCAATCTCTAGATTCCTTGCCCAGACACAAACATCCTGCCCGAAGTACAGTGCCATTAAAGACGCCAGCGTCTACCGGCTACGTCCGCCAAGGTGATCATAGGAAGCTTTATAGCGACTTGAGCACTGAAGAAGCCATCCTTCAGATTGCTGACACTATTGATAATCTCAATGGGATAGTGGAAGACGTGTTCTCTCGCTTAACGCGGCGAATCAAGCAGAATGTAGATAAAACCGCAAAATTGAATGAACGCATCGAAAACTCTAGGCTTAAAGTTGAAAAACTGACTGGAATGCAGAGAGCCATCAAAGTAATACAGGAGAAGCTACACTTCTATCATGTTAAAGTTGCGGAACCTAATTCACTCAAGCCCGATGAAGATTTTAGTTTGAACGCTGTTATTGATAACCTGGAAACCATCGGTGAACTTTTAGTATTTAACACCGATGAAAGCCCTTACTTCGGTGTTACCTCTAAAACACCAACCTACAAACCGGTCATCAAGAAATCGAATATTGAAAATAAGAGTCTAATAGAAGCGGCACCTTCGTCTATAATGAAGAAAGACGTCTTGAAGCGCGAAGTAGACGAATACATGTACGCACCTGGGATGGGAATGGTACCCGAAATGGAGATGCCGTTGGATTTACCGCATCTTCCAGGAATCGCTGAGGACATACAATATACTATCGCGACTGAAGGGTCTATAGCCCCGTCTGTCGTGACGTCACCGGTCACCGCCGATCATCGCCATTCCTGA
Protein
MKYEIRIYIKSKAAVKEEFWSDFTYTMVYTLSERPRRPPDTAVGAFDQSLYQTNRMIYIYWKQLEEYEEAGRMNFIVLSANKTMQVIELPRENRYQFAISANNGTKTSGMVWAICDISKDAIPMYGFPVKMDHGIPGKTFVKIKWAMTCTLQEGIITGYLIAYCPALDTTNVCATKNKTNLYISNPKQMEINITNLEPYTTYIFTLALNTTYGFKTVENAFTSMTTTEDTPTSPVNINITDVLSDSLTISWDPPIHKNGVIGKYVINNYGQEVYVEKVSDKDEDSKRRHIRLTGLQGFTNYSLSVQACSTAISSCSVLNPHDALFVRTRIGPPSKMRAPTVKNNPDFIKWVPPIIPGGTIDLYQIKRIKDESEAEIINTTNLSFSLIHCEGGVAGQTYQVRAVNFDEDPYHGALTDNKDVYLHNANYKGILEYPGEWSEPSFVNCTSKEHVTLTLILMGIFMIIGIMYGFIKCYKKYRKMEDIKPVLPSGLGVPEKDISKYAIGGWNPTNKDEKPSSDEMLLLPNSRTTVSSELKQKENNNSASSDHTDSTALSDTSHGPVERQISSSDDGSNSSLHLEVEPTRTADSNIDGESCNSETETSQIPSPFFNDNTFKKNPCGYVQQSVVSPNTGYVQSVQAGVTNPPQPSSLSASSSYVMAALSPPIFTTRVAQPCVTSSQSAGGYVHPEGAQTISAMNFPKLGQTTNKLFGPESLPTMQTLPPAKHTADSSYIQLQSLDSLPRHKHPARSTVPLKTPASTGYVRQGDHRKLYSDLSTEEAILQIADTIDNLNGIVEDVFSRLTRRIKQNVDKTAKLNERIENSRLKVEKLTGMQRAIKVIQEKLHFYHVKVAEPNSLKPDEDFSLNAVIDNLETIGELLVFNTDESPYFGVTSKTPTYKPVIKKSNIENKSLIEAAPSSIMKKDVLKREVDEYMYAPGMGMVPEMEMPLDLPHLPGIAEDIQYTIATEGSIAPSVVTSPVTADHRHS
Summary
Uniprot
EMBL
Proteomes
PRIDE
Pfam
PF00041 fn3
SUPFAM
SSF49265
SSF49265
Gene 3D
CDD
ProteinModelPortal
PDB
6IAA
E-value=0.00850776,
Score=96
Ontologies
GO
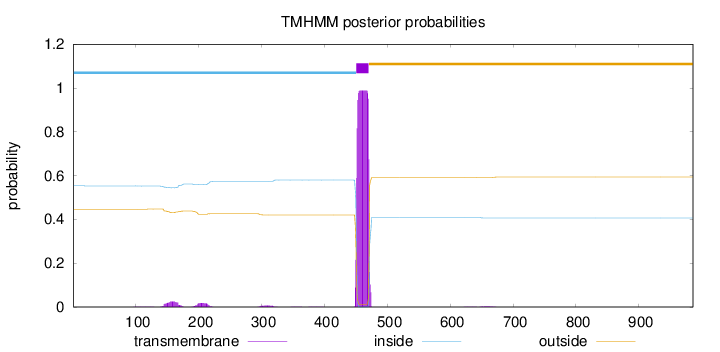
Topology
Length:
986
Number of predicted TMHs:
1
Exp number of AAs in TMHs:
21.51931
Exp number, first 60 AAs:
0.00057
Total prob of N-in:
0.55321
inside
1 - 450
TMhelix
451 - 470
outside
471 - 986
Population Genetic Test Statistics
Pi
0
Theta
0
Tajima's D
0
CLR
0
CSRT
0
Interpretation
Uncertain
