Gene
KWMTBOMO16710
Annotation
endonuclease-reverse_transcriptase_[Bombyx_mori]
Full name
Integrin beta
Location in the cell
Nuclear Reliability : 2.631
Sequence
CDS
ATGGGCGACTTTAATGCCAAAGTGGGTGTACAGATGTGTGACGAATCAAGAGTGGGACCACATGGATTTGGAAGCAGGAACGATAGGGGGCAAATGCTCGTTAACTTTCTAGAGCAAGAGGAGCTATTCTTAATGAACTCATTTTTTAAAAAGCAGCCCCAAAGAAAATGGACATGGCGAAGCCCGGACACTGTAACGAAAAATGAGATTGATTTTATCATTGCGGACAAAAAACGGTTATTTAGGGATGTCTCAGTGGTCAACAGGTTTAACACCGGAAGCGACCACCGGCTAGTAAGAGCTACGCTGAGCATAGACACTCGATTGGAAAGATACCGCATGATACAGTCTACACTCCGACCTAGCCAGCTCCAAATCAGTGCAGGTTCAGAAGTTTTCCATAAAACCTTGGAAAATCGATTTGTTAACTTGGAAACCACTGATGATGTTGACGAAAACCTCGAAAATCTGGTCACACTCATGAGGGAAGAGGGTGCAAAATTTTTCAGCGGGAAGCGAACGGGAAAAAAATCCGAACTCTCAGACGAGACGTTGCGACTCATGGCCATACGGCGACATTTCAAAAATTCTGCTTAA
Protein
MGDFNAKVGVQMCDESRVGPHGFGSRNDRGQMLVNFLEQEELFLMNSFFKKQPQRKWTWRSPDTVTKNEIDFIIADKKRLFRDVSVVNRFNTGSDHRLVRATLSIDTRLERYRMIQSTLRPSQLQISAGSEVFHKTLENRFVNLETTDDVDENLENLVTLMREEGAKFFSGKRTGKKSELSDETLRLMAIRRHFKNSA
Summary
Similarity
Belongs to the integrin beta chain family.
Feature
chain Integrin beta
Uniprot
D7F165
A0A3S2KZJ4
A0A437B0R4
D7F157
D7F160
D7F158
+ More
A0A2W1BHU6 D7F159 D7F164 D7F166 A0A3S2N805 A0A437BHD1 A0A3S2L6Q1 A0A016WTM3 A0A131YWK0 A0A131ZDH5 A0A016W0A5 A0A1W4UBD7 A0A016V9Q8 A0A016VJ99 A0A016VK03 A0A016TNL5 A0A1S3HCU5 A0A016ULA1 A0A016U8A7 A0A1S3JU44 A0A016SG46 A0A016TK11 A0A016W4R1 A0A2G8L111 A0A2H1VS24 A0A3P9HFJ3 A0A016VWP2 A0A016V4Q8 A0A016V419 A0A1S3JDY7 A0A1S3JCH2 R7VL02 A0A1S3JGK3 A0A2R2MNC1 R7VDL6 A0A016THQ0 A0A3P9KP93 A0A1S3HDJ9 A0A023GFX8 A0A1S3GZC1 R7U2Q7 A0A0L7LVR9 A0A131XMC8 A0A016VUF9 A0A016S1D2 A0A2G8K4B6 W6N9K0 A0A2W1BZV1 A0A016UYH1 W4XDC3 A0A0N4W414 A0A2R2MPD4 A0A0L7LL03 A0A2G9UVW6 A0A2W1BGL1 A0A0L7LK10 A0A2H2JJX7 A0A2H2ITM4 A0A2A4JIH6 A0A2G9UJ87 K7HM73 K7HM74 A0A016W136 A0A2H2JHZ6 A0A0P4VWD0 A0A2H2J622 A0A2H2JFT4 D7F176 K7HDV1 A0A1I7Y0E1 A0A2W1BNL4 K7HBN5
A0A2W1BHU6 D7F159 D7F164 D7F166 A0A3S2N805 A0A437BHD1 A0A3S2L6Q1 A0A016WTM3 A0A131YWK0 A0A131ZDH5 A0A016W0A5 A0A1W4UBD7 A0A016V9Q8 A0A016VJ99 A0A016VK03 A0A016TNL5 A0A1S3HCU5 A0A016ULA1 A0A016U8A7 A0A1S3JU44 A0A016SG46 A0A016TK11 A0A016W4R1 A0A2G8L111 A0A2H1VS24 A0A3P9HFJ3 A0A016VWP2 A0A016V4Q8 A0A016V419 A0A1S3JDY7 A0A1S3JCH2 R7VL02 A0A1S3JGK3 A0A2R2MNC1 R7VDL6 A0A016THQ0 A0A3P9KP93 A0A1S3HDJ9 A0A023GFX8 A0A1S3GZC1 R7U2Q7 A0A0L7LVR9 A0A131XMC8 A0A016VUF9 A0A016S1D2 A0A2G8K4B6 W6N9K0 A0A2W1BZV1 A0A016UYH1 W4XDC3 A0A0N4W414 A0A2R2MPD4 A0A0L7LL03 A0A2G9UVW6 A0A2W1BGL1 A0A0L7LK10 A0A2H2JJX7 A0A2H2ITM4 A0A2A4JIH6 A0A2G9UJ87 K7HM73 K7HM74 A0A016W136 A0A2H2JHZ6 A0A0P4VWD0 A0A2H2J622 A0A2H2JFT4 D7F176 K7HDV1 A0A1I7Y0E1 A0A2W1BNL4 K7HBN5
Pubmed
EMBL
FJ265550
ADI61818.1
RSAL01000359
RVE42210.1
RSAL01000220
RVE44071.1
+ More
FJ265542 ADI61810.1 FJ265545 ADI61813.1 FJ265543 ADI61811.1 KZ150353 PZC71243.1 FJ265544 ADI61812.1 FJ265549 ADI61817.1 FJ265551 ADI61819.1 RSAL01001818 RVE40675.1 RSAL01000057 RVE49824.1 RSAL01003087 RVE40344.1 JARK01000126 EYC42577.1 GEDV01005625 JAP82932.1 GEDV01000266 JAP88291.1 JARK01001338 EYC33045.1 JARK01001351 EYC23458.1 JARK01001345 EYC27376.1 EYC27377.1 JARK01001424 EYC04321.1 JARK01001372 EYC15676.1 JARK01001389 EYC10853.1 JARK01001565 EYB89683.1 JARK01001676 JARK01001446 JARK01001433 EYB83199.1 EYC01126.1 EYC02928.1 JARK01001337 EYC34625.1 MRZV01000269 PIK53948.1 ODYU01004088 SOQ43598.1 JARK01001340 EYC31168.1 JARK01001354 EYC21713.1 EYC21986.1 AMQN01035462 KB292293 ELU17851.1 AMQN01039158 KB295056 ELU13765.1 JARK01001435 EYC02484.1 GBBM01003763 JAC31655.1 AMQN01002170 AMQN01002171 KB308559 ELT97455.1 JTDY01000008 KOB79474.1 GEFH01000854 JAP67727.1 JARK01001341 EYC30403.1 JARK01001655 EYB84321.1 MRZV01000899 PIK42832.1 CAVP010052198 CDL93746.1 KZ149907 PZC78256.1 JARK01001358 EYC20230.1 AAGJ04050748 UZAF01016227 VDO23583.1 JTDY01000715 KOB76117.1 KZ345286 PIO74399.1 KZ150150 PZC72845.1 JTDY01000781 KOB75903.1 NWSH01001447 PCG71212.1 KZ346514 PIO69792.1 EYC33365.1 GDKW01001596 JAI54999.1 FJ265561 ADI61829.1 KZ150049 PZC74456.1
FJ265542 ADI61810.1 FJ265545 ADI61813.1 FJ265543 ADI61811.1 KZ150353 PZC71243.1 FJ265544 ADI61812.1 FJ265549 ADI61817.1 FJ265551 ADI61819.1 RSAL01001818 RVE40675.1 RSAL01000057 RVE49824.1 RSAL01003087 RVE40344.1 JARK01000126 EYC42577.1 GEDV01005625 JAP82932.1 GEDV01000266 JAP88291.1 JARK01001338 EYC33045.1 JARK01001351 EYC23458.1 JARK01001345 EYC27376.1 EYC27377.1 JARK01001424 EYC04321.1 JARK01001372 EYC15676.1 JARK01001389 EYC10853.1 JARK01001565 EYB89683.1 JARK01001676 JARK01001446 JARK01001433 EYB83199.1 EYC01126.1 EYC02928.1 JARK01001337 EYC34625.1 MRZV01000269 PIK53948.1 ODYU01004088 SOQ43598.1 JARK01001340 EYC31168.1 JARK01001354 EYC21713.1 EYC21986.1 AMQN01035462 KB292293 ELU17851.1 AMQN01039158 KB295056 ELU13765.1 JARK01001435 EYC02484.1 GBBM01003763 JAC31655.1 AMQN01002170 AMQN01002171 KB308559 ELT97455.1 JTDY01000008 KOB79474.1 GEFH01000854 JAP67727.1 JARK01001341 EYC30403.1 JARK01001655 EYB84321.1 MRZV01000899 PIK42832.1 CAVP010052198 CDL93746.1 KZ149907 PZC78256.1 JARK01001358 EYC20230.1 AAGJ04050748 UZAF01016227 VDO23583.1 JTDY01000715 KOB76117.1 KZ345286 PIO74399.1 KZ150150 PZC72845.1 JTDY01000781 KOB75903.1 NWSH01001447 PCG71212.1 KZ346514 PIO69792.1 EYC33365.1 GDKW01001596 JAI54999.1 FJ265561 ADI61829.1 KZ150049 PZC74456.1
Proteomes
Pfam
Interpro
IPR000477
RT_dom
+ More
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR027124 Swc5/CFDP2
IPR020846 MFS_dom
IPR002293 AA/rel_permease1
IPR032695 Integrin_dom_sf
IPR036465 vWFA_dom_sf
IPR015812 Integrin_bsu
IPR002035 VWF_A
IPR002369 Integrin_bsu_VWA
IPR038441 THAP_Znf_sf
IPR006612 THAP_Znf
IPR016186 C-type_lectin-like/link_sf
IPR016187 CTDL_fold
IPR007110 Ig-like_dom
IPR036179 Ig-like_dom_sf
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR027124 Swc5/CFDP2
IPR020846 MFS_dom
IPR002293 AA/rel_permease1
IPR032695 Integrin_dom_sf
IPR036465 vWFA_dom_sf
IPR015812 Integrin_bsu
IPR002035 VWF_A
IPR002369 Integrin_bsu_VWA
IPR038441 THAP_Znf_sf
IPR006612 THAP_Znf
IPR016186 C-type_lectin-like/link_sf
IPR016187 CTDL_fold
IPR007110 Ig-like_dom
IPR036179 Ig-like_dom_sf
SUPFAM
Gene 3D
CDD
ProteinModelPortal
D7F165
A0A3S2KZJ4
A0A437B0R4
D7F157
D7F160
D7F158
+ More
A0A2W1BHU6 D7F159 D7F164 D7F166 A0A3S2N805 A0A437BHD1 A0A3S2L6Q1 A0A016WTM3 A0A131YWK0 A0A131ZDH5 A0A016W0A5 A0A1W4UBD7 A0A016V9Q8 A0A016VJ99 A0A016VK03 A0A016TNL5 A0A1S3HCU5 A0A016ULA1 A0A016U8A7 A0A1S3JU44 A0A016SG46 A0A016TK11 A0A016W4R1 A0A2G8L111 A0A2H1VS24 A0A3P9HFJ3 A0A016VWP2 A0A016V4Q8 A0A016V419 A0A1S3JDY7 A0A1S3JCH2 R7VL02 A0A1S3JGK3 A0A2R2MNC1 R7VDL6 A0A016THQ0 A0A3P9KP93 A0A1S3HDJ9 A0A023GFX8 A0A1S3GZC1 R7U2Q7 A0A0L7LVR9 A0A131XMC8 A0A016VUF9 A0A016S1D2 A0A2G8K4B6 W6N9K0 A0A2W1BZV1 A0A016UYH1 W4XDC3 A0A0N4W414 A0A2R2MPD4 A0A0L7LL03 A0A2G9UVW6 A0A2W1BGL1 A0A0L7LK10 A0A2H2JJX7 A0A2H2ITM4 A0A2A4JIH6 A0A2G9UJ87 K7HM73 K7HM74 A0A016W136 A0A2H2JHZ6 A0A0P4VWD0 A0A2H2J622 A0A2H2JFT4 D7F176 K7HDV1 A0A1I7Y0E1 A0A2W1BNL4 K7HBN5
A0A2W1BHU6 D7F159 D7F164 D7F166 A0A3S2N805 A0A437BHD1 A0A3S2L6Q1 A0A016WTM3 A0A131YWK0 A0A131ZDH5 A0A016W0A5 A0A1W4UBD7 A0A016V9Q8 A0A016VJ99 A0A016VK03 A0A016TNL5 A0A1S3HCU5 A0A016ULA1 A0A016U8A7 A0A1S3JU44 A0A016SG46 A0A016TK11 A0A016W4R1 A0A2G8L111 A0A2H1VS24 A0A3P9HFJ3 A0A016VWP2 A0A016V4Q8 A0A016V419 A0A1S3JDY7 A0A1S3JCH2 R7VL02 A0A1S3JGK3 A0A2R2MNC1 R7VDL6 A0A016THQ0 A0A3P9KP93 A0A1S3HDJ9 A0A023GFX8 A0A1S3GZC1 R7U2Q7 A0A0L7LVR9 A0A131XMC8 A0A016VUF9 A0A016S1D2 A0A2G8K4B6 W6N9K0 A0A2W1BZV1 A0A016UYH1 W4XDC3 A0A0N4W414 A0A2R2MPD4 A0A0L7LL03 A0A2G9UVW6 A0A2W1BGL1 A0A0L7LK10 A0A2H2JJX7 A0A2H2ITM4 A0A2A4JIH6 A0A2G9UJ87 K7HM73 K7HM74 A0A016W136 A0A2H2JHZ6 A0A0P4VWD0 A0A2H2J622 A0A2H2JFT4 D7F176 K7HDV1 A0A1I7Y0E1 A0A2W1BNL4 K7HBN5
Ontologies
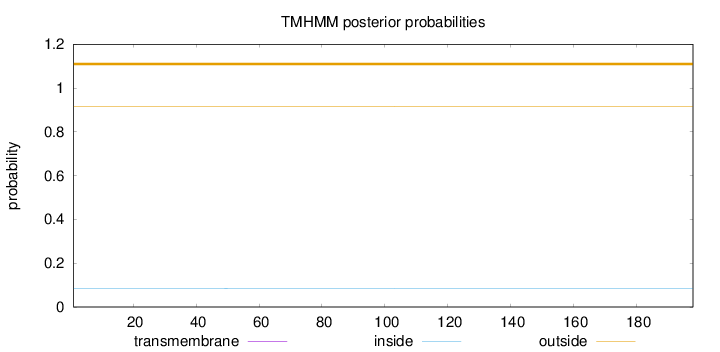
Topology
Subcellular location
Membrane
Length:
198
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00096
Exp number, first 60 AAs:
0.00096
Total prob of N-in:
0.08390
outside
1 - 198
Population Genetic Test Statistics
Pi
13.318289
Theta
19.523609
Tajima's D
-0.987787
CLR
0
CSRT
0.139643017849108
Interpretation
Uncertain
