Pre Gene Modal
BGIBMGA013860
Annotation
UDP-glycosyltransferase_UGT33D5_precursor_[Bombyx_mori]
Location in the cell
PlasmaMembrane Reliability : 2.227
Sequence
CDS
ATGATAGTGTATTTTGTGGCAATTTTACAAGAAATACAAGCGGCTCGGATCCTAGGCGTGTTCCCTATACCCTCAATCAGTCATCAAGTAGTCTTCCGTCGTCTGACCTTAGAGCTTCACAAAAGGGGTCATGAACTAGTGGTTGTGACCACTGACCCTGTGTTCAAAACAGGACAAGCACCTGCGAATTACACAGAAATAGATGTACATGACGTTTCTTATTCAGCCTGGAGAGACGGCTTCATGAAAACCTCTAAAGGAAACGCTAATGATCTTTACGAACAAATCGCCATTGCTTTAAACTTGGGCATAGATTTGCTTGAACTACAAATGAAAAACAAAGAAGTCCAAGCGTTGCTAAGGAAGAGAAAAGAGAAGAAGTTTGATTTGCTCCTTCTAGAAGCATGCATCAGATCAACGATGGTCTTGACGCATGTGTTTGATGCACCGGCAATCCTCATCAGTTCATTTGGAGGTGTAGAATTCATGTCGAAAATGATGGGTGTCCCAACCCATCCTATTTTATATCCACCACCGTTGCATCAAAGACTTTATAATTTAACGTTCTTGGAGAAAATCGGTGAGATATACACCCATTACTATATGGAGTATCTATATTGGCGATCTGAACCCCAGGAGAATCAAATGGCAAAAAGGCTTTTCGGCCCGAACACTCCAACTATCAGAGAAACCCAGAAGAACGTACAGATGGCACTCCTGAATGTACACGCGATTTGGGAGGAAAATCGTCCTGTCCCTCCGAATGTTATATACATTGGAGGTATTCATCAGAATCCAGAAAAAAACTTGCCCAAGGATTTGAAGGAATATTTAGACTCTTCAAAACATGGCGTTATATACATTAGTTTCGGCACCAACGTGGAACCATCGTTACTACCTCCAGAACGGATTCAACTCTTCATTAAAGTGTTCTCCGAACTGCCTTACGATGTGCTCTGGAAATGGGACAAAGATGAACTGCCCGGGAGCTCAAAGAACATCAGAATAGCAAAGTGGCTGCCTCAATCAGATTTACTTAGACACCCAAAGATCAAGGCGTTCATAACTCAAGGGGGTCTGCAATCGACCGAAGAAGCGATCACAGCTGGAGTGCCACTGATTGGAATGCCTATGTTGATGGACCAGTGGTACAATGTCGAGAAATACGTTCGACACAACATCGGTTTGAGACTCGATTTGGGTTCCGTTACAGAGGAATCTCTCAGAAATGCCATCAACACTATCACTGGTGACGAAAGCTACCGTCAGAATATAGCTCGTCTCCGTAGTCAGGTCTACGATCAGCCCCAGAGCTCAGTAGACCGCGCCGTGTGGTGGACTGAGCACGTGCTGCGCCACGGAGGAGCCACTCATCTCCGAGCCGCAGGAGCCTTGAAATCTTGGACTGAATATTTTGAACTAAATTTAATTGCAGTTTTACTAGTCACTTTCCTAATTGTTATTGCATTAATTGCAAGCAACGGTGTGTTCCAAAGAGTACTTGTGATGACAAATATGTTTTCTTTAAACAAGATAGTGCTTATTGTGACAATTATACACGGAACCCAAGCGGCTCGGATCCTAGGCGTGTTCCCTACACCCTCAATCAGTCATCAAGTAGTCTTCCGTCGTCTGACCTTAGAGCTTCACAAAAGGGGTCATGAACTAGTGGTTGTGACCACTGACCCGGTGTTCAAAACAGGACAAGCACCTGCGAATTACACAGAAATAGACGTACACGACGTTTCTTATTCAGCCTGGAGAGACGGCTTCATGAAAACCTCTAGGGGAAGCGCCAATGATATTTACGAACAAATTTCCAGAATGTTGAACTTAAGAATAGATTTGTTTGAATTGCAGATGAAAAATAAAGAAGTGCAGTCGCTGCTGAGTAAGAGAAAAGAGAGCAAGTTTGATTTGCTCCTTCTAGAAGCATGCATCAGACCAACGATGGTCTTGACGCATGTGTTTGATGCACCAGCAATCCTCATCAGTTCATTTGGGGGTGTAGAATTCGTCTTGAAAATGATTGGTGTCCCAACCCATCCTATATTATATCCACCTTCCTTGCACCAACGAATTTATAACTTAACATTTTCGGAGAAAATCCGAGAGGTTTACACTCATTACTATTTGGAATATCTATATTGGCGATCTGAACCCCAGGAGAATCAAATGGCAAAAAGGCTTTTCGGCCCGAACACTCCAACTATCAGAGAGACACAGAAGAACGTGCAGATGGCACTCCTGAATGTACACGCGATTTGGGAGGAAAATCGTCCTGTCCCTCCGAATGTTATATACATTGGAGGTATTCATCAGAATCCCGAAAAGGAATTGCCCAAGGATTTGAAGGAATATTTAGACTCTTCAAAACATGGCGTTATATACATTAGTTTCGGCACCAACGTGGAACCATCGCTACTACCTCCAGAACGGATTCAAATCCTCGTCAAAGTATTTTCCAAGCTGCCTTACGATGTGCTCTGGAAATGGGACAAAGATGAACTGCCCGGGAGCTCAAAGAACATCAGAATAGCAAAATGGCTGCCTCAATCAGATTTACTGAGACACCCAAAGATCAAGGCATTCATAACTCAAGGGGGTCTGCAATCGACGGAAGAAGCAATCACAGCTGGAGTGCCACTGATTGGAATGCCTATGTTGATGGACCAATGGTACAATGTCGAGAAATACGTTCGACACAACATCGGTTTGAGACTCGATTTGGGTTCCGTTACAGAGGAATCTCTCAGGAACGCCATCGACACTATCACTGGTGATGAAAGCTACCGTCAGAACATAGCTCGTCTCCGTAGTCAGGTCTACGATCAGCCCCAGAGCTCAGTTGACCGCGCCGTGTGGTGGACTGAGCACGTGCTGCGCCACGGAGGAGCCACTCATCTCCGAGCTGCAGGAGCCCTGAAATCTTGGACTGAATATTTTGAACTAAATTTAATTGCAGTATTATTAGTTACTTTCCTAATTATAATTACATTCATAGCAAAATTAATATCTTCGTTTGTGAGGTCATTGAAAATGTATTTCAATTATAATGATAAAGTAAAATTACATTGA
Protein
MIVYFVAILQEIQAARILGVFPIPSISHQVVFRRLTLELHKRGHELVVVTTDPVFKTGQAPANYTEIDVHDVSYSAWRDGFMKTSKGNANDLYEQIAIALNLGIDLLELQMKNKEVQALLRKRKEKKFDLLLLEACIRSTMVLTHVFDAPAILISSFGGVEFMSKMMGVPTHPILYPPPLHQRLYNLTFLEKIGEIYTHYYMEYLYWRSEPQENQMAKRLFGPNTPTIRETQKNVQMALLNVHAIWEENRPVPPNVIYIGGIHQNPEKNLPKDLKEYLDSSKHGVIYISFGTNVEPSLLPPERIQLFIKVFSELPYDVLWKWDKDELPGSSKNIRIAKWLPQSDLLRHPKIKAFITQGGLQSTEEAITAGVPLIGMPMLMDQWYNVEKYVRHNIGLRLDLGSVTEESLRNAINTITGDESYRQNIARLRSQVYDQPQSSVDRAVWWTEHVLRHGGATHLRAAGALKSWTEYFELNLIAVLLVTFLIVIALIASNGVFQRVLVMTNMFSLNKIVLIVTIIHGTQAARILGVFPTPSISHQVVFRRLTLELHKRGHELVVVTTDPVFKTGQAPANYTEIDVHDVSYSAWRDGFMKTSRGSANDIYEQISRMLNLRIDLFELQMKNKEVQSLLSKRKESKFDLLLLEACIRPTMVLTHVFDAPAILISSFGGVEFVLKMIGVPTHPILYPPSLHQRIYNLTFSEKIREVYTHYYLEYLYWRSEPQENQMAKRLFGPNTPTIRETQKNVQMALLNVHAIWEENRPVPPNVIYIGGIHQNPEKELPKDLKEYLDSSKHGVIYISFGTNVEPSLLPPERIQILVKVFSKLPYDVLWKWDKDELPGSSKNIRIAKWLPQSDLLRHPKIKAFITQGGLQSTEEAITAGVPLIGMPMLMDQWYNVEKYVRHNIGLRLDLGSVTEESLRNAIDTITGDESYRQNIARLRSQVYDQPQSSVDRAVWWTEHVLRHGGATHLRAAGALKSWTEYFELNLIAVLLVTFLIIITFIAKLISSFVRSLKMYFNYNDKVKLH
Summary
Similarity
Belongs to the UDP-glycosyltransferase family.
Belongs to the peptidase S1 family.
Belongs to the peptidase S1 family.
Uniprot
EMBL
BABH01029273
BABH01029274
ODYU01004485
SOQ44409.1
ODYU01004546
SOQ44548.1
+ More
KQ459547 KPJ00041.1 ODYU01010294 SOQ55304.1 ODYU01005543 SOQ46493.1 KQ459680 KPJ20473.1 BABH01029238 BABH01029239 BABH01029240 KPJ00055.1 BABH01029259 KPJ00049.1 AGBW02010679 OWR48052.1 JTDY01001195 KOB74579.1 AGBW02009962 OWR49474.1 JTDY01001352 KOB74102.1 ODYU01001030 SOQ36683.1 JTDY01003324 KOB69741.1 CVRI01000067 CRL06668.1
KQ459547 KPJ00041.1 ODYU01010294 SOQ55304.1 ODYU01005543 SOQ46493.1 KQ459680 KPJ20473.1 BABH01029238 BABH01029239 BABH01029240 KPJ00055.1 BABH01029259 KPJ00049.1 AGBW02010679 OWR48052.1 JTDY01001195 KOB74579.1 AGBW02009962 OWR49474.1 JTDY01001352 KOB74102.1 ODYU01001030 SOQ36683.1 JTDY01003324 KOB69741.1 CVRI01000067 CRL06668.1
Proteomes
Interpro
SUPFAM
SSF50494
SSF50494
CDD
ProteinModelPortal
PDB
2O6L
E-value=3.92996e-27,
Score=306
Ontologies
PATHWAY
00040
Pentose and glucuronate interconversions - Bombyx mori (domestic silkworm)
00053 Ascorbate and aldarate metabolism - Bombyx mori (domestic silkworm)
00830 Retinol metabolism - Bombyx mori (domestic silkworm)
00860 Porphyrin and chlorophyll metabolism - Bombyx mori (domestic silkworm)
00980 Metabolism of xenobiotics by cytochrome P450 - Bombyx mori (domestic silkworm)
00982 Drug metabolism - cytochrome P450 - Bombyx mori (domestic silkworm)
00983 Drug metabolism - other enzymes - Bombyx mori (domestic silkworm)
01100 Metabolic pathways - Bombyx mori (domestic silkworm)
00053 Ascorbate and aldarate metabolism - Bombyx mori (domestic silkworm)
00830 Retinol metabolism - Bombyx mori (domestic silkworm)
00860 Porphyrin and chlorophyll metabolism - Bombyx mori (domestic silkworm)
00980 Metabolism of xenobiotics by cytochrome P450 - Bombyx mori (domestic silkworm)
00982 Drug metabolism - cytochrome P450 - Bombyx mori (domestic silkworm)
00983 Drug metabolism - other enzymes - Bombyx mori (domestic silkworm)
01100 Metabolic pathways - Bombyx mori (domestic silkworm)
GO
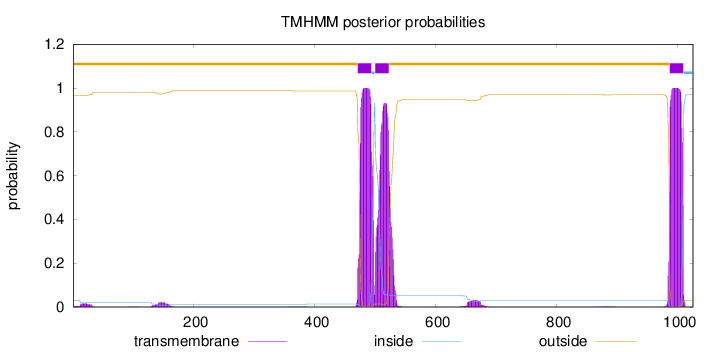
Topology
Length:
1025
Number of predicted TMHs:
3
Exp number of AAs in TMHs:
66.6076500000001
Exp number, first 60 AAs:
0.3199
Total prob of N-in:
0.03257
outside
1 - 470
TMhelix
471 - 493
inside
494 - 499
TMhelix
500 - 522
outside
523 - 986
TMhelix
987 - 1009
inside
1010 - 1025
Population Genetic Test Statistics
Pi
188.42675
Theta
175.766254
Tajima's D
1.111697
CLR
0.525218
CSRT
0.688965551722414
Interpretation
Uncertain
