Gene
KWMTBOMO16383 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA013737
Annotation
PREDICTED:_transmembrane_protease_serine_11A-like_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 1.302 PlasmaMembrane Reliability : 1.093
Sequence
CDS
ATGGAGGTGGAAACAGTATTTGTTTACGATGGATACAATTATGAAAATGGAACAAATAACATTGCGTTGTTAGAACTGAAGAACGATGTGGTGTTTAACGAAAAAGTCCAACCTGCGTGTCTGTGGGAGTTCAGCGCATACAAGAAGTTAAATTTGAAAGACATCAAGGGATCTGTCATAAGCACAACTCTCGGCGAAGATATTGATGAAGGTCTCGAGGTAGAGAATATGAAAAAGCTTAAGCCTGTTAAAGAAGTAAATGTTTATAGGGAGGAAACATTTGAATCCAAATTCAACGCCACATCTATCAATCTGTGTACGAATGCTAAAGATTATGATAAGCATTGGGCTTTGATTGGGAATCCACTTCTTCATGCAGCTCCGTGCATCGGAAACAGTAAAGTATCTATATCATATGAACCGGGCCTGGCACCGGAGGAAGAAACTAAATACAATTTATTTATAAACGGCCCTTTTCCCGAACACACGACAATCAAAATGAAATTTGATTCGCCCGCCAATGTGACTCTGAGCAACGACGTAGGTAAATTAGCAAGAGTGTCTTCCTCTGCTGACGGAATGTTCTTCATCAAATACTTTAAAGGCGGACCCTATTTCAGTGCAGATGTGATAGGTTTAAACTTATCCGTAGTACCTTACCCGACCACTATTAACTTAAACTTTGTCGAATACTGCGAGCACCCGGCACTTGGAATCTTAGATGGATATATCAACGGATATAAATCAACAGCCTCATCTACTTATACAGAACACGATGACAACTGTGGAAGAATTAAAATCTCCGAAAAACAATGCGGTGGAAACGAAGAAGAACCTATGCCCTGGCATGCTCTTATCAGAGACGACAACTCTACAATATGCGCTGGAACGTTGATATTACAGAGATATGTTTTGACAGCCGCTCAATGCGTCACAAACTTAGGCGTTGCGAAGAACGCCACAACTCTGTCTGTGGTTCTTGGGAAATTTAATAACACAAGTGATGAATGCTTACAGGAAATGGAGGTGGAAACAGTATTTGTTTACGATGGATACAATTATGAAAATGGAACAAATAACATTGCGTTGTTAGAACTGAAGAACGATGTGGTGTTTAACGAAAAAGTCCAACCTGCGTGTCTGTGGGAATTCAGCGCATACAAGAAGTTAAATTTGAAAGACATCAAGGGATCTGTCATAAGCACAACTCTCGGCGATGATATTGATGAAGGTCTCGAGGTAGTGAATATGAAAAAGCTTAAGTCTTCTAAAGAAGTAAATGTTTATAGGGAGGAGAAATTTGGATCCAAATTCAACGCCACATCTATCAATCTGTGTACGAATGCTAAGGATTATAATAAGCATTGGGCTTTGATTGGGAATCCACTTCTTCATGCAGCCCCGTGCATCGGAAACAGTAAAGTATCTATATCATATGAACCGGGCCTGGCACCGGAGGAAGAAACTAAATACAATTTATTTATAAACGGCCCTTTCCCCGAACACACGACGATCAAAATGAAATTTGATTCGCCCGCCAATGTGACTCTGAGCAACGACATAGGCAAATTAGCAAGAGTGTCTTCTTATGCTGACGGAATGTTCTTCATCAAATACTTTAAAGGCGGACCCTATTTCAGTGCAAATGTGATAGGTTTAAACTTATCCGTAGTACCTTACCCGACCACTATTAACTTAAACTTCGTCGAATACTGCGAGCACCCGGCACTTGGAATCTTAGACGAATATATCGACGGATATAAATCAACAGCCTCATCTACTTATACAGAACACGATGACAACTGTGGAAGAATTAAAATCTCCGAAAAACAATGCGGTGGAAACGAAGAAGAACCTATGCCCTGGCATGCTCTTATCAGAGACGACAACTCTACAATATGCGCTGGAACGTTGATATTACAGAGATATGTTTTGACAGCCGCTCAATGCGTCACAAACTTAGGCGTTGCGAAGAACGCCACAACTCTGTCTGTGGTTCTTGGGAAATTTAACACAAGTGATGAATGCTTACAGGAAATGGAGGTGGAAACAGTATTTGTTTACGATGGATACAATTATGAAAATGGAACAAATAACATTGCGTTGTTAGAACTGAAGAACGATGTGGTGTTTAACGAAAAAGTCCAACCTGCGTGTCTGTGGGAGTTCAGCGCATACAAGAAGTTAAATATGAAAGACATCAAGGGATCTGTCATAAGCACAACTCTCGGCGAAGATATTGATGAAGGTCTCGAGGTAGTGAATATGAAAAAGCTTAAGCCTTTTAAAGAAGTAAATGTTTATAGGGAGGAAACATTTGGATCCAAATTCAACGCCACATCTATCAATCAATCCGAAGCCTGCAAGCTAGTTGGTAGCGGATTTATGGTGTTTGTTCCGGATTCCCGTGAATTGAATAGCACCGGAGCGTGGTACATCCACGGAATCGTCGCTACGAACGATTCGACGCTAAAATGCGATTCTAGCGATCCCACTGATTTCATCAACTTAGACTACTTCAGAGGCTGGGTCCTGAACAAAAAGAACAGATCTTAA
Protein
MEVETVFVYDGYNYENGTNNIALLELKNDVVFNEKVQPACLWEFSAYKKLNLKDIKGSVISTTLGEDIDEGLEVENMKKLKPVKEVNVYREETFESKFNATSINLCTNAKDYDKHWALIGNPLLHAAPCIGNSKVSISYEPGLAPEEETKYNLFINGPFPEHTTIKMKFDSPANVTLSNDVGKLARVSSSADGMFFIKYFKGGPYFSADVIGLNLSVVPYPTTINLNFVEYCEHPALGILDGYINGYKSTASSTYTEHDDNCGRIKISEKQCGGNEEEPMPWHALIRDDNSTICAGTLILQRYVLTAAQCVTNLGVAKNATTLSVVLGKFNNTSDECLQEMEVETVFVYDGYNYENGTNNIALLELKNDVVFNEKVQPACLWEFSAYKKLNLKDIKGSVISTTLGDDIDEGLEVVNMKKLKSSKEVNVYREEKFGSKFNATSINLCTNAKDYNKHWALIGNPLLHAAPCIGNSKVSISYEPGLAPEEETKYNLFINGPFPEHTTIKMKFDSPANVTLSNDIGKLARVSSYADGMFFIKYFKGGPYFSANVIGLNLSVVPYPTTINLNFVEYCEHPALGILDEYIDGYKSTASSTYTEHDDNCGRIKISEKQCGGNEEEPMPWHALIRDDNSTICAGTLILQRYVLTAAQCVTNLGVAKNATTLSVVLGKFNTSDECLQEMEVETVFVYDGYNYENGTNNIALLELKNDVVFNEKVQPACLWEFSAYKKLNMKDIKGSVISTTLGEDIDEGLEVVNMKKLKPFKEVNVYREETFGSKFNATSINQSEACKLVGSGFMVFVPDSRELNSTGAWYIHGIVATNDSTLKCDSSDPTDFINLDYFRGWVLNKKNRS
Summary
Similarity
Belongs to the peptidase S1 family.
Uniprot
Pubmed
EMBL
Proteomes
Pfam
PF00089 Trypsin
Interpro
SUPFAM
SSF50494
SSF50494
CDD
ProteinModelPortal
PDB
2BQW
E-value=6.6546e-10,
Score=157
Ontologies
GO
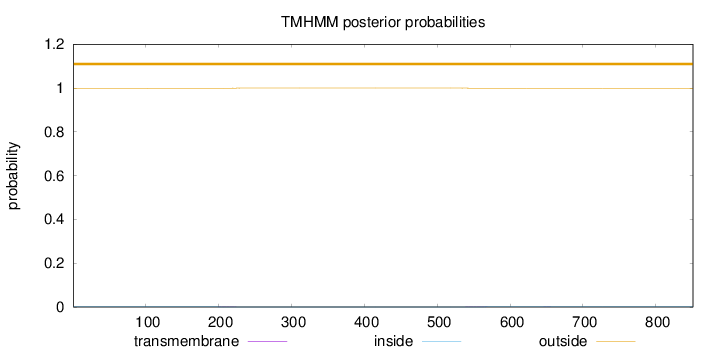
Topology
Length:
851
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.0413000000000001
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00071
outside
1 - 851
Population Genetic Test Statistics
Pi
359.027244
Theta
158.073873
Tajima's D
4.500998
CLR
0.248362
CSRT
0.99990000499975
Interpretation
Uncertain
