Gene
KWMTBOMO16360
Pre Gene Modal
BGIBMGA013795
Annotation
PREDICTED:_Down_syndrome_cell_adhesion_molecule-like_protein_Dscam2_[Amyelois_transitella]
Location in the cell
Extracellular Reliability : 1.515 Nuclear Reliability : 1.457 PlasmaMembrane Reliability : 1.205
Sequence
CDS
ATGCAAACTTTTACAGGTTTGGACAGAAGAAAAAGAATGTCAGCGAACCGGGTCATCATCACACAGCATTTCACGGAGCGAACCATCACACCCGGCGGTGATATCAGCATGCAGTGCGCAGCCACTGGCGATAAGCCTCCACAGTTTGTTTGGGAAAGAGACGGCATTGTGGTGTCAAGCAATACAGATCCTAGATATGCTTTAGGACAAGTCATGACATCAGACAATTCGGTGATCGCCCAACTAAATATAACAAGAGTCAGAGTCGAAGATGGAGGGCTTTACACGTGCACCGCTAAGGAAGGAGAATATAGTGCTAGTCACGAAAACAGACTGGATATTTATGGCCCACCATATATCAGATCCCTGCCACCAATCAAAGTGCAGAGCGGAGAATCGGTAAATTTAAGATGTCCATACTACGGATATCCAATAAGTAAAATCGATTGGGAACATCAAGGCCAAATCGTCAGCAGCAACGGCCTCTTCCAGAGCAGATACAAAAGGTCAATAACCCAACGCCGCAAAATCAGAAGAAACGCACCTACCGTCAACATTCACGGAGTGCTGACCATCTCGGAAGTCGACAAAGAACAAAGCGGTGCTGTATACACTTGCATCGTGACAAGTCCGTCAGGGGAAATGGCCAGAAGGGCTTTCGAGATCCAAGTAATCGAAGCGCCAGTCTTAGATGAACTGCTTTTGGGAAGCAACCTGCAGGAGGGGCAAATAGCAAACATTTACTGCAACGTCAGGACTGGAGATTTGCCGATACATTTCGAATGGTTGAAAGACGGGAAAAGAATACCTAGCAGTTTGAAGGTGGCAGAAAGAAACTCGGAGCTTTTCAGCGCCCTGGTAATCAAGAAAGTTTCTTTAGAGCATTGCGGTACTTACACTTGCGTCGCTTCGAACCACGTAGCAAAAGTTAATCGATCCACTGAGCTTTATATCAAAGTTGCTCCGAAATGGTTGGAAGAGCCATCAAATACATCATTACTTCTCGGCAGGAAGGGAGCAGTCTCGTGTAGTGCTAGCGGATATCCTCAACCACAAGTGCATTGGATGAAAAAGGACGCGCTGTTAGGAACGTGGCAACCAGTTCTGGAGCTAGCTGGCGGTGGAATACTCAGCCTTCCGAATGGTACGCTCGTTATTGAGGACGTGTCAGTCACGGACGAGGGTTTGTACTCGTGCAATGTTGAAAATGGCGTCGGCCAGCCTCTAAGCAAAACCATCTGGATGAGTGTGAACAAGCCGGTCCATTTCGACTCGTCTTCCTCAAACGTGACGTCACAACTCGGACATGCAGTGTCGCTGCAGTGCAGAGCTCTCGGCGACGACCCTATCCGCATCTCGTGGACTAAAGATGGGAAGATCATTAATCTTCAGAATCAGAGACTACAGCATTTGGACTCGAAGGCAAACCAAGGATTGGAAAGCACCATCAATATTCAATACTCAGAGACGACTGACGCAGGGCTTTACCAATGTCGAGCGAATAATCCATACGGTTCTGCGATTTTTAATATACACCTCACCATATTGGAGCCACCGAGTCCACCTGCCGAAGTTCGCGTAGAAGCAGTACGCCCTCGAGCCGCACTCTTGTCGTGGCGGGACTCTGCAAGGTCCTCGGTCGCGAACTACCTCGTCCAATATTGCGCTCTGAACTACGACCACGACTGGGATCGAGCAATCACCATTAATCACACTCGCAAAGACCTTGACATCCGACAGTCTCTAGAACTACAGAATCTTCGCCCGGCAACCTCATATGCAGTGAGGGTTGCTTCAACGAACCAAGTCGGCATCAGCCAATATACTGCGCCACTGCACTTCACCACCCACGAAGAAGCTCCGACGTCACCACCGGTAAACGTTCAAGTGGAACAAACCGAGAATCCCGGAGAACTGTATATCTCGTGGCTACCTCCTCTTAAAGAGACCCATAACGGTATTCTTCTCGGCTACCATCTGAAGGCGGTTGCGCAGATTAGTGGCGCTGAGGAAGCTGATTCGAGGACGTTGAAGGTCTCACAGCTTTCCGGCAAACAAGAAACCGTCGTCAGTGGACTCCTCAAAAATACTAGATATGCTGTATCGATATCAGCTTTCACTAGTGCGGGTTCCGGACCGTTCTCGTTACCAGTCTTCCGAGACACGAGAGAAGGAGCACCGGAGCAGGCTCCATCTTCGGTGGAGTGCACGGCTGTCTCTCCCACCTCGCTACGTATCGGCTGGCAGCCTCTGACGTCACCGGGACACGGCAGCTCTCTCGTGGGGTACACTTTGCTATATGCAACTGAAGAAGGACCGTGGAACAATGTAACTTCACTCCACACCGAGATCTATCTTCAATCGCTGCTGAAGTACACGAATTACACCATCAAAGTGGCCGGCTTTTCCAACTATGGTGCCGGTCCGTATTCGTACCCGATTGTATGTGCCACACTTGAGGATGTTCCGGGTCCTCCGCCTGACATCAAAGTGCTGGTGACATCAGCCACGTCACTCCTTATCAGCTGGAAAAGACCAGACCAGCCCAACGGTGAAATTCTCCACTTCACAGCATACATCAAACCAACAACAAGCAATGGTCCACCTCAGACGCACGTCGTTGAAAGCTCCAAGACCACAAAAGAGGTGCGAGAACTCAGGACAGGATCCTCATATGAGTGCTGGGTCACAGCTTCCACCCGCGCCGGGGAAGGACCTCCTAGTAGAAGAGTTTCACACACTCCAACACATCGAGTGGTAGCGGGTGTGTCGTCCCTAGGCGGGGTGGTGTCGGCTGGCGCTGGCAGCTCGGTGCTGCTGGGATGCGCGTGCGTGGGGACGCCGGCGCCGCGCACCGTGTGGTACCGACACCACCACATCATCACGCACCACCCCAGATACACCAGGAACCATGATGATAGTCTGCTCATTAACAACATAGATCAGTCTTTAAGTGGGAACTACACCTGCCTGGCCAAGAATCTCTATGGGTCTGATTCCGTTGAGTACAACGTAGTGGTGCTACCCTTGCCCGAAGCGCCAATTCTGAGGGCTACACCATACAAAGACTCAATTTTAGTAGAATGGGAACAACCTTACTACATCACCAACAGAAGTTCCATTCAAAAGATTAGTTACAGTCTCACGTGGAAGGAAGCAAACGGTCAGTGGCAAGAAGCGTGGCAGGCTAACAAGATGCCCGACTTATCCACAGTCCATAAGTACTCCTTAAAAGGGCTCAAGTGTGGTACAAAATATTCTCTAAGGATCACAGCTACAAACAAAGTTGGTACATCACAACCGGCGTATTTAGAAGTCAGCACGTTGGGAGGAGCACCTGTAGCCCCATCTACGAATGAATGGCTTTGGAGCAATAGCTCCCATATTGTGGTGCAAATGGCTGGGTGGGAGTCAGCCGGCTGTGAGCTTCGAACTTGGGAAGTTGATTACAAAGCTCTTGGAGCAAGAACTTGGATACGCACTAGAAGTAACACGGCTGTAACCGACAGTTGGAATATGCTGTACCAAGATCCAACATTTCGCTCCGGCTCTCTGGCCATCGCCGGCCTGGCTCCTGGTACTTGGTACCAGGTCAGAGTGACGGCCACCAACGAAGCCGGAAGCGTAACCTCGCTTTATAATTATGCAACAAAGAATGAGGACGGAAGTGAAATCGGCCCGCCCTCAGATCTTTTTGATCTGAACATGCTTGTGATAATCTGCAGCTCTATACTACTGACAGTGTGTCTTCTCGCTTGTCTCTACATTTTAATTAAAAGACGAAGAAGCGGCAACCTAACAGAATACAGGGATTCAATAACGAATGACAATAAATCTGACAACGGCAACCTGACGGCCAACACTTCCCACTCGAATCTGGCCAACGTGAAAGAGAATCCTGGGAATCCACAAAACAGGATATACAGTGCACCGATACATATGAGAAACAACAGTAAGCACGTGTTATTTTCAGAACTATACGAGATAAGTCCGTACGCGCAATTCGCTGTTGGCTTCCGGACATTTGGACATGTTGAAAATCAAGATCTACCAAGTCGGGTCAAGCACAGATACGACACGGAGACGAGTTTCCAAGTCTGTTCAGAATCCGAAGACAGTGACAGCATATCAAAGTCCACGTTAAAAAGCGTTCCAAGGAAAATATGCAGAACTCCGCATCGCAGATAA
Protein
MQTFTGLDRRKRMSANRVIITQHFTERTITPGGDISMQCAATGDKPPQFVWERDGIVVSSNTDPRYALGQVMTSDNSVIAQLNITRVRVEDGGLYTCTAKEGEYSASHENRLDIYGPPYIRSLPPIKVQSGESVNLRCPYYGYPISKIDWEHQGQIVSSNGLFQSRYKRSITQRRKIRRNAPTVNIHGVLTISEVDKEQSGAVYTCIVTSPSGEMARRAFEIQVIEAPVLDELLLGSNLQEGQIANIYCNVRTGDLPIHFEWLKDGKRIPSSLKVAERNSELFSALVIKKVSLEHCGTYTCVASNHVAKVNRSTELYIKVAPKWLEEPSNTSLLLGRKGAVSCSASGYPQPQVHWMKKDALLGTWQPVLELAGGGILSLPNGTLVIEDVSVTDEGLYSCNVENGVGQPLSKTIWMSVNKPVHFDSSSSNVTSQLGHAVSLQCRALGDDPIRISWTKDGKIINLQNQRLQHLDSKANQGLESTINIQYSETTDAGLYQCRANNPYGSAIFNIHLTILEPPSPPAEVRVEAVRPRAALLSWRDSARSSVANYLVQYCALNYDHDWDRAITINHTRKDLDIRQSLELQNLRPATSYAVRVASTNQVGISQYTAPLHFTTHEEAPTSPPVNVQVEQTENPGELYISWLPPLKETHNGILLGYHLKAVAQISGAEEADSRTLKVSQLSGKQETVVSGLLKNTRYAVSISAFTSAGSGPFSLPVFRDTREGAPEQAPSSVECTAVSPTSLRIGWQPLTSPGHGSSLVGYTLLYATEEGPWNNVTSLHTEIYLQSLLKYTNYTIKVAGFSNYGAGPYSYPIVCATLEDVPGPPPDIKVLVTSATSLLISWKRPDQPNGEILHFTAYIKPTTSNGPPQTHVVESSKTTKEVRELRTGSSYECWVTASTRAGEGPPSRRVSHTPTHRVVAGVSSLGGVVSAGAGSSVLLGCACVGTPAPRTVWYRHHHIITHHPRYTRNHDDSLLINNIDQSLSGNYTCLAKNLYGSDSVEYNVVVLPLPEAPILRATPYKDSILVEWEQPYYITNRSSIQKISYSLTWKEANGQWQEAWQANKMPDLSTVHKYSLKGLKCGTKYSLRITATNKVGTSQPAYLEVSTLGGAPVAPSTNEWLWSNSSHIVVQMAGWESAGCELRTWEVDYKALGARTWIRTRSNTAVTDSWNMLYQDPTFRSGSLAIAGLAPGTWYQVRVTATNEAGSVTSLYNYATKNEDGSEIGPPSDLFDLNMLVIICSSILLTVCLLACLYILIKRRRSGNLTEYRDSITNDNKSDNGNLTANTSHSNLANVKENPGNPQNRIYSAPIHMRNNSKHVLFSELYEISPYAQFAVGFRTFGHVENQDLPSRVKHRYDTETSFQVCSESEDSDSISKSTLKSVPRKICRTPHRR
Summary
Uniprot
H9JW81
A0A2A4JQ07
A0A212F605
A0A0N1I4G3
A0A3S2LWI1
A0A194PQ02
+ More
A0A0N1PIJ2 A0A212EZG7 A0A0N0P9C7 A0A0N1I745 A0A212F1J3 A0A088APY9 A0A139WPP7 A0A139WPF3 A0A139WPD8 A0A0N0BC89 A0A2J7PQP1 A0A2J7PQK1 A0A2J7PQK5 A0A0L0C0J5 A0A1B0FNH0 A0A1J1HP71 A0A151IA46 A0A151JRW6 A0A195BJQ9 A0A3L8DRQ5 F4W742 A0A158NJC3 A0A195EYM2 A0A2H1VAH3 A0A2W1BND6 A0A0Q9X218 A0A0Q9X715 B4K7M7 A0A0P8XZR3 B3LVQ3 H9J8A5 K7J3G0 A0A1W4V5H6 A0A1W4UT57 W8BHF6 A0A1W4USR1 A0A0R1E506 A0A0R1E4W5 A0A0R1E3C0 B4PVB9 B4IBG9 A0A1I8Q183 A0A1I8Q177 A0A232F8U8 B4N9U7 A0A1I8Q151 Q9VEJ5 Q7KSE9 A0A0Q9WGA5 B4LY43 A0A0M4ETQ6 A0A0Q9WFX9 A8JR35 A0A336M8I3 A0A336KKP0 Q29A88 A0A0R3NK49 B4G2L9 A0A0R3NGP1 A0A0R3NG76 A0A0R3NGB7 A0A3B0KAQ1 A0A1A9ZZV3 A0A3S2NBJ4 A0A1W7R6Y5 A0A0L7R8H0 A0A182QYC3 A0A212FEX3 A0A084WCP8 A0A182J4M0 A0A182VI12 E9J1T2 A0A182FCX3 A0A1B0BV85 A0A2H1VFK8 A0A182RKW3 W5JR02 A0A2W1BN39 N6TSR9 U4U3K5 E2A3Y9 A0A2J7PZP0 A0A0R1DV58 A0A0R1E1F0
A0A0N1PIJ2 A0A212EZG7 A0A0N0P9C7 A0A0N1I745 A0A212F1J3 A0A088APY9 A0A139WPP7 A0A139WPF3 A0A139WPD8 A0A0N0BC89 A0A2J7PQP1 A0A2J7PQK1 A0A2J7PQK5 A0A0L0C0J5 A0A1B0FNH0 A0A1J1HP71 A0A151IA46 A0A151JRW6 A0A195BJQ9 A0A3L8DRQ5 F4W742 A0A158NJC3 A0A195EYM2 A0A2H1VAH3 A0A2W1BND6 A0A0Q9X218 A0A0Q9X715 B4K7M7 A0A0P8XZR3 B3LVQ3 H9J8A5 K7J3G0 A0A1W4V5H6 A0A1W4UT57 W8BHF6 A0A1W4USR1 A0A0R1E506 A0A0R1E4W5 A0A0R1E3C0 B4PVB9 B4IBG9 A0A1I8Q183 A0A1I8Q177 A0A232F8U8 B4N9U7 A0A1I8Q151 Q9VEJ5 Q7KSE9 A0A0Q9WGA5 B4LY43 A0A0M4ETQ6 A0A0Q9WFX9 A8JR35 A0A336M8I3 A0A336KKP0 Q29A88 A0A0R3NK49 B4G2L9 A0A0R3NGP1 A0A0R3NG76 A0A0R3NGB7 A0A3B0KAQ1 A0A1A9ZZV3 A0A3S2NBJ4 A0A1W7R6Y5 A0A0L7R8H0 A0A182QYC3 A0A212FEX3 A0A084WCP8 A0A182J4M0 A0A182VI12 E9J1T2 A0A182FCX3 A0A1B0BV85 A0A2H1VFK8 A0A182RKW3 W5JR02 A0A2W1BN39 N6TSR9 U4U3K5 E2A3Y9 A0A2J7PZP0 A0A0R1DV58 A0A0R1E1F0
Pubmed
EMBL
BABH01025528
NWSH01000925
PCG73523.1
AGBW02010096
OWR49171.1
KQ459547
+ More
KPJ00028.1 RSAL01000157 RVE45570.1 KQ459602 KPI93210.1 KQ460353 KPJ15639.1 AGBW02011290 OWR46880.1 KPJ00029.1 KQ461071 KPJ09867.1 AGBW02010870 OWR47596.1 KQ971307 KYB29801.1 KYB29803.1 KYB29804.1 KQ435922 KOX68634.1 NEVH01022635 PNF18616.1 PNF18614.1 PNF18619.1 JRES01001161 KNC24949.1 CCAG010010693 CCAG010010694 CCAG010010695 CCAG010010696 CCAG010010697 CVRI01000006 CRK88017.1 KQ978255 KYM95887.1 KQ978579 KYN30034.1 KQ976453 KYM85408.1 QOIP01000005 RLU23107.1 GL887813 EGI69926.1 ADTU01017524 ADTU01017525 ADTU01017526 ADTU01017527 ADTU01017528 ADTU01017529 ADTU01017530 ADTU01017531 ADTU01017532 ADTU01017533 KQ981920 KYN32984.1 ODYU01001341 SOQ37392.1 KZ149956 PZC76468.1 CH933806 KRG01529.1 KRG01530.1 EDW15371.2 CH902617 KPU80416.1 EDV43677.1 BABH01027510 BABH01027511 BABH01027512 BABH01027513 GAMC01005845 JAC00711.1 CM000160 KRK02724.1 KRK02722.1 KRK02723.1 EDW95788.2 KRK02725.1 CH480827 EDW44727.1 NNAY01000640 OXU27246.1 CH964232 EDW80662.2 AE014297 AAF55426.3 AAS65164.2 CH940650 KRF83565.1 EDW67931.1 CP012526 ALC46292.1 KRF83564.1 ABW08696.1 UFQS01000550 UFQT01000550 SSX04800.1 SSX25163.1 SSX04801.1 SSX25164.1 CM000070 EAL27462.3 KRT00047.1 CH479179 EDW24064.1 KRT00044.1 KRT00045.1 KRT00046.1 KRT00048.1 KRT00049.1 KRT00043.1 OUUW01000005 SPP80648.1 RSAL01000273 RVE43150.1 GEHC01000744 JAV46901.1 KQ414632 KOC67128.1 AXCN02001019 AXCN02001020 AGBW02008880 OWR52305.1 ATLV01022806 ATLV01022807 ATLV01022808 ATLV01022809 ATLV01022810 ATLV01022811 KE525337 KFB47992.1 GL767674 EFZ13261.1 JXJN01021159 JXJN01021160 ODYU01002306 SOQ39628.1 ADMH02000362 ETN66827.1 PZC76469.1 APGK01020326 APGK01020327 APGK01020328 KB740193 ENN81073.1 KB631669 ERL85186.1 GL436519 EFN71841.1 NEVH01020333 PNF21807.1 CM000159 KRK01032.1 KRK01031.1
KPJ00028.1 RSAL01000157 RVE45570.1 KQ459602 KPI93210.1 KQ460353 KPJ15639.1 AGBW02011290 OWR46880.1 KPJ00029.1 KQ461071 KPJ09867.1 AGBW02010870 OWR47596.1 KQ971307 KYB29801.1 KYB29803.1 KYB29804.1 KQ435922 KOX68634.1 NEVH01022635 PNF18616.1 PNF18614.1 PNF18619.1 JRES01001161 KNC24949.1 CCAG010010693 CCAG010010694 CCAG010010695 CCAG010010696 CCAG010010697 CVRI01000006 CRK88017.1 KQ978255 KYM95887.1 KQ978579 KYN30034.1 KQ976453 KYM85408.1 QOIP01000005 RLU23107.1 GL887813 EGI69926.1 ADTU01017524 ADTU01017525 ADTU01017526 ADTU01017527 ADTU01017528 ADTU01017529 ADTU01017530 ADTU01017531 ADTU01017532 ADTU01017533 KQ981920 KYN32984.1 ODYU01001341 SOQ37392.1 KZ149956 PZC76468.1 CH933806 KRG01529.1 KRG01530.1 EDW15371.2 CH902617 KPU80416.1 EDV43677.1 BABH01027510 BABH01027511 BABH01027512 BABH01027513 GAMC01005845 JAC00711.1 CM000160 KRK02724.1 KRK02722.1 KRK02723.1 EDW95788.2 KRK02725.1 CH480827 EDW44727.1 NNAY01000640 OXU27246.1 CH964232 EDW80662.2 AE014297 AAF55426.3 AAS65164.2 CH940650 KRF83565.1 EDW67931.1 CP012526 ALC46292.1 KRF83564.1 ABW08696.1 UFQS01000550 UFQT01000550 SSX04800.1 SSX25163.1 SSX04801.1 SSX25164.1 CM000070 EAL27462.3 KRT00047.1 CH479179 EDW24064.1 KRT00044.1 KRT00045.1 KRT00046.1 KRT00048.1 KRT00049.1 KRT00043.1 OUUW01000005 SPP80648.1 RSAL01000273 RVE43150.1 GEHC01000744 JAV46901.1 KQ414632 KOC67128.1 AXCN02001019 AXCN02001020 AGBW02008880 OWR52305.1 ATLV01022806 ATLV01022807 ATLV01022808 ATLV01022809 ATLV01022810 ATLV01022811 KE525337 KFB47992.1 GL767674 EFZ13261.1 JXJN01021159 JXJN01021160 ODYU01002306 SOQ39628.1 ADMH02000362 ETN66827.1 PZC76469.1 APGK01020326 APGK01020327 APGK01020328 KB740193 ENN81073.1 KB631669 ERL85186.1 GL436519 EFN71841.1 NEVH01020333 PNF21807.1 CM000159 KRK01032.1 KRK01031.1
Proteomes
UP000005204
UP000218220
UP000007151
UP000053268
UP000283053
UP000053240
+ More
UP000005203 UP000007266 UP000053105 UP000235965 UP000037069 UP000092444 UP000183832 UP000078542 UP000078492 UP000078540 UP000279307 UP000007755 UP000005205 UP000078541 UP000009192 UP000007801 UP000002358 UP000192221 UP000002282 UP000001292 UP000095300 UP000215335 UP000007798 UP000000803 UP000008792 UP000092553 UP000001819 UP000008744 UP000268350 UP000092445 UP000053825 UP000075886 UP000030765 UP000075880 UP000075903 UP000069272 UP000092460 UP000075900 UP000000673 UP000019118 UP000030742 UP000000311
UP000005203 UP000007266 UP000053105 UP000235965 UP000037069 UP000092444 UP000183832 UP000078542 UP000078492 UP000078540 UP000279307 UP000007755 UP000005205 UP000078541 UP000009192 UP000007801 UP000002358 UP000192221 UP000002282 UP000001292 UP000095300 UP000215335 UP000007798 UP000000803 UP000008792 UP000092553 UP000001819 UP000008744 UP000268350 UP000092445 UP000053825 UP000075886 UP000030765 UP000075880 UP000075903 UP000069272 UP000092460 UP000075900 UP000000673 UP000019118 UP000030742 UP000000311
Interpro
IPR036116
FN3_sf
+ More
IPR013098 Ig_I-set
IPR013151 Immunoglobulin
IPR013783 Ig-like_fold
IPR003961 FN3_dom
IPR003599 Ig_sub
IPR007110 Ig-like_dom
IPR036179 Ig-like_dom_sf
IPR003598 Ig_sub2
IPR013106 Ig_V-set
IPR027413 GROEL-like_equatorial_sf
IPR003890 MIF4G-like_typ-3
IPR016021 MIF4-like_sf
IPR016024 ARM-type_fold
IPR018379 BEN_domain
IPR013087 Znf_C2H2_type
IPR003006 Ig/MHC_CS
IPR013098 Ig_I-set
IPR013151 Immunoglobulin
IPR013783 Ig-like_fold
IPR003961 FN3_dom
IPR003599 Ig_sub
IPR007110 Ig-like_dom
IPR036179 Ig-like_dom_sf
IPR003598 Ig_sub2
IPR013106 Ig_V-set
IPR027413 GROEL-like_equatorial_sf
IPR003890 MIF4G-like_typ-3
IPR016021 MIF4-like_sf
IPR016024 ARM-type_fold
IPR018379 BEN_domain
IPR013087 Znf_C2H2_type
IPR003006 Ig/MHC_CS
Gene 3D
CDD
ProteinModelPortal
H9JW81
A0A2A4JQ07
A0A212F605
A0A0N1I4G3
A0A3S2LWI1
A0A194PQ02
+ More
A0A0N1PIJ2 A0A212EZG7 A0A0N0P9C7 A0A0N1I745 A0A212F1J3 A0A088APY9 A0A139WPP7 A0A139WPF3 A0A139WPD8 A0A0N0BC89 A0A2J7PQP1 A0A2J7PQK1 A0A2J7PQK5 A0A0L0C0J5 A0A1B0FNH0 A0A1J1HP71 A0A151IA46 A0A151JRW6 A0A195BJQ9 A0A3L8DRQ5 F4W742 A0A158NJC3 A0A195EYM2 A0A2H1VAH3 A0A2W1BND6 A0A0Q9X218 A0A0Q9X715 B4K7M7 A0A0P8XZR3 B3LVQ3 H9J8A5 K7J3G0 A0A1W4V5H6 A0A1W4UT57 W8BHF6 A0A1W4USR1 A0A0R1E506 A0A0R1E4W5 A0A0R1E3C0 B4PVB9 B4IBG9 A0A1I8Q183 A0A1I8Q177 A0A232F8U8 B4N9U7 A0A1I8Q151 Q9VEJ5 Q7KSE9 A0A0Q9WGA5 B4LY43 A0A0M4ETQ6 A0A0Q9WFX9 A8JR35 A0A336M8I3 A0A336KKP0 Q29A88 A0A0R3NK49 B4G2L9 A0A0R3NGP1 A0A0R3NG76 A0A0R3NGB7 A0A3B0KAQ1 A0A1A9ZZV3 A0A3S2NBJ4 A0A1W7R6Y5 A0A0L7R8H0 A0A182QYC3 A0A212FEX3 A0A084WCP8 A0A182J4M0 A0A182VI12 E9J1T2 A0A182FCX3 A0A1B0BV85 A0A2H1VFK8 A0A182RKW3 W5JR02 A0A2W1BN39 N6TSR9 U4U3K5 E2A3Y9 A0A2J7PZP0 A0A0R1DV58 A0A0R1E1F0
A0A0N1PIJ2 A0A212EZG7 A0A0N0P9C7 A0A0N1I745 A0A212F1J3 A0A088APY9 A0A139WPP7 A0A139WPF3 A0A139WPD8 A0A0N0BC89 A0A2J7PQP1 A0A2J7PQK1 A0A2J7PQK5 A0A0L0C0J5 A0A1B0FNH0 A0A1J1HP71 A0A151IA46 A0A151JRW6 A0A195BJQ9 A0A3L8DRQ5 F4W742 A0A158NJC3 A0A195EYM2 A0A2H1VAH3 A0A2W1BND6 A0A0Q9X218 A0A0Q9X715 B4K7M7 A0A0P8XZR3 B3LVQ3 H9J8A5 K7J3G0 A0A1W4V5H6 A0A1W4UT57 W8BHF6 A0A1W4USR1 A0A0R1E506 A0A0R1E4W5 A0A0R1E3C0 B4PVB9 B4IBG9 A0A1I8Q183 A0A1I8Q177 A0A232F8U8 B4N9U7 A0A1I8Q151 Q9VEJ5 Q7KSE9 A0A0Q9WGA5 B4LY43 A0A0M4ETQ6 A0A0Q9WFX9 A8JR35 A0A336M8I3 A0A336KKP0 Q29A88 A0A0R3NK49 B4G2L9 A0A0R3NGP1 A0A0R3NG76 A0A0R3NGB7 A0A3B0KAQ1 A0A1A9ZZV3 A0A3S2NBJ4 A0A1W7R6Y5 A0A0L7R8H0 A0A182QYC3 A0A212FEX3 A0A084WCP8 A0A182J4M0 A0A182VI12 E9J1T2 A0A182FCX3 A0A1B0BV85 A0A2H1VFK8 A0A182RKW3 W5JR02 A0A2W1BN39 N6TSR9 U4U3K5 E2A3Y9 A0A2J7PZP0 A0A0R1DV58 A0A0R1E1F0
PDB
3DMK
E-value=9.90754e-61,
Score=597
Ontologies
GO
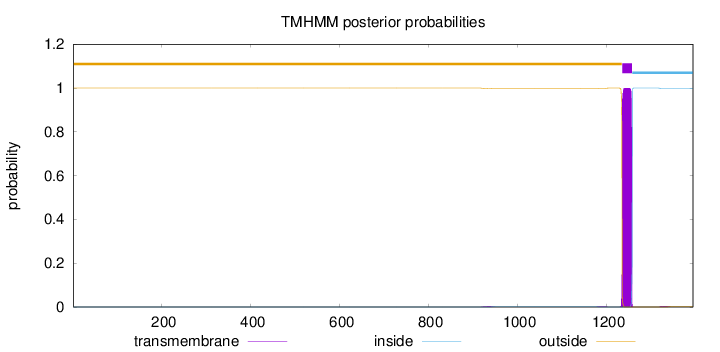
Topology
Length:
1395
Number of predicted TMHs:
1
Exp number of AAs in TMHs:
23.04249
Exp number, first 60 AAs:
0.00056
Total prob of N-in:
0.00007
outside
1 - 1235
TMhelix
1236 - 1258
inside
1259 - 1395
Population Genetic Test Statistics
Pi
268.781654
Theta
171.306194
Tajima's D
1.921106
CLR
0.110799
CSRT
0.869906504674766
Interpretation
Uncertain
