Gene
KWMTBOMO16556 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA010203
Annotation
serine_protease_[Bombyx_mandarina]
Location in the cell
Extracellular Reliability : 2.611
Sequence
CDS
ATGCGCACGCAGCGAAAGCTCTCAAAATGGAAAAAAATTCCGATTTTTCCAGCTGCTTCGGGAGCCAACAACCAACAAAAACGCCAAGTGAGCCTCCAGGTCATTACCAACGCCGTCTGCGCCCGCACGTACGGAAACTCTGTGATCATTGGCTCCACCCTCTGTGTTTCCGGCGCTAACGGTCGCAGCACCTGCAGCGGAGACTCCGGCGGCCCTCTCTCCATCGGCAGCGGCGGAGGCCGTCAGCTGGTCAGTACACTAGTTTTGTTTAACTTTACTGCTTTAACCGGTTGCCTTTTTTTTATTGCTTAG
Protein
MRTQRKLSKWKKIPIFPAASGANNQQKRQVSLQVITNAVCARTYGNSVIIGSTLCVSGANGRSTCSGDSGGPLSIGSGGGRQLVSTLVLFNFTALTGCLFFIA
Summary
Similarity
Belongs to the peptidase S1 family.
Uniprot
EMBL
BABH01011411
BABH01011412
BABH01011413
BABH01041853
AY945210
EU338400
+ More
AAX39408.1 ABY58153.1 BABH01041861 AB117641 BAD13342.1 BABH01042790 AB003670 BAA20136.1 BABH01041852 AY945211 DQ310733 AAX39409.1 ABC26003.1 BABH01041859 BABH01041860 KR024679 ALE15221.1 RSAL01000073 RVE48982.1 KR024673 ALE15215.1
AAX39408.1 ABY58153.1 BABH01041861 AB117641 BAD13342.1 BABH01042790 AB003670 BAA20136.1 BABH01041852 AY945211 DQ310733 AAX39409.1 ABC26003.1 BABH01041859 BABH01041860 KR024679 ALE15221.1 RSAL01000073 RVE48982.1 KR024673 ALE15215.1
Proteomes
Pfam
PF00089 Trypsin
Interpro
SUPFAM
SSF50494
SSF50494
CDD
ProteinModelPortal
Ontologies
GO
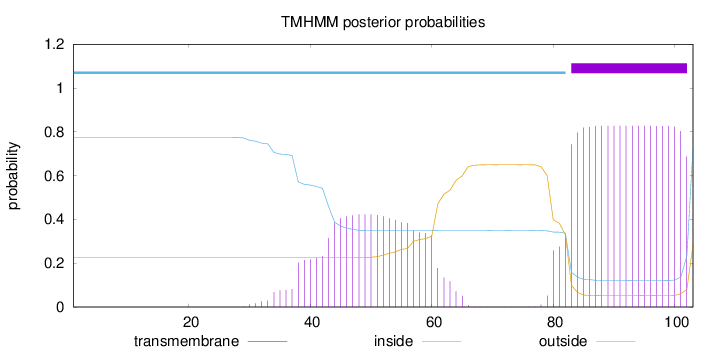
Topology
Length:
103
Number of predicted TMHs:
1
Exp number of AAs in TMHs:
26.19571
Exp number, first 60 AAs:
8.44741
Total prob of N-in:
0.77395
inside
1 - 82
TMhelix
83 - 102
outside
103 - 103
Population Genetic Test Statistics
Pi
0
Theta
0
Tajima's D
0
CLR
0
CSRT
0
Interpretation
Uncertain
