Pre Gene Modal
BGIBMGA005637
Annotation
putative_penguin_[Heliconius_melpomene]
Location in the cell
Nuclear Reliability : 3.629
Sequence
CDS
ATGCAGAAAGGTAAAAGGAAAAGTGATGAAGATGGTGTTAGTTCACCAAAAAAGAAAAAACTAAATAGTTCGCAAACAACAGTTCGCAAAGAACAAGATAGTAAAATTAAAAACATCAAAAACAAAACAAACCTAAAAGGTAAAAAAAATAATAAGGATGGATTAAAAGGTTTCAAAAAATTTACTAAAAACGACAAAGGATCTAAATTTGTTAAAACAAAAGAAAAATCGAATTTTTCTAAAGATGAAGCAAACGGAGAAAAGCCAAAATGGGCTGAAATGAAAAAAGAGAAAAAACAATTAAGGACTGTAAGACGTAAAGCTAAAATAACAGCTGAAGTATTTGAGATTTCACATAAGGCCAAGTTGTTAGCAGCACAAATACAGAGAAAAGTTGTGAAATCTGACTTCAAAATAAACTCCTGTAAAGAATTGCACAATATGTTAAAGGGACAATACAAGTCTATTGCTCTCACTCATGATCTCAGTCGAGTTATTCAAGTTCTACTAAAACATAGCCCTGAAGACATTATAAATGAAATAACGAAAGAGCTTCTAGACATTATAGTACAGATGGGTCAATCTAAGTATGCGCATCATTCCGTTAAGCGAATACTGAAATATGGAACAGATTTCATTAGGCATGAGGTTTTGAAGAAGTTTTATGGACATGTTGTATCCCTGTCAACTCATGCTATTAGTGCACCTGTCTTGGATTATGCATATGGGGAGTTTGCCTCGAAAAAGGAAAAATGCCACATGCAACAAGAATTCTATGGAGAGATTTATAAGAATACAAAAGACGACAAGGTAAAGACACTCAGAGATGCATACAAAGACAGTCCAGAAATGAAATCAGCTATACTCCAGGCCTGTAAGGCCAACTTACAGAAGATATTAGACAAAAACCTACATGATGGAGAATTGTTTCATTCTGTGTTATATGACTATATCTGTGAATGCAGCCAAGAAGATAGAGTAGATTTTATTTCTATTCTTAGCCCACTGATTGTCCCACTCAGTAATTCTTTACCTGGTGTCAATGCTGCCAGTATGTGTGTTTGGCAGGGTACAAATAAGGACAAGAAAACAATTTTAAAAGTGGTCAAGGAACATGCTATTCCTTTAAGTAAACATAAGACTGGATACCGCCTTCTGATTGCTATTTTTGACTCTGTTGACGATACTGTTCTTGTCAAGAAGACTATTGTGTCCACTCTGGCTGCCAATATAAAAGACATTGCTAAGGACCACTGGGGTAATATGGTAATTCACTGGTTAGTAAAACCAAAAGACACAGCTGCTTTCCATCCTTGTTTAATTAAGTTTTTGGAAGATGGTGCTAAGAGTGGAACATCGAAGAAAGACACAGAAATTCGTGTTTCGGAGCTTAGGGAAGCTATATTACCTGCTCTAAAGAAAGATATGGAAACAGATCCTAAATTCTGGCTAAGCAATAAATCTAATATGTTGTTGGCTGTTGCTGTATTGTCTATAGAATCATCTAAAGAGATTCTGCATGCATTGGCAAAAGTGATCTGCGAGACTGACTGGCACATGCAGGTGGAAGGGAAGGACGATACCCTGGCGATTGAGGATGCTGGTATACACATGTGCTTAAAAAAACTAGCCAGTTTAGACAAAGAAAAATCGGCTGATACTCTTGGTGAAGCTATAAGTGAACATTTAAATGATGAAACTATTAAAGTGTGGATACGTACAAATAGGGGTTGCTTTTTCCTATTGAAATTGATAGAGAGTAATAGTGAAGAAGTAGATAAGAAATTAAAGGCAAAATTAAAACCACATGTAAATATTTTGAAGCAACAATCATCGGAAGGTGCTAAATTATTGTTACAGAATGTACAGAATAAATAG
Protein
MQKGKRKSDEDGVSSPKKKKLNSSQTTVRKEQDSKIKNIKNKTNLKGKKNNKDGLKGFKKFTKNDKGSKFVKTKEKSNFSKDEANGEKPKWAEMKKEKKQLRTVRRKAKITAEVFEISHKAKLLAAQIQRKVVKSDFKINSCKELHNMLKGQYKSIALTHDLSRVIQVLLKHSPEDIINEITKELLDIIVQMGQSKYAHHSVKRILKYGTDFIRHEVLKKFYGHVVSLSTHAISAPVLDYAYGEFASKKEKCHMQQEFYGEIYKNTKDDKVKTLRDAYKDSPEMKSAILQACKANLQKILDKNLHDGELFHSVLYDYICECSQEDRVDFISILSPLIVPLSNSLPGVNAASMCVWQGTNKDKKTILKVVKEHAIPLSKHKTGYRLLIAIFDSVDDTVLVKKTIVSTLAANIKDIAKDHWGNMVIHWLVKPKDTAAFHPCLIKFLEDGAKSGTSKKDTEIRVSELREAILPALKKDMETDPKFWLSNKSNMLLAVAVLSIESSKEILHALAKVICETDWHMQVEGKDDTLAIEDAGIHMCLKKLASLDKEKSADTLGEAISEHLNDETIKVWIRTNRGCFFLLKLIESNSEEVDKKLKAKLKPHVNILKQQSSEGAKLLLQNVQNK
Summary
Uniprot
A0A2A4JDV9
A0A2H1VM78
A0A194QAR9
A0A194QTH1
D0AB89
L7X0R1
+ More
S4PGK9 A0A212EP73 A0A0L7LL06 D6WZ15 N6U5R4 A0A0T6B3C6 A0A1Y1MBF6 A0A232EX65 K7J9N0 E2BXL1 A0A088ALU9 A0A154PGZ6 A0A0J7K4J0 E9IFM9 A0A2P8XS97 E2A7S4 A0A026W9A4 A0A3L8DD65 A0A195DCV3 A0A195BCC0 A0A158NS35 A0A2J7QKP0 A0A0K8USZ1 F4WJT7 A0A195FXT3 A0A151WGY7 U5EHI9 A0A067QIM4 A0A1A9WZ92 A0A3L8DD77 W8BH01 A0A1B0AIX9 A0A336MJ44 A0A336MGJ9 A0A195BYX8 A0A1A9UHS6 A0A1B0D4D4 A0A1L8DGJ9 A0A1J1J034 A0A182H6R2 A0A1B0CMH0 A0A182X8S7 A0A182K363 A0A3Q0IPH3 A0A084VJS3 A0A2M4AJN3 A0A182L6U1 A0A182PAK3 A0A2M4AJE2 A0A2M3Z396 A0A1S4FNX6 A0A182F8N9 A0A2M3Z3B7 A0A182IVV3 A0A2M4AJW9 Q16UF7 A0A182UXD4 A0A182TG74 A0A2T7NPQ9 A0A2M4BGH9 A0A182LW26 A0A1S3JC21 A0A1S3JCA0 Q7QAB5 A0A1S3K9S0 A0A2C9JHF8 B4NCP2 A0A182ID10 A0A182XWT0 A0A182R5P3 A0A182N5H7 A0A2R7VX35 E9G064 A0A0P5W3J1
S4PGK9 A0A212EP73 A0A0L7LL06 D6WZ15 N6U5R4 A0A0T6B3C6 A0A1Y1MBF6 A0A232EX65 K7J9N0 E2BXL1 A0A088ALU9 A0A154PGZ6 A0A0J7K4J0 E9IFM9 A0A2P8XS97 E2A7S4 A0A026W9A4 A0A3L8DD65 A0A195DCV3 A0A195BCC0 A0A158NS35 A0A2J7QKP0 A0A0K8USZ1 F4WJT7 A0A195FXT3 A0A151WGY7 U5EHI9 A0A067QIM4 A0A1A9WZ92 A0A3L8DD77 W8BH01 A0A1B0AIX9 A0A336MJ44 A0A336MGJ9 A0A195BYX8 A0A1A9UHS6 A0A1B0D4D4 A0A1L8DGJ9 A0A1J1J034 A0A182H6R2 A0A1B0CMH0 A0A182X8S7 A0A182K363 A0A3Q0IPH3 A0A084VJS3 A0A2M4AJN3 A0A182L6U1 A0A182PAK3 A0A2M4AJE2 A0A2M3Z396 A0A1S4FNX6 A0A182F8N9 A0A2M3Z3B7 A0A182IVV3 A0A2M4AJW9 Q16UF7 A0A182UXD4 A0A182TG74 A0A2T7NPQ9 A0A2M4BGH9 A0A182LW26 A0A1S3JC21 A0A1S3JCA0 Q7QAB5 A0A1S3K9S0 A0A2C9JHF8 B4NCP2 A0A182ID10 A0A182XWT0 A0A182R5P3 A0A182N5H7 A0A2R7VX35 E9G064 A0A0P5W3J1
Pubmed
EMBL
NWSH01001832
PCG69966.1
ODYU01003152
SOQ41542.1
KQ459232
KPJ02633.1
+ More
KQ461155 KPJ08300.1 FP102340 CBH09284.1 KC469893 AGC92694.1 GAIX01003617 JAA88943.1 AGBW02013517 OWR43288.1 JTDY01000686 KOB76223.1 KQ971363 EFA09038.1 APGK01041594 KB740994 KB632255 ENN75986.1 ERL90654.1 LJIG01016117 KRT81565.1 GEZM01035601 JAV83159.1 NNAY01001792 OXU22907.1 AAZX01002711 GL451265 EFN79580.1 KQ434896 KZC10744.1 LBMM01014327 KMQ85219.1 GL762880 EFZ20617.1 PYGN01001432 PSN34880.1 GL437382 EFN70542.1 KK107374 EZA51594.1 QOIP01000010 RLU18102.1 KQ980989 KYN10708.1 KQ976528 KYM81867.1 ADTU01024454 NEVH01013260 PNF29156.1 GDHF01022839 JAI29475.1 GL888186 EGI65579.1 KQ981208 KYN44654.1 KQ983136 KYQ47082.1 GANO01003004 JAB56867.1 KK853308 KDR08684.1 RLU18103.1 GAMC01010297 GAMC01010295 JAB96258.1 UFQT01001201 SSX29481.1 UFQT01001175 SSX29325.1 KQ978457 KYM93817.1 AJVK01011378 AJVK01011379 GFDF01008496 JAV05588.1 CVRI01000066 CRL05875.1 JXUM01114779 JXUM01114780 JXUM01114781 JXUM01114782 JXUM01114783 JXUM01114784 KQ565817 KXJ70629.1 AJWK01018717 ATLV01013862 KE524905 KFB38217.1 GGFK01007636 MBW40957.1 GGFK01007598 MBW40919.1 GGFM01002212 MBW22963.1 GGFM01002187 MBW22938.1 GGFK01007750 MBW41071.1 CH477622 EAT38180.1 PZQS01000010 PVD23157.1 GGFJ01002986 MBW52127.1 AXCM01006361 AAAB01008898 EAA09186.4 CH964239 EDW82601.2 APCN01004963 KK854135 PTY11899.1 GL732528 EFX87453.1 GDIP01092481 JAM11234.1
KQ461155 KPJ08300.1 FP102340 CBH09284.1 KC469893 AGC92694.1 GAIX01003617 JAA88943.1 AGBW02013517 OWR43288.1 JTDY01000686 KOB76223.1 KQ971363 EFA09038.1 APGK01041594 KB740994 KB632255 ENN75986.1 ERL90654.1 LJIG01016117 KRT81565.1 GEZM01035601 JAV83159.1 NNAY01001792 OXU22907.1 AAZX01002711 GL451265 EFN79580.1 KQ434896 KZC10744.1 LBMM01014327 KMQ85219.1 GL762880 EFZ20617.1 PYGN01001432 PSN34880.1 GL437382 EFN70542.1 KK107374 EZA51594.1 QOIP01000010 RLU18102.1 KQ980989 KYN10708.1 KQ976528 KYM81867.1 ADTU01024454 NEVH01013260 PNF29156.1 GDHF01022839 JAI29475.1 GL888186 EGI65579.1 KQ981208 KYN44654.1 KQ983136 KYQ47082.1 GANO01003004 JAB56867.1 KK853308 KDR08684.1 RLU18103.1 GAMC01010297 GAMC01010295 JAB96258.1 UFQT01001201 SSX29481.1 UFQT01001175 SSX29325.1 KQ978457 KYM93817.1 AJVK01011378 AJVK01011379 GFDF01008496 JAV05588.1 CVRI01000066 CRL05875.1 JXUM01114779 JXUM01114780 JXUM01114781 JXUM01114782 JXUM01114783 JXUM01114784 KQ565817 KXJ70629.1 AJWK01018717 ATLV01013862 KE524905 KFB38217.1 GGFK01007636 MBW40957.1 GGFK01007598 MBW40919.1 GGFM01002212 MBW22963.1 GGFM01002187 MBW22938.1 GGFK01007750 MBW41071.1 CH477622 EAT38180.1 PZQS01000010 PVD23157.1 GGFJ01002986 MBW52127.1 AXCM01006361 AAAB01008898 EAA09186.4 CH964239 EDW82601.2 APCN01004963 KK854135 PTY11899.1 GL732528 EFX87453.1 GDIP01092481 JAM11234.1
Proteomes
UP000218220
UP000053268
UP000053240
UP000007151
UP000037510
UP000007266
+ More
UP000019118 UP000030742 UP000215335 UP000002358 UP000008237 UP000005203 UP000076502 UP000036403 UP000245037 UP000000311 UP000053097 UP000279307 UP000078492 UP000078540 UP000005205 UP000235965 UP000007755 UP000078541 UP000075809 UP000027135 UP000091820 UP000092445 UP000078542 UP000078200 UP000092462 UP000183832 UP000069940 UP000249989 UP000092461 UP000076407 UP000075881 UP000079169 UP000030765 UP000075882 UP000075885 UP000069272 UP000075880 UP000008820 UP000075903 UP000075902 UP000245119 UP000075883 UP000085678 UP000007062 UP000076420 UP000007798 UP000075840 UP000076408 UP000075900 UP000075884 UP000000305
UP000019118 UP000030742 UP000215335 UP000002358 UP000008237 UP000005203 UP000076502 UP000036403 UP000245037 UP000000311 UP000053097 UP000279307 UP000078492 UP000078540 UP000005205 UP000235965 UP000007755 UP000078541 UP000075809 UP000027135 UP000091820 UP000092445 UP000078542 UP000078200 UP000092462 UP000183832 UP000069940 UP000249989 UP000092461 UP000076407 UP000075881 UP000079169 UP000030765 UP000075882 UP000075885 UP000069272 UP000075880 UP000008820 UP000075903 UP000075902 UP000245119 UP000075883 UP000085678 UP000007062 UP000076420 UP000007798 UP000075840 UP000076408 UP000075900 UP000075884 UP000000305
Pfam
PF08144 CPL
Interpro
SUPFAM
SSF48371
SSF48371
Gene 3D
ProteinModelPortal
A0A2A4JDV9
A0A2H1VM78
A0A194QAR9
A0A194QTH1
D0AB89
L7X0R1
+ More
S4PGK9 A0A212EP73 A0A0L7LL06 D6WZ15 N6U5R4 A0A0T6B3C6 A0A1Y1MBF6 A0A232EX65 K7J9N0 E2BXL1 A0A088ALU9 A0A154PGZ6 A0A0J7K4J0 E9IFM9 A0A2P8XS97 E2A7S4 A0A026W9A4 A0A3L8DD65 A0A195DCV3 A0A195BCC0 A0A158NS35 A0A2J7QKP0 A0A0K8USZ1 F4WJT7 A0A195FXT3 A0A151WGY7 U5EHI9 A0A067QIM4 A0A1A9WZ92 A0A3L8DD77 W8BH01 A0A1B0AIX9 A0A336MJ44 A0A336MGJ9 A0A195BYX8 A0A1A9UHS6 A0A1B0D4D4 A0A1L8DGJ9 A0A1J1J034 A0A182H6R2 A0A1B0CMH0 A0A182X8S7 A0A182K363 A0A3Q0IPH3 A0A084VJS3 A0A2M4AJN3 A0A182L6U1 A0A182PAK3 A0A2M4AJE2 A0A2M3Z396 A0A1S4FNX6 A0A182F8N9 A0A2M3Z3B7 A0A182IVV3 A0A2M4AJW9 Q16UF7 A0A182UXD4 A0A182TG74 A0A2T7NPQ9 A0A2M4BGH9 A0A182LW26 A0A1S3JC21 A0A1S3JCA0 Q7QAB5 A0A1S3K9S0 A0A2C9JHF8 B4NCP2 A0A182ID10 A0A182XWT0 A0A182R5P3 A0A182N5H7 A0A2R7VX35 E9G064 A0A0P5W3J1
S4PGK9 A0A212EP73 A0A0L7LL06 D6WZ15 N6U5R4 A0A0T6B3C6 A0A1Y1MBF6 A0A232EX65 K7J9N0 E2BXL1 A0A088ALU9 A0A154PGZ6 A0A0J7K4J0 E9IFM9 A0A2P8XS97 E2A7S4 A0A026W9A4 A0A3L8DD65 A0A195DCV3 A0A195BCC0 A0A158NS35 A0A2J7QKP0 A0A0K8USZ1 F4WJT7 A0A195FXT3 A0A151WGY7 U5EHI9 A0A067QIM4 A0A1A9WZ92 A0A3L8DD77 W8BH01 A0A1B0AIX9 A0A336MJ44 A0A336MGJ9 A0A195BYX8 A0A1A9UHS6 A0A1B0D4D4 A0A1L8DGJ9 A0A1J1J034 A0A182H6R2 A0A1B0CMH0 A0A182X8S7 A0A182K363 A0A3Q0IPH3 A0A084VJS3 A0A2M4AJN3 A0A182L6U1 A0A182PAK3 A0A2M4AJE2 A0A2M3Z396 A0A1S4FNX6 A0A182F8N9 A0A2M3Z3B7 A0A182IVV3 A0A2M4AJW9 Q16UF7 A0A182UXD4 A0A182TG74 A0A2T7NPQ9 A0A2M4BGH9 A0A182LW26 A0A1S3JC21 A0A1S3JCA0 Q7QAB5 A0A1S3K9S0 A0A2C9JHF8 B4NCP2 A0A182ID10 A0A182XWT0 A0A182R5P3 A0A182N5H7 A0A2R7VX35 E9G064 A0A0P5W3J1
PDB
4WZW
E-value=3.83217e-50,
Score=502
Ontologies
GO
PANTHER
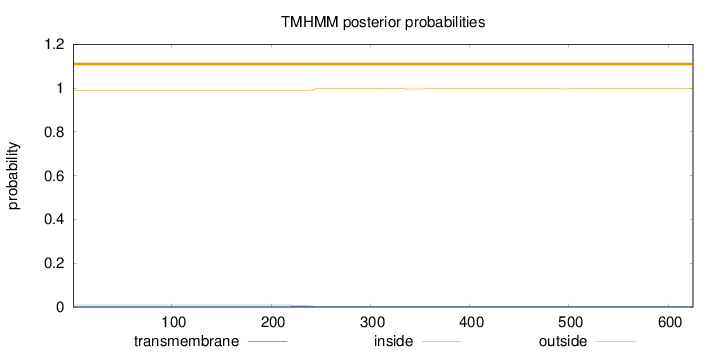
Topology
Length:
625
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.14368
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00960
outside
1 - 625
Population Genetic Test Statistics
Pi
2.314358
Theta
23.462693
Tajima's D
-1.321643
CLR
0.003159
CSRT
0.0837458127093645
Interpretation
Uncertain
