Gene
KWMTBOMO16622
Annotation
non-LTR_retrotransposon_R1Bmks_ORF2_protein_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 0.992
Sequence
CDS
ATGAGTGCCAAAGGCCGGAGGACCGGCCTGCATATCCCGGTTGCTTCTGGACTGTCGTCCTCGGCGCCTGTCACCCCGGTTGACCCAGTGGCTGGTGTCCCTAGCTTCCCCATTCCTGTTGCGTCGGGGGCTCCGACTGGACCCCTTGATGTAGCAGCGCAGGGGAGGCTGGAGCTGCTCGAGCGGGCAAACCGAGCGGTGCGCGGGATCATGTCGGTGGCGACGGCGGCATCTAAGCTCAATAAGTCGGAGGTCAATCTAATCTCTGAACTCGGTCGCGATATCCTGGCCGTAGTTGGGGCCCTCGGGATCCAGCTCTCTGATAAGGAGCTGGTGGTCGAAAGGCTCCGCTCGGCAGAGGCCTCTGCACGGCGTGACTCCGTGGCGCCTTCGATGGCTGCTGGTGCGGCGGCCGGGTCTGGGACTGGGCCTGCGACCTTCGCCACTGTCCTCAGGACGGGCCCTGGTGGGGTCCCGAGGTCCATTGGGGCCTCTCAGGGGCCTTCCTTGGCATTCTACCCGTCCGAGGGCAATGCGGAACTGAAGACAGCTGAGGACACAAAGAAAGAGGTAAAAAAGGCCATTGACCCTAAGTCAATGGCTATTGGAATACAGAGCGTAAGGAAGGTTGGGAATGCGGGGGTAGTGGTGCAGACCACGTCCCCCGCGGCCGCAGTCAAGCTTAGAAATGCTGCCCCGCCATCACTGAGAGTCACCGAACCCAGGCGCCGACAGCCCTTAGTAGCTGTCAATGGCGTGGAGGGCGACCCCTCCTTCGAGGAGGTAATCGAGTGCCTGGCTAGCCAGAACCTCGACCCGGAGGAGTGGCCCCTCACTAGGGTACGAGCGGAGCTCACGGGGGCGTTCAAAAAGGGGAGGCGACAATCCAACAACACTACTGTGGTGTTTAACGCCTCCCCTCGCATCAGGGACGCCCTCGTGAAGATTGGCAGAGTGTATGTGGGGTGGGTCGCCTGTGAGGTCACGGACTTTGTCCGGGTGACCTGCTGCAATAAGTGTCAGCAATATGGTCACCCGGAGAAATTTTGCCGGGCCAAGGAGGCCACCTGTGGCCGATGTGGAGAGGATGGCCACCGAATGGAGGCCTGTAAGGCAGCCTCCGCGTCGAGTATGGATATTAGGCCCCGACTTCGTATTGGCCAAATCAATCTGGGTGGTGCAGAGGATGCGACGAGGGAGCTACCCTCCATTGCACGGGATCTCGGCCTGGATATTGTTCTTGTACAGGAACAATATTCCATGGTCGGGTTCCTAGCCCAATGTGGAGCACACCCCAAGGCGGGTGTGTATATCCGCAATAGGGTGCTCCCCTGCGCGGTTCTGCACCACCTTAGCAGCACACATATAACGGTAGTGCACATTGGGGGGTGGGACTTATATATGGTGTCTGCGTACTTCCAGTATAGTGACCCTATTGACCCATACCTGCACCGGCTCGGGAATATTCTTGACCGGCTGCGGGGGGCTCGGGTCGTTATCTGCGCAGACACTAATGCCCACTCGCCATTGTGGCACTCGCTGCCCAGGCACTACGTCGGTCGGGGTCAGGAAGTGGCTGACCGCCGCGCCAAGATGGAGGATTTCATTGGGGCGAGGCGGTTGGTCGTCCATAACGCGGATGGCCACCTGCCGACCTTCAGTACGGCGAACGGAGAATCTTATGTCGATGTCACGCTGTCTACGCGGGGAGTACGCGTGTCTGAATGGCGTGTAACTAATGAATCATCGAGCGATCACCGGCTCATTGTGTTTGGGGTGGGGGGCGGTACAACAGGGGAGCGGGACGAGGACGAGGAGGCGCGGAGCGATTTGAGGCCGGGCGAGCCGTGCGCACTGCGTCGGTACCGGGACCGTGGGGTGGATTGGGACCTCTTCAGATCGCGTATCCACGAGCGAATGGGGAGTCTGGACCTCGAGGAACCTGTGGCTGCCCTTTGCGAAAAATTTACCGGGGTTATAACTCGCACAGCTGAAGAATGCTTAGGATCACTGAAAGCAGATAGAACTGACAGGGGTTATGAGTGGTGGACCCCAGTACTCGATAAGCTTAGGGTAGCCCAGGGTAGGGCCAGGCGTCGATGGCAAAAGGCCCGCCGAACGGGGGGTGAGGAAGAGGAGCAGTCTGGGAGAGTCTTCCGCGACCGCAGGCGCGAGTATCGAAGAGCGATGCATGACGCTGAGACCGCCTTTTACCGGGAGATCGCTGAAGGGGGAAACCGTGACCCGTGGGGACTAGCGTATCGGACGGCGAGTGGTAGGCGACGCGCACCAACTAATGTGGTTAACGGGGTGGAGTATGCGGGGCGGTGCTCGGATGATGTGAGTGGGGCCATGCGCACCTTGATGTGGGCGCTGTGTCCGGATGACTATATGTCTCGGGACACTCCGTACCATGCGCGGGTGCGTATCATGGCCGCGCTCCCCCCATCCGGGCGGGACGCAGACCCGCTGAGCAAAGACTCCCTTCGTGCCATAATTGGCTCACTGAAGAATACCGCACCGGGCATCGACGGTTTAACGGCGCGCATTATCAAAAAGGCACTTCCGGCTGCTGAGGCCGAGTTCGTGGCCGTATACGCACGGTGCGTTGTGGAGGGGACCTTCCCGCCGGTGTGGAAGGATGGCCGCCTACTTGTTCTGCCAAAGGGGAATGGCAGGCCCTTAACGGACCCTAAGGCGTATCGCCCGGTCACCTTGCTGCCGGTCCTGGGAAAGATCTTGGAGAAGGTATTATTGCAGTGTGCTCCTGGCCTCACCCATAGTATTAGTCCGCGCCAGCACGGGTTCTCTCCTGGACGCTCAACGGTGACGGCGCTGCGAACTCTGCTGGACGTGTCGCGCGCCTCGGAGCAGAGGTACGTAATGGCCATATTCTTGGACATCAGTGGAGCTTTCGATAACGCGTGGTGGCCCATGATAATGGTGAAGGCCAAGCGGAACTGTCCGCCCAACATCTATCGGATGCTGACGGACTATTTCCGCGGACGCCGTATTGCCGTTGTCGCGGGGGAATGTGCGGAATGGAAGGTGTCCACGATGGGCTGTCCGCAGGGCTCAGTGCTCGGGCCGACGCTCTGGAACGTTCTGATGGATGACCTGCTCGCCTTGCCGCAGGGGATAGAGGGAACAGAGATGGTCGCCTATGCCGATGACGTGACGGTACTGGTTAGGGGTGACTCACGGGCGCAGCTTGAGAGGAGAGCGCACGCCGTGCTAGGACTCGCAGAGGGGTGGGCGAGCAGGAATAAGCTCGATTTTGCCCCGGCGAAGTCCCGATGCATAATGCTGAGGGGAAAGTTTCAGCGTCCCCCTATAGTCCGGTACGGCAGTCATGTCATTCGGTTCGAGAACCAGGTGACGGTGTTGGGCGTCGTCTTCGACGATTGCCTCTCTTTCGCGCCGCATGCGGCGGCCATTGGCGAGAGGGCGAGCAGGTGCTTCGGCAAGATGTCTAGAGTTTCGGCTTCGGCTTGGGGGCTGCGATATAGGGCTTTGCGTGTCTTGTACATGGGCACTTATGTTACAACCCTTACCTATGCGGCGGCCGTATGGTATTTGCGGGCTGCTGTGCACGTCGTGCGCAGCGTGCTGCTTAGGACGCAGCGCCCGTCGTTGACGCTGCTAACGAAGGCCTACCGTTCGTGCAGCACGGCTGCTTTGCCGGTGTTGGCGGGCGTCCTGCCGGCGGACCTGGAGGTGACTCGTGCTGGACGGAGGATGCGGGAGTGCGAAGGATTGGCGCGGGAGTTGGCGGCGGAGAGACGACGACGGATCGACGGCGATGTCTTGGTAGTTTGGCAGAACAGGTGGGTGTCTGAGGGTAAGGGGAGGGAACTGTACAAGTTCTTTCCCGATGTTGCGGACAGGAAGAAGGCAACGTGGATGGAGCCGGACTATCAGACCTCGCAGATCCTCACGGGTCATGGGATCTTTAATAAGCGGTTGGCGGATATGCGACTGAGGGAGGGGCATGCTTGCGACTGCGGGGCGGTTGAGGAGGATAGGGACCATGTCCTGTGGGAGTGTCCTCTCTATGACGAAATCCGGGGCAGGATGCTCGATGGAATCTCGCGGTCTGAGGTGGGCCCAGTTTACCACGCGGACCTGGTCAGGGACGAGAAAAATTTTCGGCTCTTGCGCGAGTTCGCGCATACATGGCACACGGCGCGCACTGCGCGCGAATCACGAGGACTGCCTGGCGACGAAGAATGTGGGTGA
Protein
MSAKGRRTGLHIPVASGLSSSAPVTPVDPVAGVPSFPIPVASGAPTGPLDVAAQGRLELLERANRAVRGIMSVATAASKLNKSEVNLISELGRDILAVVGALGIQLSDKELVVERLRSAEASARRDSVAPSMAAGAAAGSGTGPATFATVLRTGPGGVPRSIGASQGPSLAFYPSEGNAELKTAEDTKKEVKKAIDPKSMAIGIQSVRKVGNAGVVVQTTSPAAAVKLRNAAPPSLRVTEPRRRQPLVAVNGVEGDPSFEEVIECLASQNLDPEEWPLTRVRAELTGAFKKGRRQSNNTTVVFNASPRIRDALVKIGRVYVGWVACEVTDFVRVTCCNKCQQYGHPEKFCRAKEATCGRCGEDGHRMEACKAASASSMDIRPRLRIGQINLGGAEDATRELPSIARDLGLDIVLVQEQYSMVGFLAQCGAHPKAGVYIRNRVLPCAVLHHLSSTHITVVHIGGWDLYMVSAYFQYSDPIDPYLHRLGNILDRLRGARVVICADTNAHSPLWHSLPRHYVGRGQEVADRRAKMEDFIGARRLVVHNADGHLPTFSTANGESYVDVTLSTRGVRVSEWRVTNESSSDHRLIVFGVGGGTTGERDEDEEARSDLRPGEPCALRRYRDRGVDWDLFRSRIHERMGSLDLEEPVAALCEKFTGVITRTAEECLGSLKADRTDRGYEWWTPVLDKLRVAQGRARRRWQKARRTGGEEEEQSGRVFRDRRREYRRAMHDAETAFYREIAEGGNRDPWGLAYRTASGRRRAPTNVVNGVEYAGRCSDDVSGAMRTLMWALCPDDYMSRDTPYHARVRIMAALPPSGRDADPLSKDSLRAIIGSLKNTAPGIDGLTARIIKKALPAAEAEFVAVYARCVVEGTFPPVWKDGRLLVLPKGNGRPLTDPKAYRPVTLLPVLGKILEKVLLQCAPGLTHSISPRQHGFSPGRSTVTALRTLLDVSRASEQRYVMAIFLDISGAFDNAWWPMIMVKAKRNCPPNIYRMLTDYFRGRRIAVVAGECAEWKVSTMGCPQGSVLGPTLWNVLMDDLLALPQGIEGTEMVAYADDVTVLVRGDSRAQLERRAHAVLGLAEGWASRNKLDFAPAKSRCIMLRGKFQRPPIVRYGSHVIRFENQVTVLGVVFDDCLSFAPHAAAIGERASRCFGKMSRVSASAWGLRYRALRVLYMGTYVTTLTYAAAVWYLRAAVHVVRSVLLRTQRPSLTLLTKAYRSCSTAALPVLAGVLPADLEVTRAGRRMRECEGLARELAAERRRRIDGDVLVVWQNRWVSEGKGRELYKFFPDVADRKKATWMEPDYQTSQILTGHGIFNKRLADMRLREGHACDCGAVEEDRDHVLWECPLYDEIRGRMLDGISRSEVGPVYHADLVRDEKNFRLLREFAHTWHTARTARESRGLPGDEECG
Summary
Uniprot
Q5NTZ0
Q7M4J4
V9H050
A0A1B6DRM7
A0A1B6E1N0
A0A023EYP7
+ More
K7JMI9 A0A0J7KTH2 J9M8T8 A0A224XAL8 J9MA04 K7JAI1 J9K5I1 J9M4Z2 J9L798 J9KMZ9 J9LMF6 J9LPJ1 X1WL18 X1XHQ0 J9KPX6 A0A224X653 J9LNF5 J9L0V3 J9LPX1 J9LVC2 A0A087UHT1 J9LPX0 J9K2L0 Q9BLI5 J9KA38 J9KRH3 J9KVB7 J9KPC7 J9KPE5 J9KJG8 A0A142LX43 X1WVB5 Q868T0 A0A0J7KB28 J9KT61 A0A2M4CS98 A0A0K8VPH1 A0A2S2NQQ6
K7JMI9 A0A0J7KTH2 J9M8T8 A0A224XAL8 J9MA04 K7JAI1 J9K5I1 J9M4Z2 J9L798 J9KMZ9 J9LMF6 J9LPJ1 X1WL18 X1XHQ0 J9KPX6 A0A224X653 J9LNF5 J9L0V3 J9LPX1 J9LVC2 A0A087UHT1 J9LPX0 J9K2L0 Q9BLI5 J9KA38 J9KRH3 J9KVB7 J9KPC7 J9KPE5 J9KJG8 A0A142LX43 X1WVB5 Q868T0 A0A0J7KB28 J9KT61 A0A2M4CS98 A0A0K8VPH1 A0A2S2NQQ6
EMBL
AB182560
BAD82946.1
M19755
AAC13649.1
GEDC01008950
JAS28348.1
+ More
GEDC01005462 JAS31836.1 GBBI01004973 JAC13739.1 AAZX01008070 LBMM01003366 KMQ93606.1 ABLF02007471 ABLF02007472 GFTR01008452 JAW07974.1 ABLF02008888 ABLF02016960 ABLF02032572 AAZX01006426 ABLF02027069 ABLF02027071 ABLF02010919 ABLF02010923 ABLF02010931 ABLF02007045 ABLF02039177 ABLF02016436 ABLF02012872 ABLF02012873 ABLF02012874 ABLF02012875 ABLF02042485 ABLF02010037 ABLF02010039 ABLF02066755 ABLF02019936 ABLF02057951 GFTR01008469 JAW07957.1 ABLF02006133 ABLF02012458 ABLF02012459 ABLF02012124 ABLF02008516 ABLF02041501 KK119855 KFM76920.1 ABLF02020044 ABLF02004512 ABLF02004513 ABLF02046624 ABLF02049075 ABLF02059998 ABLF02065990 AB046668 BAB21511.1 ABLF02004773 ABLF02020082 ABLF02020086 ABLF02020092 ABLF02022279 ABLF02034348 ABLF02008515 ABLF02013103 ABLF02013105 ABLF02013107 ABLF02061460 ABLF02041886 KU543682 AMS38369.1 ABLF02007851 ABLF02016357 ABLF02064424 AB090812 BAC57900.1 LBMM01010435 KMQ87461.1 ABLF02009073 ABLF02030702 ABLF02042963 GGFL01004044 MBW68222.1 GDHF01029467 GDHF01011533 JAI22847.1 JAI40781.1 GGMR01006838 MBY19457.1
GEDC01005462 JAS31836.1 GBBI01004973 JAC13739.1 AAZX01008070 LBMM01003366 KMQ93606.1 ABLF02007471 ABLF02007472 GFTR01008452 JAW07974.1 ABLF02008888 ABLF02016960 ABLF02032572 AAZX01006426 ABLF02027069 ABLF02027071 ABLF02010919 ABLF02010923 ABLF02010931 ABLF02007045 ABLF02039177 ABLF02016436 ABLF02012872 ABLF02012873 ABLF02012874 ABLF02012875 ABLF02042485 ABLF02010037 ABLF02010039 ABLF02066755 ABLF02019936 ABLF02057951 GFTR01008469 JAW07957.1 ABLF02006133 ABLF02012458 ABLF02012459 ABLF02012124 ABLF02008516 ABLF02041501 KK119855 KFM76920.1 ABLF02020044 ABLF02004512 ABLF02004513 ABLF02046624 ABLF02049075 ABLF02059998 ABLF02065990 AB046668 BAB21511.1 ABLF02004773 ABLF02020082 ABLF02020086 ABLF02020092 ABLF02022279 ABLF02034348 ABLF02008515 ABLF02013103 ABLF02013105 ABLF02013107 ABLF02061460 ABLF02041886 KU543682 AMS38369.1 ABLF02007851 ABLF02016357 ABLF02064424 AB090812 BAC57900.1 LBMM01010435 KMQ87461.1 ABLF02009073 ABLF02030702 ABLF02042963 GGFL01004044 MBW68222.1 GDHF01029467 GDHF01011533 JAI22847.1 JAI40781.1 GGMR01006838 MBY19457.1
Proteomes
Pfam
Interpro
IPR000477
RT_dom
+ More
IPR005135 Endo/exonuclease/phosphatase
IPR036691 Endo/exonu/phosph_ase_sf
IPR001878 Znf_CCHC
IPR036875 Znf_CCHC_sf
IPR003959 ATPase_AAA_core
IPR027417 P-loop_NTPase
IPR003960 ATPase_AAA_CS
IPR031996 NVL2_nucleolin-bd
IPR041569 AAA_lid_3
IPR003593 AAA+_ATPase
IPR038100 NLV2_N_sf
IPR007889 HTH_Psq
IPR005312 DUF1759
IPR008737 Peptidase_asp_put
IPR036397 RNaseH_sf
IPR002156 RNaseH_domain
IPR012337 RNaseH-like_sf
IPR005135 Endo/exonuclease/phosphatase
IPR036691 Endo/exonu/phosph_ase_sf
IPR001878 Znf_CCHC
IPR036875 Znf_CCHC_sf
IPR003959 ATPase_AAA_core
IPR027417 P-loop_NTPase
IPR003960 ATPase_AAA_CS
IPR031996 NVL2_nucleolin-bd
IPR041569 AAA_lid_3
IPR003593 AAA+_ATPase
IPR038100 NLV2_N_sf
IPR007889 HTH_Psq
IPR005312 DUF1759
IPR008737 Peptidase_asp_put
IPR036397 RNaseH_sf
IPR002156 RNaseH_domain
IPR012337 RNaseH-like_sf
Gene 3D
ProteinModelPortal
Q5NTZ0
Q7M4J4
V9H050
A0A1B6DRM7
A0A1B6E1N0
A0A023EYP7
+ More
K7JMI9 A0A0J7KTH2 J9M8T8 A0A224XAL8 J9MA04 K7JAI1 J9K5I1 J9M4Z2 J9L798 J9KMZ9 J9LMF6 J9LPJ1 X1WL18 X1XHQ0 J9KPX6 A0A224X653 J9LNF5 J9L0V3 J9LPX1 J9LVC2 A0A087UHT1 J9LPX0 J9K2L0 Q9BLI5 J9KA38 J9KRH3 J9KVB7 J9KPC7 J9KPE5 J9KJG8 A0A142LX43 X1WVB5 Q868T0 A0A0J7KB28 J9KT61 A0A2M4CS98 A0A0K8VPH1 A0A2S2NQQ6
K7JMI9 A0A0J7KTH2 J9M8T8 A0A224XAL8 J9MA04 K7JAI1 J9K5I1 J9M4Z2 J9L798 J9KMZ9 J9LMF6 J9LPJ1 X1WL18 X1XHQ0 J9KPX6 A0A224X653 J9LNF5 J9L0V3 J9LPX1 J9LVC2 A0A087UHT1 J9LPX0 J9K2L0 Q9BLI5 J9KA38 J9KRH3 J9KVB7 J9KPC7 J9KPE5 J9KJG8 A0A142LX43 X1WVB5 Q868T0 A0A0J7KB28 J9KT61 A0A2M4CS98 A0A0K8VPH1 A0A2S2NQQ6
PDB
2EI9
E-value=2.28684e-127,
Score=1172
Ontologies
GO
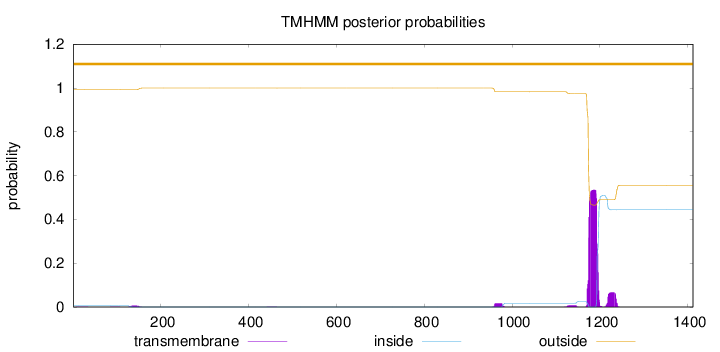
Topology
Length:
1412
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
13.67875
Exp number, first 60 AAs:
0.01321
Total prob of N-in:
0.00705
outside
1 - 1412
Population Genetic Test Statistics
Pi
0
Theta
0
Tajima's D
0
CLR
0
CSRT
0
Interpretation
Uncertain
