Gene
KWMTBOMO16420
Pre Gene Modal
BGIBMGA013830
Annotation
UDP-glucosyltransferase_precursor_[Bombyx_mori]
Full name
UDP-glucuronosyltransferase
Location in the cell
Cytoplasmic Reliability : 1.782
Sequence
CDS
ATGCTCTTAATCACAATTCTGCAAGGAACTAAAGCGGCTAGAATTTTAGGAGTGTTCCCTACACCATCAATCAGTCATCAAGTAGTCTTCCGACGTCTCACCTTAGAGCTTCACAAAAGAGGTCACGAACTAGTAATCGTCACAACTGACCCTATGTACCAGAAAGGCGAAGCCCCTCAAAACTATACAGAAATCGACGTGCATGACATTTCGTACACAACTTGGAGAAATGACTTCATGAAACTTTCCAGAGGAAGCTCCGATGATCTTTTCGAACAATCAGCTGTTATATTGGAATTAACTACGAATTTGTTTGAAATGCAACTGAAAAGCAAAGAAGTTCAAGCACTGATCAAAGTTAAAGATGCAAAGAAGTTTGATTTGCTCCTTCTAGAAGCATGCATTAGACCAGCAATTATTTTAACGCACGTTTTCGATGCACCAGCAATCCTTGTCAGTTCATTTGGAGGTGTGGAATATGTGTTTCGTATACTTGGCGTCCCAACGCATCCTGTACTATACCCGCCTCCGCTGCACCAAAGAATTTTTAATTTGACATTTTGGGAGAAAACGCACGAAATTTTTACTCATTACTATTTGGAATATCTTTTTTGGAAGGCAGAATATAAGGTCGATGAAATGGTAAAAAGGATTTTCGGACCCAGCACTCCAACTGTGAGAGATACCTATAAAAATGTAGAGATGATACTCTTGAATGCGTATGCTGTTTGGGAGAATAATAGACCTGTCCCTCCTAATGTCATATACGTTGGTGGCTTACATCAGAAACCAGAGAAGGATTTACCCGGGGATTTGAAAGAATATTTAGATTCTTCAAAACACGGCGTTGTATACATCAGTTTCGGCACAAACGTGGAACCATCGCTGCTGCCTCCAGAACGGATTCAACTCCTCATCAAAGTATTCTCCGAGCTGCCTTACGATGTACTCTGGAAATGGGACCAAGATGAATTGCCGGGCAAATCGGAAAACATCAAAATAGCAAAGTGGCTGCCACAATCAGATTTACTAAGGCATCCAAAGATCAAGGTGTTTATAACTCAAGGAGGTCTGCAATCGACCGAAGAGGCAATCACAGCTGGAGTGCCACTGATAGGGATACCTATGTTGATGGATCAATGGTACAATGTAGAGAAATACGTTCAGCTAAACATTGGCTTGAAACTCGATTTGGGCTCCATTACAGAGGATTCTTTCAGAAATGCTATCAACACAGTTACTGGTGATGAAAGCTATCGTCAGAACGTAGCTCGTCTCCGTAGCCAGGTCTTCGACCAGCCCCAGGGCCCTCTTGAGCGCGCCGTGTGGTGGACTGAGCACGTACTGCGCCACGGAGGGGCCACTCATCTCCGAGCCGCAGGAGCCCTGAAATCTTGGACTGAATATTTTGAACTAAATTTAATTGCAGTATTATTATTATAA
Protein
MLLITILQGTKAARILGVFPTPSISHQVVFRRLTLELHKRGHELVIVTTDPMYQKGEAPQNYTEIDVHDISYTTWRNDFMKLSRGSSDDLFEQSAVILELTTNLFEMQLKSKEVQALIKVKDAKKFDLLLLEACIRPAIILTHVFDAPAILVSSFGGVEYVFRILGVPTHPVLYPPPLHQRIFNLTFWEKTHEIFTHYYLEYLFWKAEYKVDEMVKRIFGPSTPTVRDTYKNVEMILLNAYAVWENNRPVPPNVIYVGGLHQKPEKDLPGDLKEYLDSSKHGVVYISFGTNVEPSLLPPERIQLLIKVFSELPYDVLWKWDQDELPGKSENIKIAKWLPQSDLLRHPKIKVFITQGGLQSTEEAITAGVPLIGIPMLMDQWYNVEKYVQLNIGLKLDLGSITEDSFRNAINTVTGDESYRQNVARLRSQVFDQPQGPLERAVWWTEHVLRHGGATHLRAAGALKSWTEYFELNLIAVLLL
Summary
Catalytic Activity
glucuronate acceptor + UDP-alpha-D-glucuronate = acceptor beta-D-glucuronoside + H(+) + UDP
Similarity
Belongs to the UDP-glycosyltransferase family.
Feature
chain UDP-glucuronosyltransferase
Uniprot
F1DG68
G9LPS2
H9JWB6
G9LPS6
Q1HPX7
H9JWE6
+ More
G9LPS5 G9LPS9 B7TZA3 G9LPS4 G9LPS8 G9LPS7 H9JWB9 H9JWE5 G9LPS3 H9JWB7 A0A2H1W7S2 A0A2W1BL20 G9LPP3 A0A191T2F8 A0A2W1BEB8 A0A2H1VU94 M4M1W9 G9LPN9 A0A2A4IVU8 A0A2W1BE55 G9LPP5 A0A2H1WTF5 A0A191T1K8 A0A191T2J2 A0A024AG00 A0A2W1BHP8 G9LPN7 A0A2W1B475 A0A024AH14 A0A2W1BA70 A0A2W1B3V4 A0A2W1B485 A0A191T1L2 A0A191T1L8 A0A191T2J5 A0A2H1VUS4 G9LPN6 A0A2H4WB73 G9LPN3 A0A1L8D6T8 G9LPP0 A0A2H1W0F9 A0A2W1BBH1 A0A0N1IB27 G9LPN2 A0A0L7LF59 A0A2W1BEA5 A0A2W1BCY8 A0A2W1B8D3 G9LPP1 A0A191T2G3 G9LPN8 A0A2W1B675 A0A2H4WB75 A0A2W1B3X2 G9LPP7 A0A2W1B7N5 A0A2W1B8G2 G9LPP2 G9LPT1 A0A2W1BQP4 A0A2A4JWQ0 A0A024AGF9 G9LPP4 A0A0N1I2R8 A0A191T1L3 A0A024AGL3 G9LPN5 A0A2W1B9W6 A0A140EA35 A0A0N0PE43 A0A2W1BDA5 A0A2W1B6A2 A0A2I6RJL8 A0A2W1BHY3 A0A191T1M4 G9LPN4 A0A191T1K4 A0A0N1ICS6 A0A2H1VUM4 A0A2H1WSA7 A0A191T2G1 A0A2H1V761 A0A191T1N0 A0A0N0P9C8 A0A2W1BAD4 G9LPP6 A0A1L8D6R8 A0A0N1IQG9 A0A0N0P9D1 A0A212F6X6 D2JLK8 A0A0N1IE25 G9LPT0
G9LPS5 G9LPS9 B7TZA3 G9LPS4 G9LPS8 G9LPS7 H9JWB9 H9JWE5 G9LPS3 H9JWB7 A0A2H1W7S2 A0A2W1BL20 G9LPP3 A0A191T2F8 A0A2W1BEB8 A0A2H1VU94 M4M1W9 G9LPN9 A0A2A4IVU8 A0A2W1BE55 G9LPP5 A0A2H1WTF5 A0A191T1K8 A0A191T2J2 A0A024AG00 A0A2W1BHP8 G9LPN7 A0A2W1B475 A0A024AH14 A0A2W1BA70 A0A2W1B3V4 A0A2W1B485 A0A191T1L2 A0A191T1L8 A0A191T2J5 A0A2H1VUS4 G9LPN6 A0A2H4WB73 G9LPN3 A0A1L8D6T8 G9LPP0 A0A2H1W0F9 A0A2W1BBH1 A0A0N1IB27 G9LPN2 A0A0L7LF59 A0A2W1BEA5 A0A2W1BCY8 A0A2W1B8D3 G9LPP1 A0A191T2G3 G9LPN8 A0A2W1B675 A0A2H4WB75 A0A2W1B3X2 G9LPP7 A0A2W1B7N5 A0A2W1B8G2 G9LPP2 G9LPT1 A0A2W1BQP4 A0A2A4JWQ0 A0A024AGF9 G9LPP4 A0A0N1I2R8 A0A191T1L3 A0A024AGL3 G9LPN5 A0A2W1B9W6 A0A140EA35 A0A0N0PE43 A0A2W1BDA5 A0A2W1B6A2 A0A2I6RJL8 A0A2W1BHY3 A0A191T1M4 G9LPN4 A0A191T1K4 A0A0N1ICS6 A0A2H1VUM4 A0A2H1WSA7 A0A191T2G1 A0A2H1V761 A0A191T1N0 A0A0N0P9C8 A0A2W1BAD4 G9LPP6 A0A1L8D6R8 A0A0N1IQG9 A0A0N0P9D1 A0A212F6X6 D2JLK8 A0A0N1IE25 G9LPT0
EC Number
2.4.1.17
Pubmed
EMBL
HQ833816
ADY17534.1
JQ070229
AEW43145.1
BABH01029271
JQ070233
+ More
AEW43149.1 DQ443275 ABF51364.1 BABH01029273 BABH01029274 JQ070232 AEW43148.1 JQ070236 AEW43152.1 FJ418685 ACJ48963.1 JQ070231 AEW43147.1 JQ070235 AEW43151.1 JQ070234 AEW43150.1 BABH01029259 BABH01029260 BABH01029272 JQ070230 AEW43146.1 BABH01029269 BABH01029270 ODYU01006890 SOQ49139.1 KZ150193 PZC72373.1 JQ070200 AEW43116.1 KU680280 ANI21986.1 KZ150360 PZC71200.1 ODYU01004485 SOQ44409.1 JQ747500 AGG36457.1 JQ070196 AEW43112.1 NWSH01005967 PCG63789.1 PZC71196.1 JQ070202 AEW43118.1 ODYU01010933 SOQ56350.1 KU680282 ANI21988.1 KU680284 ANI21990.1 KF777109 AHY99680.1 PZC71193.1 JQ070194 AEW43110.1 PZC71189.1 KF777111 AHY99682.1 PZC71195.1 PZC71192.1 PZC71199.1 KU680287 ANI21993.1 KU680281 ANI21987.1 KU680289 ANI21995.1 ODYU01004546 SOQ44548.1 JQ070193 AEW43109.1 KY202933 AUC64277.1 JQ070190 AEW43106.1 GEYN01000079 JAV02050.1 JQ070197 AEW43113.1 ODYU01005543 SOQ46493.1 PZC71194.1 KQ459547 KPJ00043.1 JQ070189 AEW43105.1 JTDY01001352 KOB74102.1 PZC71190.1 PZC71197.1 PZC71191.1 JQ070198 AEW43114.1 KU680290 ANI21996.1 JQ070195 AEW43111.1 KZ150704 PZC70375.1 KY202945 AUC64289.1 PZC71202.1 JQ070204 AEW43120.1 PZC70374.1 PZC71201.1 JQ070199 AEW43115.1 JQ070238 AEW43154.1 KZ149974 PZC75964.1 NWSH01000435 PCG76447.1 KF777110 AHY99681.1 JQ070201 AEW43117.1 KPJ00048.1 KU680288 ANI21994.1 KF777112 AHY99683.1 JQ070192 AEW43108.1 PZC72369.1 KU214506 AMK97472.1 KQ459937 KPJ19195.1 PZC72371.1 PZC71198.1 KY304474 AUN86405.1 PZC72370.1 KU680286 ANI21992.1 JQ070191 AEW43107.1 KU680283 ANI21989.1 KPJ00041.1 SOQ44549.1 ODYU01010294 SOQ55304.1 KU680285 ANI21991.1 ODYU01001030 SOQ36683.1 KU680291 ANI21997.1 KPJ00042.1 PZC72372.1 JQ070203 AEW43119.1 GEYN01000099 JAV02030.1 KQ459680 KPJ20473.1 KPJ00052.1 AGBW02009962 OWR49474.1 GQ915324 ACZ97418.2 KQ460636 KPJ13397.1 JQ070237 AEW43153.1
AEW43149.1 DQ443275 ABF51364.1 BABH01029273 BABH01029274 JQ070232 AEW43148.1 JQ070236 AEW43152.1 FJ418685 ACJ48963.1 JQ070231 AEW43147.1 JQ070235 AEW43151.1 JQ070234 AEW43150.1 BABH01029259 BABH01029260 BABH01029272 JQ070230 AEW43146.1 BABH01029269 BABH01029270 ODYU01006890 SOQ49139.1 KZ150193 PZC72373.1 JQ070200 AEW43116.1 KU680280 ANI21986.1 KZ150360 PZC71200.1 ODYU01004485 SOQ44409.1 JQ747500 AGG36457.1 JQ070196 AEW43112.1 NWSH01005967 PCG63789.1 PZC71196.1 JQ070202 AEW43118.1 ODYU01010933 SOQ56350.1 KU680282 ANI21988.1 KU680284 ANI21990.1 KF777109 AHY99680.1 PZC71193.1 JQ070194 AEW43110.1 PZC71189.1 KF777111 AHY99682.1 PZC71195.1 PZC71192.1 PZC71199.1 KU680287 ANI21993.1 KU680281 ANI21987.1 KU680289 ANI21995.1 ODYU01004546 SOQ44548.1 JQ070193 AEW43109.1 KY202933 AUC64277.1 JQ070190 AEW43106.1 GEYN01000079 JAV02050.1 JQ070197 AEW43113.1 ODYU01005543 SOQ46493.1 PZC71194.1 KQ459547 KPJ00043.1 JQ070189 AEW43105.1 JTDY01001352 KOB74102.1 PZC71190.1 PZC71197.1 PZC71191.1 JQ070198 AEW43114.1 KU680290 ANI21996.1 JQ070195 AEW43111.1 KZ150704 PZC70375.1 KY202945 AUC64289.1 PZC71202.1 JQ070204 AEW43120.1 PZC70374.1 PZC71201.1 JQ070199 AEW43115.1 JQ070238 AEW43154.1 KZ149974 PZC75964.1 NWSH01000435 PCG76447.1 KF777110 AHY99681.1 JQ070201 AEW43117.1 KPJ00048.1 KU680288 ANI21994.1 KF777112 AHY99683.1 JQ070192 AEW43108.1 PZC72369.1 KU214506 AMK97472.1 KQ459937 KPJ19195.1 PZC72371.1 PZC71198.1 KY304474 AUN86405.1 PZC72370.1 KU680286 ANI21992.1 JQ070191 AEW43107.1 KU680283 ANI21989.1 KPJ00041.1 SOQ44549.1 ODYU01010294 SOQ55304.1 KU680285 ANI21991.1 ODYU01001030 SOQ36683.1 KU680291 ANI21997.1 KPJ00042.1 PZC72372.1 JQ070203 AEW43119.1 GEYN01000099 JAV02030.1 KQ459680 KPJ20473.1 KPJ00052.1 AGBW02009962 OWR49474.1 GQ915324 ACZ97418.2 KQ460636 KPJ13397.1 JQ070237 AEW43153.1
Proteomes
PRIDE
Pfam
PF00201 UDPGT
ProteinModelPortal
F1DG68
G9LPS2
H9JWB6
G9LPS6
Q1HPX7
H9JWE6
+ More
G9LPS5 G9LPS9 B7TZA3 G9LPS4 G9LPS8 G9LPS7 H9JWB9 H9JWE5 G9LPS3 H9JWB7 A0A2H1W7S2 A0A2W1BL20 G9LPP3 A0A191T2F8 A0A2W1BEB8 A0A2H1VU94 M4M1W9 G9LPN9 A0A2A4IVU8 A0A2W1BE55 G9LPP5 A0A2H1WTF5 A0A191T1K8 A0A191T2J2 A0A024AG00 A0A2W1BHP8 G9LPN7 A0A2W1B475 A0A024AH14 A0A2W1BA70 A0A2W1B3V4 A0A2W1B485 A0A191T1L2 A0A191T1L8 A0A191T2J5 A0A2H1VUS4 G9LPN6 A0A2H4WB73 G9LPN3 A0A1L8D6T8 G9LPP0 A0A2H1W0F9 A0A2W1BBH1 A0A0N1IB27 G9LPN2 A0A0L7LF59 A0A2W1BEA5 A0A2W1BCY8 A0A2W1B8D3 G9LPP1 A0A191T2G3 G9LPN8 A0A2W1B675 A0A2H4WB75 A0A2W1B3X2 G9LPP7 A0A2W1B7N5 A0A2W1B8G2 G9LPP2 G9LPT1 A0A2W1BQP4 A0A2A4JWQ0 A0A024AGF9 G9LPP4 A0A0N1I2R8 A0A191T1L3 A0A024AGL3 G9LPN5 A0A2W1B9W6 A0A140EA35 A0A0N0PE43 A0A2W1BDA5 A0A2W1B6A2 A0A2I6RJL8 A0A2W1BHY3 A0A191T1M4 G9LPN4 A0A191T1K4 A0A0N1ICS6 A0A2H1VUM4 A0A2H1WSA7 A0A191T2G1 A0A2H1V761 A0A191T1N0 A0A0N0P9C8 A0A2W1BAD4 G9LPP6 A0A1L8D6R8 A0A0N1IQG9 A0A0N0P9D1 A0A212F6X6 D2JLK8 A0A0N1IE25 G9LPT0
G9LPS5 G9LPS9 B7TZA3 G9LPS4 G9LPS8 G9LPS7 H9JWB9 H9JWE5 G9LPS3 H9JWB7 A0A2H1W7S2 A0A2W1BL20 G9LPP3 A0A191T2F8 A0A2W1BEB8 A0A2H1VU94 M4M1W9 G9LPN9 A0A2A4IVU8 A0A2W1BE55 G9LPP5 A0A2H1WTF5 A0A191T1K8 A0A191T2J2 A0A024AG00 A0A2W1BHP8 G9LPN7 A0A2W1B475 A0A024AH14 A0A2W1BA70 A0A2W1B3V4 A0A2W1B485 A0A191T1L2 A0A191T1L8 A0A191T2J5 A0A2H1VUS4 G9LPN6 A0A2H4WB73 G9LPN3 A0A1L8D6T8 G9LPP0 A0A2H1W0F9 A0A2W1BBH1 A0A0N1IB27 G9LPN2 A0A0L7LF59 A0A2W1BEA5 A0A2W1BCY8 A0A2W1B8D3 G9LPP1 A0A191T2G3 G9LPN8 A0A2W1B675 A0A2H4WB75 A0A2W1B3X2 G9LPP7 A0A2W1B7N5 A0A2W1B8G2 G9LPP2 G9LPT1 A0A2W1BQP4 A0A2A4JWQ0 A0A024AGF9 G9LPP4 A0A0N1I2R8 A0A191T1L3 A0A024AGL3 G9LPN5 A0A2W1B9W6 A0A140EA35 A0A0N0PE43 A0A2W1BDA5 A0A2W1B6A2 A0A2I6RJL8 A0A2W1BHY3 A0A191T1M4 G9LPN4 A0A191T1K4 A0A0N1ICS6 A0A2H1VUM4 A0A2H1WSA7 A0A191T2G1 A0A2H1V761 A0A191T1N0 A0A0N0P9C8 A0A2W1BAD4 G9LPP6 A0A1L8D6R8 A0A0N1IQG9 A0A0N0P9D1 A0A212F6X6 D2JLK8 A0A0N1IE25 G9LPT0
PDB
2O6L
E-value=5.82856e-25,
Score=284
Ontologies
PATHWAY
00040
Pentose and glucuronate interconversions - Bombyx mori (domestic silkworm)
00053 Ascorbate and aldarate metabolism - Bombyx mori (domestic silkworm)
00830 Retinol metabolism - Bombyx mori (domestic silkworm)
00860 Porphyrin and chlorophyll metabolism - Bombyx mori (domestic silkworm)
00980 Metabolism of xenobiotics by cytochrome P450 - Bombyx mori (domestic silkworm)
00982 Drug metabolism - cytochrome P450 - Bombyx mori (domestic silkworm)
00983 Drug metabolism - other enzymes - Bombyx mori (domestic silkworm)
01100 Metabolic pathways - Bombyx mori (domestic silkworm)
00053 Ascorbate and aldarate metabolism - Bombyx mori (domestic silkworm)
00830 Retinol metabolism - Bombyx mori (domestic silkworm)
00860 Porphyrin and chlorophyll metabolism - Bombyx mori (domestic silkworm)
00980 Metabolism of xenobiotics by cytochrome P450 - Bombyx mori (domestic silkworm)
00982 Drug metabolism - cytochrome P450 - Bombyx mori (domestic silkworm)
00983 Drug metabolism - other enzymes - Bombyx mori (domestic silkworm)
01100 Metabolic pathways - Bombyx mori (domestic silkworm)
GO
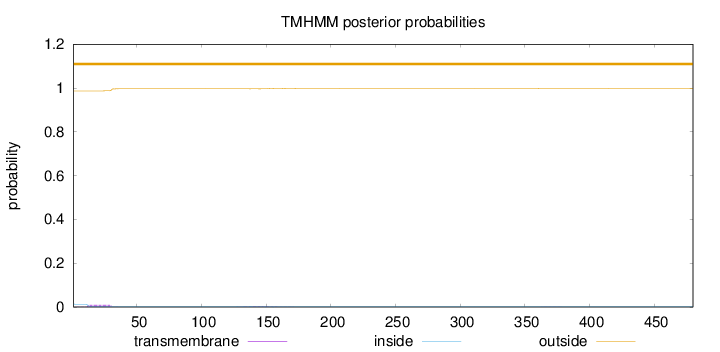
Topology
Subcellular location
Membrane
Length:
480
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.255279999999999
Exp number, first 60 AAs:
0.18212
Total prob of N-in:
0.01298
outside
1 - 480
Population Genetic Test Statistics
Pi
242.080138
Theta
152.16595
Tajima's D
1.037207
CLR
1321.878681
CSRT
0.667566621668917
Interpretation
Uncertain
