Gene
KWMTBOMO16602
Annotation
PREDICTED:_small_G_protein_signaling_modulator_3_homolog_[Amyelois_transitella]
Location in the cell
Cytoplasmic Reliability : 2.653
Sequence
CDS
ATGGATCTTGCCAAATCGTTATTCAGTTCAAGAGATCATGTTGGGTATGTCGGTCGTGAAGACCACGAACGTAAGATTAGTGCAGCGCTGAACGATGATGACCCTGAAGACGTGATCGACGCGATTAGAGGACTAAACATAGCTGATGAACTTCTTCCTGCGCCAGGAGGGCCATTTTCTGCTCTAACGCCTAGTATGGTACCTCAAGACATAATGGCTAAACTAGCACAACCAGAATCCGAAGGACACGGGGGTGGACTACCAGACTACAGTTTCGACGAGTTTGGCTTTTGTATTGAAGAAGAAGATGGTCCGGAACAAAGCTCAAACAAATTATTAGCTAATGCATTTGTTGAAGATGATCAACATAGACTGCAATGGGAGTTTTACAGCAAAGAAATTAACTTTACCGAAGGCAGTGGTAAGCTTGAAAAATCAGATAAACTACAGAAAATGGTCAAAGATGGTGTACCACATTCACTTAGGCCTCAAGTCTGGATGCATTTATGTGGAGCTAACAAGAAACGAGCATCAACTGAAATAACATATCATGAAATTGTAAGGGCATCAAGTGATGATGGTTTGGTAACTAGCAAACAAATAGAAAAAGATTTAGTCCAAATTCTGCCTAATAATGTTTGTTTCTCACATCCAACTAGCACTGGTGTACCTCGGCTTAGACGCATCCTCAGAGCCTTAGCATGGCTCTATCCAGACATTGGATACTGTCAAGGGACAGGAATGATTGCAGCATCACTATTACTATTAATGGAAGAAGAAGAAGCATTCTGGATAATGTGTACAACAGTGGAAGATCTTTTACCAGCTTCTTATTATTCATCAACACTGATTGGAATCCAAGCAGATCAAAAAGTTTTAAGGAGTCTTATATCAAACTATTTACCAGGCATCGACCAGGTCCTTAATGCTCACGACATTGAACTATCACTTATCACAATGCATTGGTTTCTAACTTTGTATGCTAATGTAGTCCATATGAAAATTTTATTAAGAATATGGGATCTATTTTTCTTTGATGGCTCTATTGTCTTGTTTAAAGTAACATTAGCGATGTTGAAAATAAAAGAAAATAGATTCACAGAAATCTCCAACTCCGCCCAAATCTTCAATGCTCTCTCTGATATACCTGGGGAAATTGATGATGTTGAACTTCTAATAAAAACCATTGATGAGGTATGTGGTGATACTCTAAGTCCTGCATTAATAGAAACACACAGAAGAAGACACTTGGCATACTTAATGGCAGATCAAGGTGCTCTTGTTGGAAATCCAAATGTTGTACCAAACTTGCCTAAGCAGCAATTGCAGAGGCGTCAAATAAAAAAGAGCAAGTCTGTAATACAAACTATTTTGTTTGGGGAAGATGGTAGTAATGAAGATGCTATTTCTAAGAATGTGAAGCAAACTGAAATACTAGTGGACTTGAGAGAAGCAATACTACAGGTTGCAAGGCACTTCCTAGCTTTAGAACCACAACTGACTTCAGAAGTTCAAATGACAGCTGATTACACCATGCAGAGTCATGCTGCTGATAGGGAAAGATACGTTAATGTTTCCAGAAGCAAGAGGCGAAGGGCAAAAGCTCTATTAGACTTTGAGAGAAGAGATGGTGATGAATTAGGATTCAGAAAAAATGATGTTATTGAAGTTATAAGTTCAAGAGATGAACACTGTTGGATTGGAGAACTAAACGGGTTGAGGGGCTGGTTTCCTGCTAGATTTGTCAAATTGCTCGATGAAAAGGGTGTAAAATATAGCAAGGCTGGTGATGATTCTGTCACGGAAAAAGCTGCACACTTAGTGAGAGGTGTATTAGCTCCAGCATTTAAAAGAGTTCTCTTGCATGGTATAAAGAGGCCAAGCTTTCTAGATGGACCATGTCATCCGTGGCTATTCATTGAAGAAGCCTCTTCTCGCGAAGTTGAAAAAGATTTTAATTCAGTTTACAGCCGCTTGGTTCTCTGTAAGACATATAGATTAGATGAAGATGGGAAGGTGTTGACTCCTGAAGAGTTATTATACAGATGTGTCCAAGCAATAAATTTATCACATGACAATGCTCATGCTCAGATGGATGTGAAATTGCGCACATTAATATGTATGGGCTTAAATGAAGAAGCATTACATCTGTGGCTTGAAGTATTGTGTTCTTGTGTTGATGTCGTTCAAAAATGGTATCATCCATGGTCATTTATAAATTCGCCTGGATGGGTACAGATCAAATGTGAATTACGTATACTTTCTCAGTTTGGTTTCAATTTGAACCCCGATTGGGAGCTTCCATTGAAAAAAGATACACAAAATCAGCCACTGAAGGAAGGTGTAAGAGATATGCTTGTTAAACACCATTTATTCTCATGGGACTTGTGA
Protein
MDLAKSLFSSRDHVGYVGREDHERKISAALNDDDPEDVIDAIRGLNIADELLPAPGGPFSALTPSMVPQDIMAKLAQPESEGHGGGLPDYSFDEFGFCIEEEDGPEQSSNKLLANAFVEDDQHRLQWEFYSKEINFTEGSGKLEKSDKLQKMVKDGVPHSLRPQVWMHLCGANKKRASTEITYHEIVRASSDDGLVTSKQIEKDLVQILPNNVCFSHPTSTGVPRLRRILRALAWLYPDIGYCQGTGMIAASLLLLMEEEEAFWIMCTTVEDLLPASYYSSTLIGIQADQKVLRSLISNYLPGIDQVLNAHDIELSLITMHWFLTLYANVVHMKILLRIWDLFFFDGSIVLFKVTLAMLKIKENRFTEISNSAQIFNALSDIPGEIDDVELLIKTIDEVCGDTLSPALIETHRRRHLAYLMADQGALVGNPNVVPNLPKQQLQRRQIKKSKSVIQTILFGEDGSNEDAISKNVKQTEILVDLREAILQVARHFLALEPQLTSEVQMTADYTMQSHAADRERYVNVSRSKRRRAKALLDFERRDGDELGFRKNDVIEVISSRDEHCWIGELNGLRGWFPARFVKLLDEKGVKYSKAGDDSVTEKAAHLVRGVLAPAFKRVLLHGIKRPSFLDGPCHPWLFIEEASSREVEKDFNSVYSRLVLCKTYRLDEDGKVLTPEELLYRCVQAINLSHDNAHAQMDVKLRTLICMGLNEEALHLWLEVLCSCVDVVQKWYHPWSFINSPGWVQIKCELRILSQFGFNLNPDWELPLKKDTQNQPLKEGVRDMLVKHHLFSWDL
Summary
Uniprot
A0A2H1VYA1
A0A2A4K9C8
A0A1E1WS27
A0A212EXS9
A0A0L7K4G6
A0A194RKB8
+ More
A0A194PJH2 A0A151I9N6 A0A195DKL1 F4WQG0 A0A1Y1K076 A0A195F775 A0A2A3EM99 A0A158NNI4 A0A087ZZH9 A0A0N0U6A0 A0A151WWS4 A0A1B0GJX0 E2BSQ7 D6W7G3 A0A0L7QNB1 A0A154P6V1 K7IN72 A0A1L8DVY6 A0A232EV58 Q17FM5 A0A0L0C869 A0A182G3G4 A0A182KH75 A0A067RC13 T1PHB3 A0A2J7PLR1 A0A1I8M973 A0A182LQW0 Q7PYV2 A0A182UP80 A0A224XIU0 E2AUU8 A0A1W4X420 A0A182FRA2 W5JDV0 A0A069DX50 A0A195B3I6 A0A182TVG4 A0A182NZ96 A0A182W014 A0A0C9R602 A0A182MJT8 A0A182R0S2 A0A1I8Q8D8 A0A1A9ZR04 B0WTA5 A0A1A9UF38 A0A182J5J6 A0A0J7KUH0 A0A1A9WPE1 A0A1B0FJF4 A0A1A9YRN8 N6U6M8 A0A182PND3 A0A1B0B4A0 A0A182HZU2 A0A182XGI4 B4KAG5 B3MXA1 B3P0R8 A0A1W4VN90 B4QZ41 B4PS79 B4HE08 Q9VFB6 B4GMM2 Q294T8 E0V8Z8 B4JGB2 A0A182Y458 B4LY26 A0A1W4WT53 A0A3B0K6J4 W8BMW7 A0A084W3H3 A0A0K8UJT7 A0A034VDE6 A0A0R3NJJ4 A0A0K8WI48 B4NBB4 A0A0A1X830 A0A0M4EW80 A0A182RNW4 A0A3R7PS56 A0A1B6KJ19 A0A2R5LBM2 A0A210Q0E7 A0A0B7ACK4 T1J9C7 K1Q6P1 A0A0B7AA75 A0A1S3HG43 L7M9L0 B7P0S7
A0A194PJH2 A0A151I9N6 A0A195DKL1 F4WQG0 A0A1Y1K076 A0A195F775 A0A2A3EM99 A0A158NNI4 A0A087ZZH9 A0A0N0U6A0 A0A151WWS4 A0A1B0GJX0 E2BSQ7 D6W7G3 A0A0L7QNB1 A0A154P6V1 K7IN72 A0A1L8DVY6 A0A232EV58 Q17FM5 A0A0L0C869 A0A182G3G4 A0A182KH75 A0A067RC13 T1PHB3 A0A2J7PLR1 A0A1I8M973 A0A182LQW0 Q7PYV2 A0A182UP80 A0A224XIU0 E2AUU8 A0A1W4X420 A0A182FRA2 W5JDV0 A0A069DX50 A0A195B3I6 A0A182TVG4 A0A182NZ96 A0A182W014 A0A0C9R602 A0A182MJT8 A0A182R0S2 A0A1I8Q8D8 A0A1A9ZR04 B0WTA5 A0A1A9UF38 A0A182J5J6 A0A0J7KUH0 A0A1A9WPE1 A0A1B0FJF4 A0A1A9YRN8 N6U6M8 A0A182PND3 A0A1B0B4A0 A0A182HZU2 A0A182XGI4 B4KAG5 B3MXA1 B3P0R8 A0A1W4VN90 B4QZ41 B4PS79 B4HE08 Q9VFB6 B4GMM2 Q294T8 E0V8Z8 B4JGB2 A0A182Y458 B4LY26 A0A1W4WT53 A0A3B0K6J4 W8BMW7 A0A084W3H3 A0A0K8UJT7 A0A034VDE6 A0A0R3NJJ4 A0A0K8WI48 B4NBB4 A0A0A1X830 A0A0M4EW80 A0A182RNW4 A0A3R7PS56 A0A1B6KJ19 A0A2R5LBM2 A0A210Q0E7 A0A0B7ACK4 T1J9C7 K1Q6P1 A0A0B7AA75 A0A1S3HG43 L7M9L0 B7P0S7
Pubmed
22118469
26227816
26354079
21719571
28004739
21347285
+ More
20798317 18362917 19820115 20075255 28648823 17510324 26108605 26483478 24845553 25315136 20966253 12364791 14747013 17210077 20920257 23761445 26334808 23537049 17994087 17550304 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 15632085 23185243 20566863 25244985 24495485 24438588 25348373 25830018 28812685 22992520 25576852
20798317 18362917 19820115 20075255 28648823 17510324 26108605 26483478 24845553 25315136 20966253 12364791 14747013 17210077 20920257 23761445 26334808 23537049 17994087 17550304 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 15632085 23185243 20566863 25244985 24495485 24438588 25348373 25830018 28812685 22992520 25576852
EMBL
ODYU01005012
SOQ45482.1
NWSH01000012
PCG80885.1
GDQN01001443
JAT89611.1
+ More
AGBW02011690 OWR46296.1 JTDY01011872 KOB52372.1 KQ460106 KPJ17780.1 KQ459602 KPI93571.1 KQ978273 KYM95786.1 KQ980765 KYN13418.1 GL888273 EGI63560.1 GEZM01098364 GEZM01098363 JAV53911.1 KQ981744 KYN36236.1 KZ288212 PBC32810.1 ADTU01021606 KQ435732 KOX77328.1 KQ982691 KYQ52266.1 AJWK01022430 GL450241 EFN81303.1 KQ971307 EFA11199.1 KQ414855 KOC60115.1 KQ434816 KZC06860.1 GFDF01003511 JAV10573.1 NNAY01002052 OXU22236.1 CH477270 EAT45318.1 JRES01000776 KNC28427.1 JXUM01141446 KQ569247 KXJ68704.1 KK852605 KDR20408.1 KA648161 AFP62790.1 NEVH01024424 PNF17267.1 AAAB01008987 EAA00950.3 GFTR01008066 JAW08360.1 GL442888 EFN62823.1 ADMH02001839 ETN60964.1 GBGD01000508 JAC88381.1 KQ976641 KYM78759.1 GBYB01008217 JAG77984.1 AXCM01001384 AXCN02000272 DS232083 EDS34333.1 LBMM01003103 KMQ93968.1 CCAG010018530 APGK01047197 APGK01047198 KB741077 ENN74217.1 JXJN01008224 APCN01002011 CH933806 EDW15678.1 CH902629 EDV35324.1 CH954181 EDV48966.1 CM000364 EDX12869.1 CM000160 EDW97499.1 CH480815 EDW42100.1 AE014297 AY058625 AAF55144.1 AAL13854.1 CH479185 EDW38096.1 CM000070 CH673789 EAL28876.1 EDY71553.1 DS234988 EEB09854.1 CH916369 EDV92581.1 CH940650 EDW66892.1 OUUW01000008 SPP83690.1 GAMC01006423 JAC00133.1 ATLV01020011 KE525292 KFB44767.1 GDHF01025508 JAI26806.1 GAKP01017631 JAC41321.1 KRT01146.1 KRT05560.1 GDHF01001491 JAI50823.1 CH964232 EDW81078.1 GBXI01007221 JAD07071.1 CP012526 ALC47742.1 QCYY01001796 ROT75270.1 GEBQ01028534 JAT11443.1 GGLE01002742 MBY06868.1 NEDP02005310 OWF42207.1 HACG01030830 CEK77695.1 JH431971 JH818393 EKC32427.1 HACG01030828 CEK77693.1 GACK01005196 JAA59838.1 ABJB010212080 ABJB010437114 ABJB010923066 DS612847 EEC00207.1
AGBW02011690 OWR46296.1 JTDY01011872 KOB52372.1 KQ460106 KPJ17780.1 KQ459602 KPI93571.1 KQ978273 KYM95786.1 KQ980765 KYN13418.1 GL888273 EGI63560.1 GEZM01098364 GEZM01098363 JAV53911.1 KQ981744 KYN36236.1 KZ288212 PBC32810.1 ADTU01021606 KQ435732 KOX77328.1 KQ982691 KYQ52266.1 AJWK01022430 GL450241 EFN81303.1 KQ971307 EFA11199.1 KQ414855 KOC60115.1 KQ434816 KZC06860.1 GFDF01003511 JAV10573.1 NNAY01002052 OXU22236.1 CH477270 EAT45318.1 JRES01000776 KNC28427.1 JXUM01141446 KQ569247 KXJ68704.1 KK852605 KDR20408.1 KA648161 AFP62790.1 NEVH01024424 PNF17267.1 AAAB01008987 EAA00950.3 GFTR01008066 JAW08360.1 GL442888 EFN62823.1 ADMH02001839 ETN60964.1 GBGD01000508 JAC88381.1 KQ976641 KYM78759.1 GBYB01008217 JAG77984.1 AXCM01001384 AXCN02000272 DS232083 EDS34333.1 LBMM01003103 KMQ93968.1 CCAG010018530 APGK01047197 APGK01047198 KB741077 ENN74217.1 JXJN01008224 APCN01002011 CH933806 EDW15678.1 CH902629 EDV35324.1 CH954181 EDV48966.1 CM000364 EDX12869.1 CM000160 EDW97499.1 CH480815 EDW42100.1 AE014297 AY058625 AAF55144.1 AAL13854.1 CH479185 EDW38096.1 CM000070 CH673789 EAL28876.1 EDY71553.1 DS234988 EEB09854.1 CH916369 EDV92581.1 CH940650 EDW66892.1 OUUW01000008 SPP83690.1 GAMC01006423 JAC00133.1 ATLV01020011 KE525292 KFB44767.1 GDHF01025508 JAI26806.1 GAKP01017631 JAC41321.1 KRT01146.1 KRT05560.1 GDHF01001491 JAI50823.1 CH964232 EDW81078.1 GBXI01007221 JAD07071.1 CP012526 ALC47742.1 QCYY01001796 ROT75270.1 GEBQ01028534 JAT11443.1 GGLE01002742 MBY06868.1 NEDP02005310 OWF42207.1 HACG01030830 CEK77695.1 JH431971 JH818393 EKC32427.1 HACG01030828 CEK77693.1 GACK01005196 JAA59838.1 ABJB010212080 ABJB010437114 ABJB010923066 DS612847 EEC00207.1
Proteomes
UP000218220
UP000007151
UP000037510
UP000053240
UP000053268
UP000078542
+ More
UP000078492 UP000007755 UP000078541 UP000242457 UP000005205 UP000005203 UP000053105 UP000075809 UP000092461 UP000008237 UP000007266 UP000053825 UP000076502 UP000002358 UP000215335 UP000008820 UP000037069 UP000069940 UP000249989 UP000075881 UP000027135 UP000235965 UP000095301 UP000075882 UP000007062 UP000075903 UP000000311 UP000192223 UP000069272 UP000000673 UP000078540 UP000075902 UP000075884 UP000075920 UP000075883 UP000075886 UP000095300 UP000092445 UP000002320 UP000078200 UP000075880 UP000036403 UP000091820 UP000092444 UP000092443 UP000019118 UP000075885 UP000092460 UP000075840 UP000076407 UP000009192 UP000007801 UP000008711 UP000192221 UP000000304 UP000002282 UP000001292 UP000000803 UP000008744 UP000001819 UP000009046 UP000001070 UP000076408 UP000008792 UP000268350 UP000030765 UP000007798 UP000092553 UP000075900 UP000283509 UP000242188 UP000005408 UP000085678 UP000001555
UP000078492 UP000007755 UP000078541 UP000242457 UP000005205 UP000005203 UP000053105 UP000075809 UP000092461 UP000008237 UP000007266 UP000053825 UP000076502 UP000002358 UP000215335 UP000008820 UP000037069 UP000069940 UP000249989 UP000075881 UP000027135 UP000235965 UP000095301 UP000075882 UP000007062 UP000075903 UP000000311 UP000192223 UP000069272 UP000000673 UP000078540 UP000075902 UP000075884 UP000075920 UP000075883 UP000075886 UP000095300 UP000092445 UP000002320 UP000078200 UP000075880 UP000036403 UP000091820 UP000092444 UP000092443 UP000019118 UP000075885 UP000092460 UP000075840 UP000076407 UP000009192 UP000007801 UP000008711 UP000192221 UP000000304 UP000002282 UP000001292 UP000000803 UP000008744 UP000001819 UP000009046 UP000001070 UP000076408 UP000008792 UP000268350 UP000030765 UP000007798 UP000092553 UP000075900 UP000283509 UP000242188 UP000005408 UP000085678 UP000001555
PRIDE
Interpro
SUPFAM
Gene 3D
ProteinModelPortal
A0A2H1VYA1
A0A2A4K9C8
A0A1E1WS27
A0A212EXS9
A0A0L7K4G6
A0A194RKB8
+ More
A0A194PJH2 A0A151I9N6 A0A195DKL1 F4WQG0 A0A1Y1K076 A0A195F775 A0A2A3EM99 A0A158NNI4 A0A087ZZH9 A0A0N0U6A0 A0A151WWS4 A0A1B0GJX0 E2BSQ7 D6W7G3 A0A0L7QNB1 A0A154P6V1 K7IN72 A0A1L8DVY6 A0A232EV58 Q17FM5 A0A0L0C869 A0A182G3G4 A0A182KH75 A0A067RC13 T1PHB3 A0A2J7PLR1 A0A1I8M973 A0A182LQW0 Q7PYV2 A0A182UP80 A0A224XIU0 E2AUU8 A0A1W4X420 A0A182FRA2 W5JDV0 A0A069DX50 A0A195B3I6 A0A182TVG4 A0A182NZ96 A0A182W014 A0A0C9R602 A0A182MJT8 A0A182R0S2 A0A1I8Q8D8 A0A1A9ZR04 B0WTA5 A0A1A9UF38 A0A182J5J6 A0A0J7KUH0 A0A1A9WPE1 A0A1B0FJF4 A0A1A9YRN8 N6U6M8 A0A182PND3 A0A1B0B4A0 A0A182HZU2 A0A182XGI4 B4KAG5 B3MXA1 B3P0R8 A0A1W4VN90 B4QZ41 B4PS79 B4HE08 Q9VFB6 B4GMM2 Q294T8 E0V8Z8 B4JGB2 A0A182Y458 B4LY26 A0A1W4WT53 A0A3B0K6J4 W8BMW7 A0A084W3H3 A0A0K8UJT7 A0A034VDE6 A0A0R3NJJ4 A0A0K8WI48 B4NBB4 A0A0A1X830 A0A0M4EW80 A0A182RNW4 A0A3R7PS56 A0A1B6KJ19 A0A2R5LBM2 A0A210Q0E7 A0A0B7ACK4 T1J9C7 K1Q6P1 A0A0B7AA75 A0A1S3HG43 L7M9L0 B7P0S7
A0A194PJH2 A0A151I9N6 A0A195DKL1 F4WQG0 A0A1Y1K076 A0A195F775 A0A2A3EM99 A0A158NNI4 A0A087ZZH9 A0A0N0U6A0 A0A151WWS4 A0A1B0GJX0 E2BSQ7 D6W7G3 A0A0L7QNB1 A0A154P6V1 K7IN72 A0A1L8DVY6 A0A232EV58 Q17FM5 A0A0L0C869 A0A182G3G4 A0A182KH75 A0A067RC13 T1PHB3 A0A2J7PLR1 A0A1I8M973 A0A182LQW0 Q7PYV2 A0A182UP80 A0A224XIU0 E2AUU8 A0A1W4X420 A0A182FRA2 W5JDV0 A0A069DX50 A0A195B3I6 A0A182TVG4 A0A182NZ96 A0A182W014 A0A0C9R602 A0A182MJT8 A0A182R0S2 A0A1I8Q8D8 A0A1A9ZR04 B0WTA5 A0A1A9UF38 A0A182J5J6 A0A0J7KUH0 A0A1A9WPE1 A0A1B0FJF4 A0A1A9YRN8 N6U6M8 A0A182PND3 A0A1B0B4A0 A0A182HZU2 A0A182XGI4 B4KAG5 B3MXA1 B3P0R8 A0A1W4VN90 B4QZ41 B4PS79 B4HE08 Q9VFB6 B4GMM2 Q294T8 E0V8Z8 B4JGB2 A0A182Y458 B4LY26 A0A1W4WT53 A0A3B0K6J4 W8BMW7 A0A084W3H3 A0A0K8UJT7 A0A034VDE6 A0A0R3NJJ4 A0A0K8WI48 B4NBB4 A0A0A1X830 A0A0M4EW80 A0A182RNW4 A0A3R7PS56 A0A1B6KJ19 A0A2R5LBM2 A0A210Q0E7 A0A0B7ACK4 T1J9C7 K1Q6P1 A0A0B7AA75 A0A1S3HG43 L7M9L0 B7P0S7
PDB
4P17
E-value=3.30556e-27,
Score=306
Ontologies
KEGG
GO
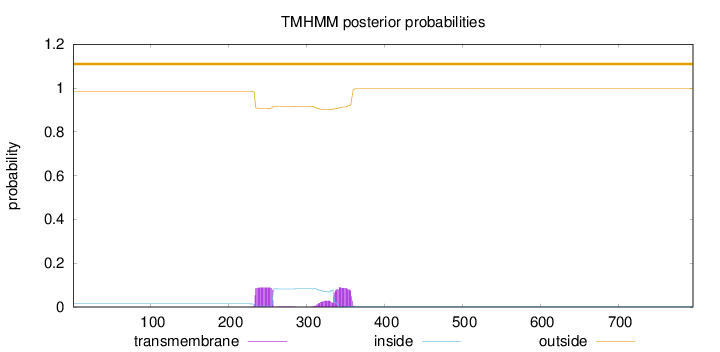
Topology
Length:
796
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
4.54366999999999
Exp number, first 60 AAs:
0.00011
Total prob of N-in:
0.01428
outside
1 - 796
Population Genetic Test Statistics
Pi
5.972249
Theta
12.056915
Tajima's D
-1.420845
CLR
17.691098
CSRT
0.0705964701764912
Interpretation
Uncertain
