Gene
KWMTBOMO16220
Annotation
endonuclease-reverse_transcriptase_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 2.744
Sequence
CDS
ATGATGATCGTAGATCGTGCTAACATCAACCAACCGGAAGTCCAACATATAGCGGGATGCGAGGTAGTTAATAGCTATGTATATCTAGGCTCCACTATCACCAACGCTGGAGGCTGCGAAGATGAGATCCGAAGACGTTGTGCCGTGACAAGATCCTCAGTTGAGAGGCTAACAAAGATTTGGAGAGACCGCAGAATAACCAAAAATACAAAAGTTCGGTTAATGAGATGCCTGGTCTTTCCAATCTTTCTGTATGGGGCTGAGACGTGGACGCTGAGGATGCAAGACCGTCGTAAGATCGACGCACTAGAAATGTGGTGTTGGAGACGTCTCCTCAGAATTCCGTGGACCGCGCACCGTACCAACATTTCGATCCTGAAAGAGCTCAACATCAAGGAGAGACTATCGTCAACAGTCCAATTACGAATCCTCAAGTTCTTCGGGCACATCGCTCGTAATGAGGACTCCATTGAGCGGTTGGTGGTGCAAGGGAAAATCGAGGGCAAAAGATCTCGCGGACGATCTCCAACACGATGGACCGACCTCATAAAATCTGTTACTCACTCCAACATAAATGTTTGCTCTCACTCTGCGAAACATCGTGCCACATGGCGTCGCATCGCGAGAGCATCGGTATCGCAGGATCCGACTGATACACCACGACCACTCTGTCAAGAGTGTACGAATAGGAAAAATTGGTAA
Protein
MMIVDRANINQPEVQHIAGCEVVNSYVYLGSTITNAGGCEDEIRRRCAVTRSSVERLTKIWRDRRITKNTKVRLMRCLVFPIFLYGAETWTLRMQDRRKIDALEMWCWRRLLRIPWTAHRTNISILKELNIKERLSSTVQLRILKFFGHIARNEDSIERLVVQGKIEGKRSRGRSPTRWTDLIKSVTHSNINVCSHSAKHRATWRRIARASVSQDPTDTPRPLCQECTNRKNW
Summary
Uniprot
D7F162
D7F167
D7F175
D7F163
A0A3S2L0R7
D7F172
+ More
D7F161 A0A2W1BGD9 D7F168 D5LB39 A0A2G8JE23 A0A2G8JVA1 A0A2G8K183 A0A2H1VU13 A0A2G8K796 A0A2G8JVI4 A0A2G8L6P4 A0A2H1V742 A0A3S3NR17 A0A3S3RLW1 W4YPU3 A0A2H1VNK5 A0A0B7BSZ9 A0A0B7BUR9 A0A0B7BT80 A0A0B7BVD9 H2YWZ6 A0A0B7BVD3 A0A0B7BSB6 W5NMM5 A0A3Q1LLH3 A0A3Q1MCX3 A0A3Q1LR75 A0A3Q1M4E7 A0A3S3P648 A0A3Q1LT03 H2YWZ5 W5MM77 A0A3S1HJZ0 O97916 A0A3Q1MDY8 A0A3Q1MXT5 A0A3Q1NB85 X1D4L4 A0A3Q1M2T8 A0A3Q1M4J9 A0A2H6KKM9 A0A3Q1M8I7 A0A3Q1MGA3 A0A3Q1MIT2 A0A3Q1M7G0 A0A3Q1MJK8 A0A3Q1MN71 A0A3Q1MQ02 A0A3Q1MMS2 A0A3Q1LT63 A0A3Q1N9K5 A0A3Q1NBK7 A0A3Q1LXQ5 A0A3Q1LWU3 A0A3Q1ML90 A0A3Q1NE26 A0A3Q1LKN4 F6R5D0 A0A3Q1MVY4 A0A3Q1N3Y9 A0A3Q1ML42 A0A3Q1MJZ2 A0A3Q1MVF7 A0A3Q1LQF3 A0A3Q1M6Y4 W4Y921 A0A3Q1LUA6 A0A3Q1NDI4 A0A3Q1LGI2 A0A3Q1MJR3 A0A061BKJ9 A0A3Q1NJH7
D7F161 A0A2W1BGD9 D7F168 D5LB39 A0A2G8JE23 A0A2G8JVA1 A0A2G8K183 A0A2H1VU13 A0A2G8K796 A0A2G8JVI4 A0A2G8L6P4 A0A2H1V742 A0A3S3NR17 A0A3S3RLW1 W4YPU3 A0A2H1VNK5 A0A0B7BSZ9 A0A0B7BUR9 A0A0B7BT80 A0A0B7BVD9 H2YWZ6 A0A0B7BVD3 A0A0B7BSB6 W5NMM5 A0A3Q1LLH3 A0A3Q1MCX3 A0A3Q1LR75 A0A3Q1M4E7 A0A3S3P648 A0A3Q1LT03 H2YWZ5 W5MM77 A0A3S1HJZ0 O97916 A0A3Q1MDY8 A0A3Q1MXT5 A0A3Q1NB85 X1D4L4 A0A3Q1M2T8 A0A3Q1M4J9 A0A2H6KKM9 A0A3Q1M8I7 A0A3Q1MGA3 A0A3Q1MIT2 A0A3Q1M7G0 A0A3Q1MJK8 A0A3Q1MN71 A0A3Q1MQ02 A0A3Q1MMS2 A0A3Q1LT63 A0A3Q1N9K5 A0A3Q1NBK7 A0A3Q1LXQ5 A0A3Q1LWU3 A0A3Q1ML90 A0A3Q1NE26 A0A3Q1LKN4 F6R5D0 A0A3Q1MVY4 A0A3Q1N3Y9 A0A3Q1ML42 A0A3Q1MJZ2 A0A3Q1MVF7 A0A3Q1LQF3 A0A3Q1M6Y4 W4Y921 A0A3Q1LUA6 A0A3Q1NDI4 A0A3Q1LGI2 A0A3Q1MJR3 A0A061BKJ9 A0A3Q1NJH7
EMBL
FJ265547
ADI61815.1
FJ265552
ADI61820.1
FJ265560
ADI61828.1
+ More
FJ265548 ADI61816.1 RSAL01000289 RVE42931.1 FJ265557 ADI61825.1 FJ265546 ADI61814.1 KZ150203 PZC72267.1 FJ265553 ADI61821.1 GU815090 ADF18553.1 MRZV01002322 PIK33998.1 MRZV01001216 PIK39650.1 MRZV01000994 PIK41715.1 ODYU01004401 SOQ44236.1 MRZV01000817 PIK43880.1 MRZV01001207 PIK39730.1 MRZV01000196 PIK55924.1 ODYU01001007 SOQ36619.1 NCKU01012182 RWS00163.1 NCKU01008836 RWS01655.1 AAGJ04055194 ODYU01003523 SOQ42419.1 HACG01049248 HACG01049253 CEK96113.1 CEK96118.1 HACG01049251 HACG01049252 CEK96116.1 CEK96117.1 HACG01049244 HACG01049247 CEK96109.1 CEK96112.1 HACG01049255 CEK96120.1 HACG01049245 CEK96110.1 HACG01049249 CEK96114.1 AHAT01022938 NCKU01004395 RWS06003.1 AHAT01015286 RQTK01000360 RUS81011.1 AJ132772 CAA10770.1 BART01031562 GAH15691.1 BDSA01000081 GBE63537.1 AAGJ04161766 LK055282 CDR71995.1
FJ265548 ADI61816.1 RSAL01000289 RVE42931.1 FJ265557 ADI61825.1 FJ265546 ADI61814.1 KZ150203 PZC72267.1 FJ265553 ADI61821.1 GU815090 ADF18553.1 MRZV01002322 PIK33998.1 MRZV01001216 PIK39650.1 MRZV01000994 PIK41715.1 ODYU01004401 SOQ44236.1 MRZV01000817 PIK43880.1 MRZV01001207 PIK39730.1 MRZV01000196 PIK55924.1 ODYU01001007 SOQ36619.1 NCKU01012182 RWS00163.1 NCKU01008836 RWS01655.1 AAGJ04055194 ODYU01003523 SOQ42419.1 HACG01049248 HACG01049253 CEK96113.1 CEK96118.1 HACG01049251 HACG01049252 CEK96116.1 CEK96117.1 HACG01049244 HACG01049247 CEK96109.1 CEK96112.1 HACG01049255 CEK96120.1 HACG01049245 CEK96110.1 HACG01049249 CEK96114.1 AHAT01022938 NCKU01004395 RWS06003.1 AHAT01015286 RQTK01000360 RUS81011.1 AJ132772 CAA10770.1 BART01031562 GAH15691.1 BDSA01000081 GBE63537.1 AAGJ04161766 LK055282 CDR71995.1
Proteomes
Interpro
SUPFAM
SSF56219
SSF56219
Gene 3D
ProteinModelPortal
D7F162
D7F167
D7F175
D7F163
A0A3S2L0R7
D7F172
+ More
D7F161 A0A2W1BGD9 D7F168 D5LB39 A0A2G8JE23 A0A2G8JVA1 A0A2G8K183 A0A2H1VU13 A0A2G8K796 A0A2G8JVI4 A0A2G8L6P4 A0A2H1V742 A0A3S3NR17 A0A3S3RLW1 W4YPU3 A0A2H1VNK5 A0A0B7BSZ9 A0A0B7BUR9 A0A0B7BT80 A0A0B7BVD9 H2YWZ6 A0A0B7BVD3 A0A0B7BSB6 W5NMM5 A0A3Q1LLH3 A0A3Q1MCX3 A0A3Q1LR75 A0A3Q1M4E7 A0A3S3P648 A0A3Q1LT03 H2YWZ5 W5MM77 A0A3S1HJZ0 O97916 A0A3Q1MDY8 A0A3Q1MXT5 A0A3Q1NB85 X1D4L4 A0A3Q1M2T8 A0A3Q1M4J9 A0A2H6KKM9 A0A3Q1M8I7 A0A3Q1MGA3 A0A3Q1MIT2 A0A3Q1M7G0 A0A3Q1MJK8 A0A3Q1MN71 A0A3Q1MQ02 A0A3Q1MMS2 A0A3Q1LT63 A0A3Q1N9K5 A0A3Q1NBK7 A0A3Q1LXQ5 A0A3Q1LWU3 A0A3Q1ML90 A0A3Q1NE26 A0A3Q1LKN4 F6R5D0 A0A3Q1MVY4 A0A3Q1N3Y9 A0A3Q1ML42 A0A3Q1MJZ2 A0A3Q1MVF7 A0A3Q1LQF3 A0A3Q1M6Y4 W4Y921 A0A3Q1LUA6 A0A3Q1NDI4 A0A3Q1LGI2 A0A3Q1MJR3 A0A061BKJ9 A0A3Q1NJH7
D7F161 A0A2W1BGD9 D7F168 D5LB39 A0A2G8JE23 A0A2G8JVA1 A0A2G8K183 A0A2H1VU13 A0A2G8K796 A0A2G8JVI4 A0A2G8L6P4 A0A2H1V742 A0A3S3NR17 A0A3S3RLW1 W4YPU3 A0A2H1VNK5 A0A0B7BSZ9 A0A0B7BUR9 A0A0B7BT80 A0A0B7BVD9 H2YWZ6 A0A0B7BVD3 A0A0B7BSB6 W5NMM5 A0A3Q1LLH3 A0A3Q1MCX3 A0A3Q1LR75 A0A3Q1M4E7 A0A3S3P648 A0A3Q1LT03 H2YWZ5 W5MM77 A0A3S1HJZ0 O97916 A0A3Q1MDY8 A0A3Q1MXT5 A0A3Q1NB85 X1D4L4 A0A3Q1M2T8 A0A3Q1M4J9 A0A2H6KKM9 A0A3Q1M8I7 A0A3Q1MGA3 A0A3Q1MIT2 A0A3Q1M7G0 A0A3Q1MJK8 A0A3Q1MN71 A0A3Q1MQ02 A0A3Q1MMS2 A0A3Q1LT63 A0A3Q1N9K5 A0A3Q1NBK7 A0A3Q1LXQ5 A0A3Q1LWU3 A0A3Q1ML90 A0A3Q1NE26 A0A3Q1LKN4 F6R5D0 A0A3Q1MVY4 A0A3Q1N3Y9 A0A3Q1ML42 A0A3Q1MJZ2 A0A3Q1MVF7 A0A3Q1LQF3 A0A3Q1M6Y4 W4Y921 A0A3Q1LUA6 A0A3Q1NDI4 A0A3Q1LGI2 A0A3Q1MJR3 A0A061BKJ9 A0A3Q1NJH7
Ontologies
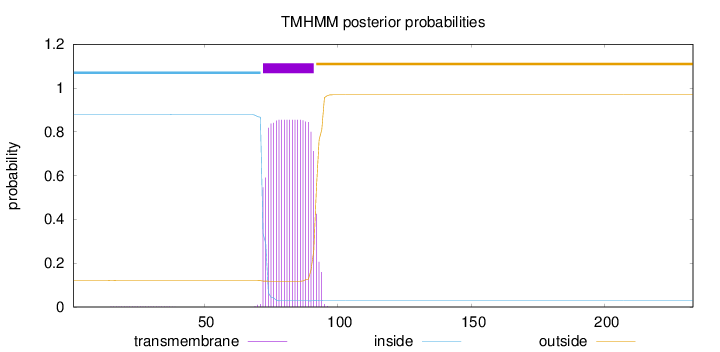
Topology
Length:
233
Number of predicted TMHs:
1
Exp number of AAs in TMHs:
17.10246
Exp number, first 60 AAs:
0.03548
Total prob of N-in:
0.87800
inside
1 - 71
TMhelix
72 - 91
outside
92 - 233
Population Genetic Test Statistics
Pi
250.188914
Theta
79.251898
Tajima's D
4.978188
CLR
1.230962
CSRT
0.999950002499875
Interpretation
Possibly Balancing Selection
