Gene
KWMTBOMO16266
Annotation
endonuclease-reverse_transcriptase_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 1.463 Mitochondrial Reliability : 1.268 Nuclear Reliability : 1.476
Sequence
CDS
ATGAAACAGCGAAGGGAAACTGCAGATGCTGCTCCGTCAGATGGAAAAACTATAAACACTAAAATCAGAAAGCTGAGTAGGCGTGACCTTCGCCAATTCAACACAACAAAGATAAAAGAGGCAATAGAACAAAACAAAGGCACCAAAGTGTTTATACGGCAACAACAACTAGGCCGAAGCCATCTGACGAAGCTCAAAACTGCCGATGGAAGAATTGTTGACTCAAGATCAGAAGTCCTCACTGAAGTGGAGAGCTACTACTGTAGTCTCTATGCCTCACATGCTCCCAAACCCTTTGCCCACCCAGAGAATGATCACCGAGCCCCTTTAGTCCGCCACTATACGGAAGACATACCCGACGTCAGTATAGAGGAGATAAGGGCTGCTTTGGAACAGCTTAGAAAATATAACAGAGCTACAGGGGATGACGGACTCAGTACAGAATTGTTGAAAGCAAGTGGTACACCAGTTCTCAACCAATTATGTACAACGTTCAACACCGTCATCCAAAAAACGACTACACCGGAAGCCTGGAGCAAGGGTACTGTCATTCTTTTCTTTAAAAAGGGAGACCGTACTCTGCTCAAGAACTACAGGCCTATTTCACTTCTTAGCCATATTTATAAGCTGTTCTCGAGAGTCCTTACAAATCGCTTGGCCTCTAGACTTGACGAGTTCCAACCTCCGGAGCAGGCTGGGTTTCGCAAAGGCTACAGTACGGTAGACCACATCCACACGCTGAGACAGGTTATTCAGAAGACTGAAGAATACAATCGTCCTCTGTGTCTCGCGTTTGTGGACTACGAAAAAGCCTTCGACCCCGTCGAAACTTGGGCTGTTCTGGAATCCTTACAGCGATGTCTAGTCGATTATCGGTACGTTGAGGTGTTGAAGAGTTTATATAAAGCCGCCAAAATGACAGTTCAGATCCAAAACCAGCAGACAAACCCGATTGAGCTTCACAGAGGGGTGCGACAGGGTGACGTTATATCGCCGAAGCTGTTCACTGCTGCTCTCGAGGATGTCTTCAAGACTCTGGACTGGAGTAAGCTGGGCATCAATGTCAATGGCGAGTATCTTTCACACCTCCGATTTGCGTACGATATCGTTATAATGGCAGAGTCGCTAGAGGATCTCAGCTGTATGCTAGGTGAGCTCAATGCCGCGTCACGACGTGTTGGTCTGGGAATGAATCTGGACAAAATTAAGGTCATGTTCAACGACCATATCATTCCTGGACCGGTAATAGTCGAATCTGCGGTACTCGAAGTTGTGTCTGAATACACCTATCTAGGCCAAATTATTCAGCTGGGCAGGGCCAACTTTGAGAAGGAAGCTGATAGGCGCATACAGTTGGGCTGGGCTGCGTATGGAAAACTCCGTCACGTCTTGAACTCAGCAATCCCGCAATGCTTAAAGACAAAAGTTTTCAACGCTTGCGTGCTGCAGATCATGACATTTGGCGCTGAAACGTGGACATTGCCGTAG
Protein
MKQRRETADAAPSDGKTINTKIRKLSRRDLRQFNTTKIKEAIEQNKGTKVFIRQQQLGRSHLTKLKTADGRIVDSRSEVLTEVESYYCSLYASHAPKPFAHPENDHRAPLVRHYTEDIPDVSIEEIRAALEQLRKYNRATGDDGLSTELLKASGTPVLNQLCTTFNTVIQKTTTPEAWSKGTVILFFKKGDRTLLKNYRPISLLSHIYKLFSRVLTNRLASRLDEFQPPEQAGFRKGYSTVDHIHTLRQVIQKTEEYNRPLCLAFVDYEKAFDPVETWAVLESLQRCLVDYRYVEVLKSLYKAAKMTVQIQNQQTNPIELHRGVRQGDVISPKLFTAALEDVFKTLDWSKLGINVNGEYLSHLRFAYDIVIMAESLEDLSCMLGELNAASRRVGLGMNLDKIKVMFNDHIIPGPVIVESAVLEVVSEYTYLGQIIQLGRANFEKEADRRIQLGWAAYGKLRHVLNSAIPQCLKTKVFNACVLQIMTFGAETWTLP
Summary
Uniprot
D7F166
D7F157
A0A3S2KX72
D7F164
D7F159
D7F165
+ More
D7F160 D7F158 A0A3S2TAR4 A4KWF2 A0A2G8JFM9 W4YXJ0 A0A2W1BLC9 A0A1E1XRS4 A0A2G8L2C7 A0A2G8K4F2 A0A2W1BDX2 A0A0N5C0B1 A0A1S3HCU5 A0A2H1VS24 A0A0K0FAW8 A0A2W1B5F8 A0A016S403 D7F177 A0A016U784 A0A2W1BZV1 D7F179 A0A069DX96 A0A016VV56 D7F176 A0A016UWI5 A0A016UTS3 A0A016RYK0 A0A023EWU8 A0A016V7H3 A0A016RXU5 A0A016WPY4 A0A2W1BCS2 A0A016WCB8 A0A016T8K4 A0A016ULU9 A0A016S1D2 A0A3P9LQX3 A0A0P4VWD0 A0A016TIA8 A0A016VBB9 A0A016VGM4 A0A016VN75 A0A016VRX3 A0A016TH80 A0A016SSP6 A0A016UUF2 A0A016T590 A0A016UGM6 A0A016TXK2 A0A016SGC3 A0A016VMK8 A0A016SX67 A0A016WKQ8 A0A016THC6 A0A016S5A1 A0A016VEC1 A0A016WEQ1 A0A016VCD9 A0A016VX03 A0A016WKT9 A0A016WVQ3 A0A016UEM0 A0A016TIG5 A0A016T5Z1 A0A016VCU3 A0A016UY78 A0A016T636 A0A016VBP7 A0A016VPL5 A0A016VVN3 A0A016WA79 A0A016US53 A0A016S795 A0A016SPR9 A0A016W5S4 A0A016TQJ1 A0A016VW83 A0A016S208 A0A016W1X2 A0A016WUN2 A0A016WRX9 A0A016X207 A0A016SCL7 A0A016VLP0
D7F160 D7F158 A0A3S2TAR4 A4KWF2 A0A2G8JFM9 W4YXJ0 A0A2W1BLC9 A0A1E1XRS4 A0A2G8L2C7 A0A2G8K4F2 A0A2W1BDX2 A0A0N5C0B1 A0A1S3HCU5 A0A2H1VS24 A0A0K0FAW8 A0A2W1B5F8 A0A016S403 D7F177 A0A016U784 A0A2W1BZV1 D7F179 A0A069DX96 A0A016VV56 D7F176 A0A016UWI5 A0A016UTS3 A0A016RYK0 A0A023EWU8 A0A016V7H3 A0A016RXU5 A0A016WPY4 A0A2W1BCS2 A0A016WCB8 A0A016T8K4 A0A016ULU9 A0A016S1D2 A0A3P9LQX3 A0A0P4VWD0 A0A016TIA8 A0A016VBB9 A0A016VGM4 A0A016VN75 A0A016VRX3 A0A016TH80 A0A016SSP6 A0A016UUF2 A0A016T590 A0A016UGM6 A0A016TXK2 A0A016SGC3 A0A016VMK8 A0A016SX67 A0A016WKQ8 A0A016THC6 A0A016S5A1 A0A016VEC1 A0A016WEQ1 A0A016VCD9 A0A016VX03 A0A016WKT9 A0A016WVQ3 A0A016UEM0 A0A016TIG5 A0A016T5Z1 A0A016VCU3 A0A016UY78 A0A016T636 A0A016VBP7 A0A016VPL5 A0A016VVN3 A0A016WA79 A0A016US53 A0A016S795 A0A016SPR9 A0A016W5S4 A0A016TQJ1 A0A016VW83 A0A016S208 A0A016W1X2 A0A016WUN2 A0A016WRX9 A0A016X207 A0A016SCL7 A0A016VLP0
Pubmed
EMBL
FJ265551
ADI61819.1
FJ265542
ADI61810.1
RSAL01002130
RVE40575.1
+ More
FJ265549 ADI61817.1 FJ265544 ADI61812.1 FJ265550 ADI61818.1 FJ265545 ADI61813.1 FJ265543 ADI61811.1 RSAL01002362 RVE40503.1 EF113398 EF113401 ABO45231.1 MRZV01002136 PIK34564.1 AAGJ04050748 KZ149985 PZC75698.1 GFAA01001426 JAU02009.1 MRZV01000249 PIK54411.1 MRZV01000899 PIK42833.1 KZ150108 PZC73382.1 ODYU01004088 SOQ43598.1 KZ150317 PZC71439.1 JARK01001639 EYB85201.1 FJ265562 ADI61830.1 JARK01001389 EYC11015.1 KZ149907 PZC78256.1 FJ265564 ADI61832.1 GBGD01000453 JAC88436.1 JARK01001340 EYC31295.1 FJ265561 ADI61829.1 JARK01001362 EYC18843.1 EYC18844.1 JARK01001677 EYB83167.1 GBBI01004877 JAC13835.1 JARK01001351 EYC23395.1 EYB83168.1 JARK01000173 EYC41342.1 KZ150485 PZC70710.1 JARK01000388 EYC37469.1 JARK01001462 EYB98997.1 JARK01001370 EYC16150.1 JARK01001655 EYB84321.1 GDKW01001596 JAI54999.1 JARK01001434 EYC02684.1 JARK01001349 EYC24735.1 JARK01001346 EYC26784.1 JARK01001344 EYC28218.1 JARK01001341 EYC29807.1 JARK01001438 EYC02012.1 JARK01001517 EYB93530.1 JARK01001361 EYC19034.1 JARK01001473 EYB97806.1 JARK01001378 EYC13977.1 JARK01001406 EYC07496.1 JARK01001568 EYB89432.1 JARK01001343 EYC28635.1 JARK01001500 EYB95040.1 EYC28825.1 JARK01000224 EYC40225.1 EYC02111.1 JARK01001630 EYB85641.1 JARK01001348 EYC25048.1 JARK01000347 EYC38051.1 EYC25050.1 EYC31303.1 JARK01000248 EYC39628.1 JARK01000077 EYC43894.1 JARK01001380 EYC13322.1 JARK01001435 EYC02496.1 JARK01001470 JARK01000204 EYB98082.1 EYC40602.1 EYC25046.1 JARK01001359 EYC20105.1 EYB98081.1 EYC40601.1 EYC25049.1 JARK01001342 EYC29360.1 EYC30828.1 JARK01000460 EYC36749.1 JARK01001365 EYC18010.1 JARK01001621 EYB86109.1 JARK01001531 EYB92324.1 JARK01000802 EYC34990.1 JARK01001421 EYC04912.1 EYC31302.1 JARK01001648 EYB84653.1 JARK01001338 EYC33297.1 JARK01000109 EYC42952.1 JARK01000132 EYC42415.1 JARK01000008 EYC46079.1 JARK01001628 JARK01001583 JARK01001528 JARK01001506 JARK01001467 JARK01001463 JARK01001457 JARK01001432 JARK01001420 JARK01001403 JARK01001371 JARK01001369 JARK01001360 JARK01001353 JARK01000603 JARK01000080 EYB85741.1 EYB88438.1 EYB92544.1 EYB94540.1 EYB98080.1 EYB98468.1 EYB98899.1 EYB99568.1 EYC03088.1 EYC05014.1 EYC08153.1 EYC15852.1 EYC16408.1 EYC19456.1 EYC22445.1 EYC24734.1 EYC24778.1 EYC24866.1 EYC33488.1 EYC35672.1 EYC43798.1 EYC28216.1
FJ265549 ADI61817.1 FJ265544 ADI61812.1 FJ265550 ADI61818.1 FJ265545 ADI61813.1 FJ265543 ADI61811.1 RSAL01002362 RVE40503.1 EF113398 EF113401 ABO45231.1 MRZV01002136 PIK34564.1 AAGJ04050748 KZ149985 PZC75698.1 GFAA01001426 JAU02009.1 MRZV01000249 PIK54411.1 MRZV01000899 PIK42833.1 KZ150108 PZC73382.1 ODYU01004088 SOQ43598.1 KZ150317 PZC71439.1 JARK01001639 EYB85201.1 FJ265562 ADI61830.1 JARK01001389 EYC11015.1 KZ149907 PZC78256.1 FJ265564 ADI61832.1 GBGD01000453 JAC88436.1 JARK01001340 EYC31295.1 FJ265561 ADI61829.1 JARK01001362 EYC18843.1 EYC18844.1 JARK01001677 EYB83167.1 GBBI01004877 JAC13835.1 JARK01001351 EYC23395.1 EYB83168.1 JARK01000173 EYC41342.1 KZ150485 PZC70710.1 JARK01000388 EYC37469.1 JARK01001462 EYB98997.1 JARK01001370 EYC16150.1 JARK01001655 EYB84321.1 GDKW01001596 JAI54999.1 JARK01001434 EYC02684.1 JARK01001349 EYC24735.1 JARK01001346 EYC26784.1 JARK01001344 EYC28218.1 JARK01001341 EYC29807.1 JARK01001438 EYC02012.1 JARK01001517 EYB93530.1 JARK01001361 EYC19034.1 JARK01001473 EYB97806.1 JARK01001378 EYC13977.1 JARK01001406 EYC07496.1 JARK01001568 EYB89432.1 JARK01001343 EYC28635.1 JARK01001500 EYB95040.1 EYC28825.1 JARK01000224 EYC40225.1 EYC02111.1 JARK01001630 EYB85641.1 JARK01001348 EYC25048.1 JARK01000347 EYC38051.1 EYC25050.1 EYC31303.1 JARK01000248 EYC39628.1 JARK01000077 EYC43894.1 JARK01001380 EYC13322.1 JARK01001435 EYC02496.1 JARK01001470 JARK01000204 EYB98082.1 EYC40602.1 EYC25046.1 JARK01001359 EYC20105.1 EYB98081.1 EYC40601.1 EYC25049.1 JARK01001342 EYC29360.1 EYC30828.1 JARK01000460 EYC36749.1 JARK01001365 EYC18010.1 JARK01001621 EYB86109.1 JARK01001531 EYB92324.1 JARK01000802 EYC34990.1 JARK01001421 EYC04912.1 EYC31302.1 JARK01001648 EYB84653.1 JARK01001338 EYC33297.1 JARK01000109 EYC42952.1 JARK01000132 EYC42415.1 JARK01000008 EYC46079.1 JARK01001628 JARK01001583 JARK01001528 JARK01001506 JARK01001467 JARK01001463 JARK01001457 JARK01001432 JARK01001420 JARK01001403 JARK01001371 JARK01001369 JARK01001360 JARK01001353 JARK01000603 JARK01000080 EYB85741.1 EYB88438.1 EYB92544.1 EYB94540.1 EYB98080.1 EYB98468.1 EYB98899.1 EYB99568.1 EYC03088.1 EYC05014.1 EYC08153.1 EYC15852.1 EYC16408.1 EYC19456.1 EYC22445.1 EYC24734.1 EYC24778.1 EYC24866.1 EYC33488.1 EYC35672.1 EYC43798.1 EYC28216.1
Proteomes
Interpro
Gene 3D
ProteinModelPortal
D7F166
D7F157
A0A3S2KX72
D7F164
D7F159
D7F165
+ More
D7F160 D7F158 A0A3S2TAR4 A4KWF2 A0A2G8JFM9 W4YXJ0 A0A2W1BLC9 A0A1E1XRS4 A0A2G8L2C7 A0A2G8K4F2 A0A2W1BDX2 A0A0N5C0B1 A0A1S3HCU5 A0A2H1VS24 A0A0K0FAW8 A0A2W1B5F8 A0A016S403 D7F177 A0A016U784 A0A2W1BZV1 D7F179 A0A069DX96 A0A016VV56 D7F176 A0A016UWI5 A0A016UTS3 A0A016RYK0 A0A023EWU8 A0A016V7H3 A0A016RXU5 A0A016WPY4 A0A2W1BCS2 A0A016WCB8 A0A016T8K4 A0A016ULU9 A0A016S1D2 A0A3P9LQX3 A0A0P4VWD0 A0A016TIA8 A0A016VBB9 A0A016VGM4 A0A016VN75 A0A016VRX3 A0A016TH80 A0A016SSP6 A0A016UUF2 A0A016T590 A0A016UGM6 A0A016TXK2 A0A016SGC3 A0A016VMK8 A0A016SX67 A0A016WKQ8 A0A016THC6 A0A016S5A1 A0A016VEC1 A0A016WEQ1 A0A016VCD9 A0A016VX03 A0A016WKT9 A0A016WVQ3 A0A016UEM0 A0A016TIG5 A0A016T5Z1 A0A016VCU3 A0A016UY78 A0A016T636 A0A016VBP7 A0A016VPL5 A0A016VVN3 A0A016WA79 A0A016US53 A0A016S795 A0A016SPR9 A0A016W5S4 A0A016TQJ1 A0A016VW83 A0A016S208 A0A016W1X2 A0A016WUN2 A0A016WRX9 A0A016X207 A0A016SCL7 A0A016VLP0
D7F160 D7F158 A0A3S2TAR4 A4KWF2 A0A2G8JFM9 W4YXJ0 A0A2W1BLC9 A0A1E1XRS4 A0A2G8L2C7 A0A2G8K4F2 A0A2W1BDX2 A0A0N5C0B1 A0A1S3HCU5 A0A2H1VS24 A0A0K0FAW8 A0A2W1B5F8 A0A016S403 D7F177 A0A016U784 A0A2W1BZV1 D7F179 A0A069DX96 A0A016VV56 D7F176 A0A016UWI5 A0A016UTS3 A0A016RYK0 A0A023EWU8 A0A016V7H3 A0A016RXU5 A0A016WPY4 A0A2W1BCS2 A0A016WCB8 A0A016T8K4 A0A016ULU9 A0A016S1D2 A0A3P9LQX3 A0A0P4VWD0 A0A016TIA8 A0A016VBB9 A0A016VGM4 A0A016VN75 A0A016VRX3 A0A016TH80 A0A016SSP6 A0A016UUF2 A0A016T590 A0A016UGM6 A0A016TXK2 A0A016SGC3 A0A016VMK8 A0A016SX67 A0A016WKQ8 A0A016THC6 A0A016S5A1 A0A016VEC1 A0A016WEQ1 A0A016VCD9 A0A016VX03 A0A016WKT9 A0A016WVQ3 A0A016UEM0 A0A016TIG5 A0A016T5Z1 A0A016VCU3 A0A016UY78 A0A016T636 A0A016VBP7 A0A016VPL5 A0A016VVN3 A0A016WA79 A0A016US53 A0A016S795 A0A016SPR9 A0A016W5S4 A0A016TQJ1 A0A016VW83 A0A016S208 A0A016W1X2 A0A016WUN2 A0A016WRX9 A0A016X207 A0A016SCL7 A0A016VLP0
PDB
6AR3
E-value=0.016205,
Score=91
Ontologies
Topology
Length:
495
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.04896
Exp number, first 60 AAs:
0
Total prob of N-in:
0.02236
outside
1 - 495
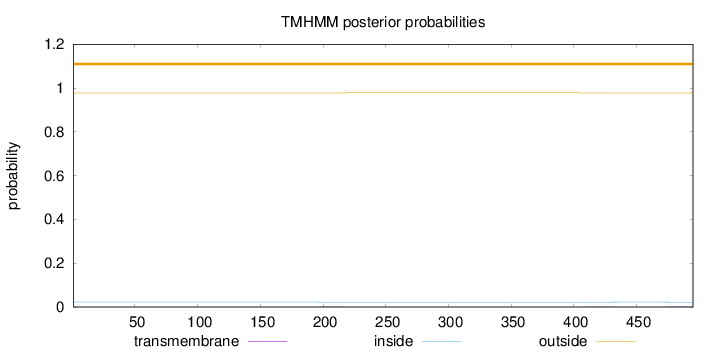
Population Genetic Test Statistics
Pi
315.198932
Theta
212.017884
Tajima's D
1.519094
CLR
0.01602
CSRT
0.794160291985401
Interpretation
Uncertain
