Gene
KWMTBOMO16492 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA004585
Annotation
vitellogenin_precursor_[Bombyx_mori]
Full name
Vitellogenin
Location in the cell
Nuclear Reliability : 3.302
Sequence
CDS
ATGAAGCTATTCGTCTTGGCGGCAATTATAGCCGCCGTGTCCTCGGATCGGTTTAGCAGCCAGAGCCAGTCCACAGGCGGACAAACATATCCATCACCTTGGCAGGTTGGCAAACAATACCGGTATGAGGTCACTTCTCGCACTTTGGCGCATTTGCAGGAGGGTCCAAGCAGCGGAAGCGCTTTCAAAGCACAATTCACAATCCGGGTAAAGAGTCCTGGCCGACTCCAAGCGAAACTAGAGAATCCACAGCATGGTAACTTCAATGAGCAACTCCCTGACCCGAGGGAACTGCCTGTTGATCTCAAGTACCAACCTACGCCGAACATCGATAAGGTATTCGAAATTGAAATTGACGGAGGTCGAATCGTAAGCCTCGATTTTCCAACTTCAGTTCCTGTCCCCCAAGAGAATTTAATAAAGGGTCTTATCAGCGCTCTGCAACTTGACACGTCGGCTCACAGAATCATTCACGACTCTCAAAACAACTACGACAGAGAACAGCAACAGGGCCTTTTCAGGAAAATGGAGACTGATGTGACTGGTGATTGTGAGACGCTGTACACGGTATCTCCTGTTGCTTCAGAATGGCGGCGAGAGCTTCCGAAGTTCGCTAACGAACAGGACCCTGTTGAGGTTACAAAGAGCACAAATTACGGTCACTGTCACCACCGGGTCGCTTATCACTTCGGAGTGCCGGTCGGTGCTGAGTGGACTGGAACAGCGCACAAAACCCAAGAACAACAGCTCATTGGCCGCGCTACTTACTCCCGTATCTTAACAGGCAAAGAGGGACCTATTTACAAAGCTGAAACCACGAGCACCGTTCATGTCCACCCTCATCTTTATGGTAAACAGAAAGCCGAAGTATACAGTCACGTCCATATGGAACTGATTTCAGTAGATCAAGACAGTGGGGCTGAATGGCCGAGAGCTGAAGCGATGCGTCCTGCTCAAAGCATTTTATATTCCTTAAGCACAAAACAAATGACTAAGCATTACGAGTCTTCTTCGTCATCGTCTTCTTCGGAATCTCACGAATTCAATTTCCCAGAGCAACACGAACACCCGCATCAGTCCAATCAACGATCGCGTCGTTCTTACATGAGATCTAAATTGGTGACTGTTCACAAAGTACTCAAAAAACGCAACAGCGAAAGCTCGTCTGGTTCTTCAAGTTCAAGTGCGGATTCATCTTCAGCGTACATTAATGATGACATCCCGGACATTGATGAGCCGGCGTATGCAGCATTATATATGAGCCCTCAACCTCACGCGGACAAAAAGCAGAATGCAATGAACGCACAAAAGATTCTTCAAGACATTGCACAACAGTTGCAAAATCCTAATAATATGCCTAAATCTGATTTCCTATCCAAATTTAATATCCTCGTTCGACTTATCGCCAGTATGAGCACCGAACAGCTCAGCCAAACTAGTAGAAGCATTGAAACTGCGAAGACTTCGAACAATATCATTAAATCTGACATGTGGATGATATTCCGTGATGGTGTCACTCAAGCTGGTACGTTGCCAGCTTTCAAACAAATTCAGTCTTGGATTGAGAACAAAAAAATCCAAGAGGAAGAAGCTGCACAAGTAGTCGCTGCTTTGCCTAGGACCCTACGCTACCCCACTAAGCAAATTATGACACAGTTCTTCAATTTTGCCAGAAGTCCTGCAGTCAAAGATCAAATGTTCCTCAATTCGAGTGCCCTGATGGCTGCCACAAAATTGATAAATTTGGGCCAAGTTAATAACTATACCGCTCACTCTTATTATCCGACACACATGTATGGTCGCCTTACGCACAAACATGACGCTTTTGTTCTCGAAGAAATTTTGCCTACTCTAGCAGCGGACTTGAAAGCATCCGTTGAATATAAAGATAGTACGAAAGCTCAAGTTTACATTCAAGCTATTGGTAATTTGGGTCATCGCGAAATACTCAAAGTCTTCGCTCCATATTTGGAAGGAAAAGTTGAAATATCTACTTATCTTCGCACTCATATTGTGAAGAATTTAAAATCTTTGGCTAAGCTAAGAGATCGGCATGTACGTGCTGTCCTGTTTAGCATTTTAAGAAACACGGCTGAGCCATATCCTGTTAGAGTTGCAGCGATCCAGAGTATATTCATAAGCCACCCCACCGGAGAAATGATGCAAGCTATGGCTGAAATGACCCACAACGATCCAAGTGTAGAAGTTCGGGCTGTTCTTAAGTCCGCCATTTTATCAGCTGCTGAATTACAACACCCACGCAATTTTTATCTATCTAGAACGGCTCAAGCTGCTAGATACTTGGTTACAAATGAAGAATTTGGTTACCAGCATTCATTTAAGTTTATTGATGATAGCTACGACGAAGATAATGACATTGGGACTTTCGTTATTTCCCACATCGGTAGCGAGGATAGTCTTCTGCCAAAAGATTTTAAAATCGTCACCAATTCCAAGGGAGGAGCCTGGGAAAGAAACACAATTGAAGCATCATTCTCTAGTGCTGAACGTTTCTTAGACTACTTAAGGGACTCGGTATTCGCACCTCACCCTAAATTCGATAGAGCTCACAAATATTCCGCTGAGAAAATCGCTAAGCTCTTAAATATAAAGAATGATGAGGAAGAGCCTTTAGAAGCATCTTTTTATGTCGATTTCATGAACAACCAGAGATTATTCAGTTTCAGCGAATCCGACCTACAGCAGTTGTCCCAATATATTAGCGAATACATGAAAAAAGTAGAGAGTGGCGCTGAAAAGCATTACACAAAAGTTTATAACCAGGATCAAGTTTCAATAATGTTCCCAGTAGCATCTGGTATGCCATTCATCTTTAAATATAAAGAACCGGCTGTGATTCATTTCCAAAGCAAACTCAAAGGGAAATTTTCATTCCCGTCAAAGGATAACAAATACTACGAGGCAAACATGATTAAAGACGTGCAATTCACATACGCCAGGAATATTGACGGAAACGTTGGCTTTATGGATACCCTTAGCAACCAATATTCCAGTGTAGGTGTTGTGAATAAACTCCAATTTAACATACCGTTCAAATTCGGAATCGAAATTAAATCTGGACTAATAAAGTTCAGAGTCGAACCTCTGCATCCAGACCAGGATCAGACCTTAGTTCACTATAGTGTTTGGCCGTACAGTGCCAGCCAAAAGAAGGATTCCCTTGTTGCAATCTCTCAAGACCCAGCTACTAAAATTGTCGAACGTAGAAGCAAGGTATTTTCAGTAGACTCTAAGTACGGTCAATCCACACATGCCGTAATCTACGCGCAAGGATACACATACTCTTCCGACTGGAGGAACTTTGGCGCTAAGTTTACTTCCCGTGACTATTTTACCAACTTAGCGTCACTACTGACACAGGAGGATATCGCTCTGACCCACTTCAACTTAAAACACCTTTGCAAACAATCACAGAGTAAGGCTTTGACCATTACCGCTTATTATGATGAATACTACAACCAACAGAACAGCGGTATATTAACAGACGCTACTGATCGCAACGACCTCAGTCCGAACAGCGAAACACGTCGAGCCGAAATGGTGAAACTAGTTTCTGCTGGCATTAACAAAGCTAGAGTGAGAGTTGTGGATTTGAGTGCTAGCTTTGAGGGCAGTCAAGATCAGAATTACGTTTTCACTGGCACCTGGGGTGATAGTCCGGTCGATTCAAAGGTTCAAGGAATGTTATTTGCTGGCACAAAGTCTGCAACGCAAGGAAACCAACAAATAAACGCTGTATTTGCCACTACGAAGCCGGAGATTCATTCCCTAAGCTTTTCGAAGGCTTTACAGAGCGATTTGAGGGCTCCTTTCGGAATGCACTTCAAGTACGGCCAGAGTGGAGAGATCCGTGTCTCTGGAAGCTTCGATCGTACCAAAAAATATACAACTGAACTAGAAAATCATCCTCTCGCCAAGCAATGCTCTCAACAAACCACCCTCAATAACTTCTACCAAGATTCTTGTCACAAAGCTATCGTTATGGCTCATGCGCCTGATCATGTGGAATTCTCCGTCTCATTCCAAGACATGAGTCCTCAATACAGAAACTTCTCCTACCACACATACAGACTCTATGAGTACTTGGGTTACTGGTATACTGAAGCTAATCCTTTGAAACTAACTCAAAATGGTAAAATGGACTTCAAAATTGATTTCTCGTACTTCGATCGTACTTACACCGTCGATATTGCATCCCCATCTGGAGAGGCACGAATGAGGGACATGCCAATTGCAACAATGGCACCTGGTGCTTTGTCTTTCTACCAACCGCTAAAAGCTTATGAACTCGTTGCCAATTACTTCACTGGCCACCAATATCAACCCTACTGTTCTATCGATGGCACAAGGATACATACCTTCAGTAATCGCAGCTACGAATATCCCCTATCTCGATCCTGGCACGTAGTCATGCAGGACGAGTCTACTCAACGCGGCAACTGGCACGAGTTGGCTATCTTGTCCAGGAGGCAACAGAGAGATCAACAGGAGATCTATATTTCTTACAAGTCTGAGTCAGGACAGGATCTTGAGATCGAAATACAACCAGCGTCTGGAGACTCTGCATATCAAGTGAAAGTCACGACGAATACCAAGAAGATCACCGATGATGACCTCACCATGTACTGGGATGATGTCAAGGAGCAACCGTTCTTACAATACCACACCCACAAAGACGGCGTTCTGGTCATAAATATAGAAGACGACAGGATCAGAGCTATATATGACGGCCAGAGATTTGTCGTTTTCACACAGGACTATCGTAATTCGACCAGAGGCATCTGTGGACGTATGAGCGGCGAGCAGCGCGACGACTATCTCACTCCAGAAGGTTTGGTCGACAAGCCTGAGCTATATGCCGCAGCCTACAGCCTAAATGAGGAGAACAGTGACCCCAAAACCCAGGAACTGAAGGCTCTAGCAACACAACAAGCTTACTACCCAGAATATAAGTACACAAGCATCCTACGCTCTGACCCAACTTGGCAAGAAGAATCACAGTCCTCCGGTGAGGATCAATGGCAGTCTGAAACCGTCTATAAGTCTAGAAGCTACGACAAACACAAGGGCGCTTGTGAAGTTAGGCAACAGGTGCAGTTTTATGAGAACCACGGCGATATCTGTATCACAACTTCGAGAGTCCCCTCGTGTCAGTCGCATTGCCGTGCTGGAGACTACAAAATACAGCACGTTCAAGTGACGTGTAAATCGAAACTCGACCACGACTTCCGCATGTACAAAGAACAGATCAAAAAGGGCCAGAACCCCGAGGTATCTGGAATACCTAGCGTCAAGCAATTCAAAGTACCGGTAACTTGCCAGCCTTAA
Protein
MKLFVLAAIIAAVSSDRFSSQSQSTGGQTYPSPWQVGKQYRYEVTSRTLAHLQEGPSSGSAFKAQFTIRVKSPGRLQAKLENPQHGNFNEQLPDPRELPVDLKYQPTPNIDKVFEIEIDGGRIVSLDFPTSVPVPQENLIKGLISALQLDTSAHRIIHDSQNNYDREQQQGLFRKMETDVTGDCETLYTVSPVASEWRRELPKFANEQDPVEVTKSTNYGHCHHRVAYHFGVPVGAEWTGTAHKTQEQQLIGRATYSRILTGKEGPIYKAETTSTVHVHPHLYGKQKAEVYSHVHMELISVDQDSGAEWPRAEAMRPAQSILYSLSTKQMTKHYESSSSSSSSESHEFNFPEQHEHPHQSNQRSRRSYMRSKLVTVHKVLKKRNSESSSGSSSSSADSSSAYINDDIPDIDEPAYAALYMSPQPHADKKQNAMNAQKILQDIAQQLQNPNNMPKSDFLSKFNILVRLIASMSTEQLSQTSRSIETAKTSNNIIKSDMWMIFRDGVTQAGTLPAFKQIQSWIENKKIQEEEAAQVVAALPRTLRYPTKQIMTQFFNFARSPAVKDQMFLNSSALMAATKLINLGQVNNYTAHSYYPTHMYGRLTHKHDAFVLEEILPTLAADLKASVEYKDSTKAQVYIQAIGNLGHREILKVFAPYLEGKVEISTYLRTHIVKNLKSLAKLRDRHVRAVLFSILRNTAEPYPVRVAAIQSIFISHPTGEMMQAMAEMTHNDPSVEVRAVLKSAILSAAELQHPRNFYLSRTAQAARYLVTNEEFGYQHSFKFIDDSYDEDNDIGTFVISHIGSEDSLLPKDFKIVTNSKGGAWERNTIEASFSSAERFLDYLRDSVFAPHPKFDRAHKYSAEKIAKLLNIKNDEEEPLEASFYVDFMNNQRLFSFSESDLQQLSQYISEYMKKVESGAEKHYTKVYNQDQVSIMFPVASGMPFIFKYKEPAVIHFQSKLKGKFSFPSKDNKYYEANMIKDVQFTYARNIDGNVGFMDTLSNQYSSVGVVNKLQFNIPFKFGIEIKSGLIKFRVEPLHPDQDQTLVHYSVWPYSASQKKDSLVAISQDPATKIVERRSKVFSVDSKYGQSTHAVIYAQGYTYSSDWRNFGAKFTSRDYFTNLASLLTQEDIALTHFNLKHLCKQSQSKALTITAYYDEYYNQQNSGILTDATDRNDLSPNSETRRAEMVKLVSAGINKARVRVVDLSASFEGSQDQNYVFTGTWGDSPVDSKVQGMLFAGTKSATQGNQQINAVFATTKPEIHSLSFSKALQSDLRAPFGMHFKYGQSGEIRVSGSFDRTKKYTTELENHPLAKQCSQQTTLNNFYQDSCHKAIVMAHAPDHVEFSVSFQDMSPQYRNFSYHTYRLYEYLGYWYTEANPLKLTQNGKMDFKIDFSYFDRTYTVDIASPSGEARMRDMPIATMAPGALSFYQPLKAYELVANYFTGHQYQPYCSIDGTRIHTFSNRSYEYPLSRSWHVVMQDESTQRGNWHELAILSRRQQRDQQEIYISYKSESGQDLEIEIQPASGDSAYQVKVTTNTKKITDDDLTMYWDDVKEQPFLQYHTHKDGVLVINIEDDRIRAIYDGQRFVVFTQDYRNSTRGICGRMSGEQRDDYLTPEGLVDKPELYAAAYSLNEENSDPKTQELKALATQQAYYPEYKYTSILRSDPTWQEESQSSGEDQWQSETVYKSRSYDKHKGACEVRQQVQFYENHGDICITTSRVPSCQSHCRAGDYKIQHVQVTCKSKLDHDFRMYKEQIKKGQNPEVSGIPSVKQFKVPVTCQP
Summary
Subunit
Heterotetramer of two heavy and two light chains.
Keywords
Cleavage on pair of basic residues
Complete proteome
Direct protein sequencing
Glycoprotein
Phosphoprotein
Reference proteome
Secreted
Signal
Storage protein
Feature
chain Vitellogenin
Uniprot
H9J4Z5
Q27309
Q33CE5
Q9BPP5
D3K369
A5A3J2
+ More
Q59IU2 Q9BPP6 Q9GUX5 A2A0V4 Q9BPP7 Q59IU3 A0A0L7LRX8 A0A2W1BQW6 R4KZ43 A0A2A4JEQ8 S4T709 A0A0N1I5M7 W0M288 A0A0N1ICG0 A0A0S2C3S6 G3LSH9 A0A2H1WVX0 A0A1D5ARX5 A7XS62 A0A0S3G570 A0A059VBJ4 A0A385HVZ9 A0A3S2PE51 A0A212ES36 A0A125RTZ3 Q25269 Q25265 A0A3G1S403 A0A2A4IZG8 A0A2A4J1L8 A0A2H1W8V4 A0A2W1BKE7 A0A2A4JFI3 A0A2W1BMM7 A0A2W1BK92 A0A2S1XV30 A0A2H1VRD1 A0A1J1I180 E0VZ54 Q17083 E0VZ52 Q9NAW9 Q7PQM2 G3K483 Q49MF3 Q19TW4 X2G7C6 A0A2M3YWZ3 U5TQT8 A0A2M4B8F1 A0A2M4B8G2 A0A182F7T5 A0A2M3ZYE0 A0A232ETW9 W8S9N9 A0A2M3YX18 A0A2M4B8F5 W5JQP3 Q49MF2 A0A2M3ZXU1 A0A2M3ZY53 T1E7K3 B2BD67 A0A1Q1N9R4 K7J0R2 A0A336LYD9 A0A059PA41 A0A182HDP7 A0A1L8DJD8 A0A182YRT3 A0A1L8DJ24 T2B338 B1B5Z4 D7P6M0 Q6UBM1 A0A0F6PB07 A0A1S4FCM3 Q177I5 A0A0G2QNM8 A0A182GE67 D7P6M1 A0A1B6C1N7 A0A0U4XB12 A0A2M3YWZ8 A0A1B6BWN0 A0A2M4CYH9 T1IB40
Q59IU2 Q9BPP6 Q9GUX5 A2A0V4 Q9BPP7 Q59IU3 A0A0L7LRX8 A0A2W1BQW6 R4KZ43 A0A2A4JEQ8 S4T709 A0A0N1I5M7 W0M288 A0A0N1ICG0 A0A0S2C3S6 G3LSH9 A0A2H1WVX0 A0A1D5ARX5 A7XS62 A0A0S3G570 A0A059VBJ4 A0A385HVZ9 A0A3S2PE51 A0A212ES36 A0A125RTZ3 Q25269 Q25265 A0A3G1S403 A0A2A4IZG8 A0A2A4J1L8 A0A2H1W8V4 A0A2W1BKE7 A0A2A4JFI3 A0A2W1BMM7 A0A2W1BK92 A0A2S1XV30 A0A2H1VRD1 A0A1J1I180 E0VZ54 Q17083 E0VZ52 Q9NAW9 Q7PQM2 G3K483 Q49MF3 Q19TW4 X2G7C6 A0A2M3YWZ3 U5TQT8 A0A2M4B8F1 A0A2M4B8G2 A0A182F7T5 A0A2M3ZYE0 A0A232ETW9 W8S9N9 A0A2M3YX18 A0A2M4B8F5 W5JQP3 Q49MF2 A0A2M3ZXU1 A0A2M3ZY53 T1E7K3 B2BD67 A0A1Q1N9R4 K7J0R2 A0A336LYD9 A0A059PA41 A0A182HDP7 A0A1L8DJD8 A0A182YRT3 A0A1L8DJ24 T2B338 B1B5Z4 D7P6M0 Q6UBM1 A0A0F6PB07 A0A1S4FCM3 Q177I5 A0A0G2QNM8 A0A182GE67 D7P6M1 A0A1B6C1N7 A0A0U4XB12 A0A2M3YWZ8 A0A1B6BWN0 A0A2M4CYH9 T1IB40
Pubmed
EMBL
BABH01025711
D13160
D30733
AB239763
BAE47146.1
AB055845
+ More
BAB32642.2 GU361974 ADB94560.1 EF523567 ABP63663.1 AB190810 BAD91196.1 AB055844 BAB32641.1 EF683091 AB049631 ABS19573.1 BAB16412.1 AB247378 BAF45319.1 AB055843 BAB32640.1 AB190809 BAD91195.1 JTDY01000218 KOB78233.1 KZ150067 PZC74123.1 JX504706 AGL08685.1 NWSH01001771 PCG70188.1 JQ723600 AFV40972.1 KQ458842 KPJ04900.1 KF501044 AHG29547.1 KQ459890 KPJ19580.1 KT947112 ALN38805.1 JN408698 AEM75020.1 ODYU01011461 SOQ57205.1 KT599434 AOH73254.1 EU095334 ABU68426.1 KT274176 ALR35190.1 KJ540279 AHZ89334.1 MG799570 AXY55008.1 RSAL01000078 RVE48729.1 AGBW02012867 OWR44310.1 KT724958 AMD78107.1 U60186 AAB03336.1 U90756 AAC02818.1 MH553377 AXM43802.1 NWSH01004650 PCG64818.1 NWSH01003613 PCG66057.1 ODYU01007081 SOQ49535.1 PZC74124.1 PCG70190.1 PZC74126.1 PZC74125.1 MF622538 AWJ95280.1 ODYU01003950 SOQ43336.1 CVRI01000037 CRK93348.1 DS235849 EEB18660.1 AB007850 BAA22791.1 EEB18658.1 AF281078 AAF82131.1 AAAB01008880 EAA08615.3 JN113091 AEO51020.1 AY691326 AAV31932.1 DQ442990 ABD97990.1 KJ000491 AHN13887.1 GGFM01000032 MBW20783.1 KF577805 AGZ02020.1 GGFJ01000131 MBW49272.1 GGFJ01000130 MBW49271.1 GGFK01000087 MBW33408.1 NNAY01002208 OXU21803.1 KJ000061 AHM10344.1 GGFM01000045 MBW20796.1 GGFJ01000129 MBW49270.1 ADMH02000727 ETN65235.1 AY691327 AAV31933.1 GGFK01000066 MBW33387.1 GGFK01000069 MBW33390.1 GAMD01002962 JAA98628.1 EF468683 ABO70318.1 KX752432 AQM52239.1 UFQT01000300 SSX22992.1 JQ867181 AFW97644.1 JXUM01034631 KQ561030 KXJ79966.1 GFDF01007630 JAV06454.1 GFDF01007631 JAV06453.1 KF113336 AGV05363.1 AB425334 BAG12118.1 GU017909 ADH04224.1 AY373377 AAQ92366.1 KJ652904 AJI43739.1 CH477376 EAT42292.1 KC136271 AGT39945.1 JXUM01057208 KQ561944 KXJ77091.1 GU017910 ADH04225.1 GEDC01030487 GEDC01029876 JAS06811.1 JAS07422.1 LC115022 BAU36889.1 GGFM01000031 MBW20782.1 GEDC01031612 JAS05686.1 GGFL01006214 MBW70392.1 ACPB03007241
BAB32642.2 GU361974 ADB94560.1 EF523567 ABP63663.1 AB190810 BAD91196.1 AB055844 BAB32641.1 EF683091 AB049631 ABS19573.1 BAB16412.1 AB247378 BAF45319.1 AB055843 BAB32640.1 AB190809 BAD91195.1 JTDY01000218 KOB78233.1 KZ150067 PZC74123.1 JX504706 AGL08685.1 NWSH01001771 PCG70188.1 JQ723600 AFV40972.1 KQ458842 KPJ04900.1 KF501044 AHG29547.1 KQ459890 KPJ19580.1 KT947112 ALN38805.1 JN408698 AEM75020.1 ODYU01011461 SOQ57205.1 KT599434 AOH73254.1 EU095334 ABU68426.1 KT274176 ALR35190.1 KJ540279 AHZ89334.1 MG799570 AXY55008.1 RSAL01000078 RVE48729.1 AGBW02012867 OWR44310.1 KT724958 AMD78107.1 U60186 AAB03336.1 U90756 AAC02818.1 MH553377 AXM43802.1 NWSH01004650 PCG64818.1 NWSH01003613 PCG66057.1 ODYU01007081 SOQ49535.1 PZC74124.1 PCG70190.1 PZC74126.1 PZC74125.1 MF622538 AWJ95280.1 ODYU01003950 SOQ43336.1 CVRI01000037 CRK93348.1 DS235849 EEB18660.1 AB007850 BAA22791.1 EEB18658.1 AF281078 AAF82131.1 AAAB01008880 EAA08615.3 JN113091 AEO51020.1 AY691326 AAV31932.1 DQ442990 ABD97990.1 KJ000491 AHN13887.1 GGFM01000032 MBW20783.1 KF577805 AGZ02020.1 GGFJ01000131 MBW49272.1 GGFJ01000130 MBW49271.1 GGFK01000087 MBW33408.1 NNAY01002208 OXU21803.1 KJ000061 AHM10344.1 GGFM01000045 MBW20796.1 GGFJ01000129 MBW49270.1 ADMH02000727 ETN65235.1 AY691327 AAV31933.1 GGFK01000066 MBW33387.1 GGFK01000069 MBW33390.1 GAMD01002962 JAA98628.1 EF468683 ABO70318.1 KX752432 AQM52239.1 UFQT01000300 SSX22992.1 JQ867181 AFW97644.1 JXUM01034631 KQ561030 KXJ79966.1 GFDF01007630 JAV06454.1 GFDF01007631 JAV06453.1 KF113336 AGV05363.1 AB425334 BAG12118.1 GU017909 ADH04224.1 AY373377 AAQ92366.1 KJ652904 AJI43739.1 CH477376 EAT42292.1 KC136271 AGT39945.1 JXUM01057208 KQ561944 KXJ77091.1 GU017910 ADH04225.1 GEDC01030487 GEDC01029876 JAS06811.1 JAS07422.1 LC115022 BAU36889.1 GGFM01000031 MBW20782.1 GEDC01031612 JAS05686.1 GGFL01006214 MBW70392.1 ACPB03007241
Proteomes
Interpro
Gene 3D
ProteinModelPortal
H9J4Z5
Q27309
Q33CE5
Q9BPP5
D3K369
A5A3J2
+ More
Q59IU2 Q9BPP6 Q9GUX5 A2A0V4 Q9BPP7 Q59IU3 A0A0L7LRX8 A0A2W1BQW6 R4KZ43 A0A2A4JEQ8 S4T709 A0A0N1I5M7 W0M288 A0A0N1ICG0 A0A0S2C3S6 G3LSH9 A0A2H1WVX0 A0A1D5ARX5 A7XS62 A0A0S3G570 A0A059VBJ4 A0A385HVZ9 A0A3S2PE51 A0A212ES36 A0A125RTZ3 Q25269 Q25265 A0A3G1S403 A0A2A4IZG8 A0A2A4J1L8 A0A2H1W8V4 A0A2W1BKE7 A0A2A4JFI3 A0A2W1BMM7 A0A2W1BK92 A0A2S1XV30 A0A2H1VRD1 A0A1J1I180 E0VZ54 Q17083 E0VZ52 Q9NAW9 Q7PQM2 G3K483 Q49MF3 Q19TW4 X2G7C6 A0A2M3YWZ3 U5TQT8 A0A2M4B8F1 A0A2M4B8G2 A0A182F7T5 A0A2M3ZYE0 A0A232ETW9 W8S9N9 A0A2M3YX18 A0A2M4B8F5 W5JQP3 Q49MF2 A0A2M3ZXU1 A0A2M3ZY53 T1E7K3 B2BD67 A0A1Q1N9R4 K7J0R2 A0A336LYD9 A0A059PA41 A0A182HDP7 A0A1L8DJD8 A0A182YRT3 A0A1L8DJ24 T2B338 B1B5Z4 D7P6M0 Q6UBM1 A0A0F6PB07 A0A1S4FCM3 Q177I5 A0A0G2QNM8 A0A182GE67 D7P6M1 A0A1B6C1N7 A0A0U4XB12 A0A2M3YWZ8 A0A1B6BWN0 A0A2M4CYH9 T1IB40
Q59IU2 Q9BPP6 Q9GUX5 A2A0V4 Q9BPP7 Q59IU3 A0A0L7LRX8 A0A2W1BQW6 R4KZ43 A0A2A4JEQ8 S4T709 A0A0N1I5M7 W0M288 A0A0N1ICG0 A0A0S2C3S6 G3LSH9 A0A2H1WVX0 A0A1D5ARX5 A7XS62 A0A0S3G570 A0A059VBJ4 A0A385HVZ9 A0A3S2PE51 A0A212ES36 A0A125RTZ3 Q25269 Q25265 A0A3G1S403 A0A2A4IZG8 A0A2A4J1L8 A0A2H1W8V4 A0A2W1BKE7 A0A2A4JFI3 A0A2W1BMM7 A0A2W1BK92 A0A2S1XV30 A0A2H1VRD1 A0A1J1I180 E0VZ54 Q17083 E0VZ52 Q9NAW9 Q7PQM2 G3K483 Q49MF3 Q19TW4 X2G7C6 A0A2M3YWZ3 U5TQT8 A0A2M4B8F1 A0A2M4B8G2 A0A182F7T5 A0A2M3ZYE0 A0A232ETW9 W8S9N9 A0A2M3YX18 A0A2M4B8F5 W5JQP3 Q49MF2 A0A2M3ZXU1 A0A2M3ZY53 T1E7K3 B2BD67 A0A1Q1N9R4 K7J0R2 A0A336LYD9 A0A059PA41 A0A182HDP7 A0A1L8DJD8 A0A182YRT3 A0A1L8DJ24 T2B338 B1B5Z4 D7P6M0 Q6UBM1 A0A0F6PB07 A0A1S4FCM3 Q177I5 A0A0G2QNM8 A0A182GE67 D7P6M1 A0A1B6C1N7 A0A0U4XB12 A0A2M3YWZ8 A0A1B6BWN0 A0A2M4CYH9 T1IB40
PDB
1LSH
E-value=2.1116e-05,
Score=121
Ontologies
Topology
Subcellular location
Secreted
SignalP
Position: 1 - 15,
Likelihood: 0.991085
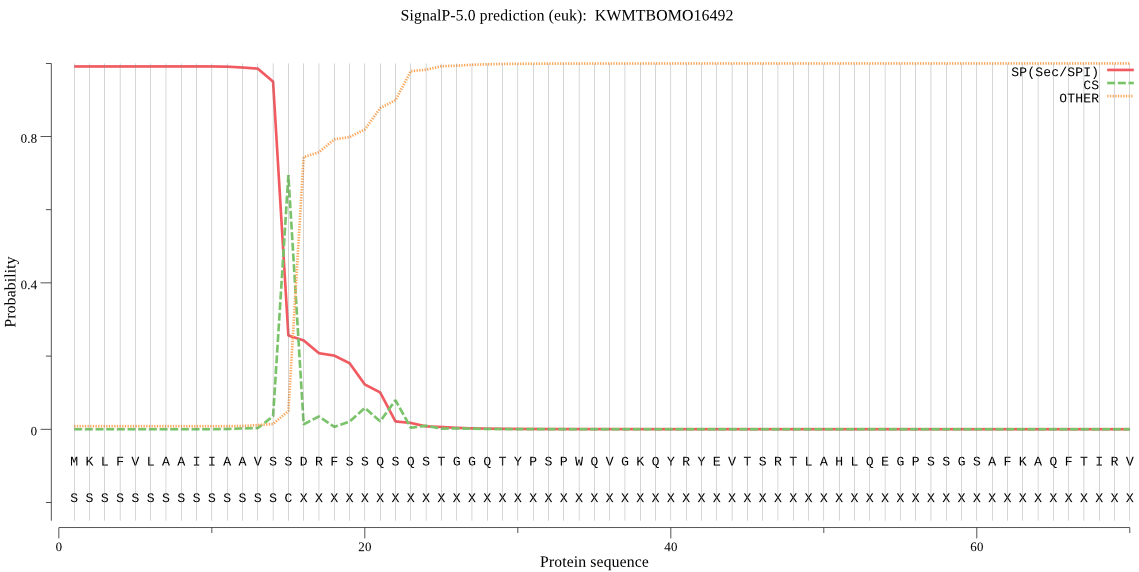
Length:
1782
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00167
Exp number, first 60 AAs:
0.00144
Total prob of N-in:
0.00008
outside
1 - 1782
Population Genetic Test Statistics
Pi
11.298257
Theta
17.506995
Tajima's D
-1.073127
CLR
0
CSRT
0.125943702814859
Interpretation
Uncertain
