Gene
KWMTBOMO16069
Pre Gene Modal
BGIBMGA009519
Annotation
Yokozuna_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 3.292
Sequence
CDS
ATGCGCTATGATAATGAAAGCAACTTACAGCAATATTTCCTAAAATTTGAAGGAATTCTTAGCGAATTAGCGGCAATCAACGTAAACGTCGATGTAGAAGATCAAATTTGCTATCTGCTGTCTTCTTTACCAAAGATATACGATCAATTAATTACATCAATTGAGACTATGGCTTCCGAGAAACAGCTGAACCTAGAATTTGTTAAAGCAAGACTACTCGATATGGAGACAAAGTTAAAAATGGACAGAAATAGTGAAAGCTATGAGAATAATGCCTTCATCACGTGCCACTCCTGCGGAAAACGTGGACACAAATCATTCGAATGCCGAAGATCAGGACGAGGACAAGGGCCGAGTAGAGGACAGTCATTGACACGAGGATGGAGCTATCCAAGATCGCGTGGACGCTCATTTAGAGGAAATTCAAGTTCACGACATACAACGTATCAACATACTTCAGCAGCAGGCAGAGACGACAAAGAAATAGGTTTCATTGCTAGCGATATTATTGCTAATGTGGGTGAATGTGGTAACATAGAACTAATTATAGACTCAGGATGTACGGAGCATTTAATAAATGAAAAATATGAAAATTATATGACGGATATTACGTACTTAGATAAACATGTCAAAATATTCGTAGCTAACGGCCAATGTATTACAACTAGTAAAAGAGGTAAACTTAAAGTAAAATATCAAAATACTACGATTAATATTGATGCTCTAATTGTGAAAGATGTTTCATACAATTTATTATCAGTAAAAAAGATTGTCGAGGCAGGTTTTAACATAAACTTTAATAAATACAATGTTAAAATATTTAATAAAAATACATTTTTACGCGGCACCGTGCAAGGAAAGTTGTATAAGCTACAAGCCGAATTATGTAAGTCCGAAGTTTGTAATGTTACTGATGCGACAGATTTATGGCACAAACATTTAGGTCACTTAAATAGGAAAAGCCTTAATATACTAAACTTACCTGTATCTAATAGAATTTGCAGTCCTTGTATGGAGGGCAAAGCCACCCGAAGCCCATTCACGCCTATCCCAAAGCCTAGAACGAAAATGACAGCCGAGATGCTTCACACAGATATAGCAGGACCGATCAGACATCAAGGACCTAATGGTGAAAAATATTTTATGACAATTATTGACGATTTCTCTCATTTTTGCGTAGTGTATCCCCTGAAGAGCAAATCTGAAGCAACAGAACGACTGATGAACTATGTATTAAGAATTGAAAACGAAACGGGGAATAAAGTTAAAAGAATTAGATGCGATAATGGTGGAGAGTACACTGCTAATAGTTTGAAACGTTTTTGCATAAATAGAGGTATTAGTTTAGAATATACATTACCATACACGCCACAACTTAATGGCATATCAGAACGGATGATCCGCACATTATGCGACAAAACGAGGACTCTATTAGCGGAAACAAATTTGCCAAAACATCTGTGGTGCGAAGCGATACAATGTGCAGTTTACCAATTAAACAGATCGCCATCATGGGCCATAAACTTCAAGACACCCAGCGAAATATTTTTTGGTAAAAACGACCTATCGAGATTGAGAGTATTTGGTTCCAAAGCTTGGGCTACTATAGTACCCAAACCGGACAAGTTTGCCAAGAGAGCAAAAGAAACAAGAATGGTAGGTTATGGAACTGTTGGTTACAGATTATGGGATCCAGAAGAAGATAAAATAGTTATTTCCAGAGATGTAAGATTTGATGAAAGCGATATTAAATTCAAAAATAACGAAGAAGTGAAACAAGTTAGATATATTCAAGACAAAGAAGAAGAAATCAATAAAGAAAAGCAAGATATTGTAGAACAGGAAGATGAATTTAAAGATGCAGAAATGAACACTGATCAAACTGAAGAAGCAATAATCAGACCCAAAAGAACGATAAATCCTCCTGCTTATCTTAAAGATTACGAGACGAATATAGAAAACATAAATGAAACTAATATTGTTATATGTCTGAGTGCACATGCACCAGAAACTTTCGAAGAAGCTATAAAAGAACCAGAATGGAAAAACGCAATCAATAAAGAGCTAGATGCACTCCAAAGACTAGAAACGTGGGAAGAACAAGAAATAACCGATGGAAGAAAACTCATAGATACAAAATGGATATTTAAACAAAAAGAAGATGGAACAAAAAAGGCAAGACTGGTAGCTAGAGGCTTCCAACAAAAGAAAAAAGATATTTTTGACAGCGTATATGCACCCGTTGCTAGACATTCAACCATAAGAATGATCTTATCAAAAGCTGTACAAGATGATTTAAAGATTATTCAGCTTGACATTCCCACTGCATTCCTAAATGGAAAATTAAAGTCTGATGTATTTATCAAAATACCCCAAGGATTAGAAATAAAAAATAATAATACAGCATTAAAACTAAAGAGAGCATTGTATGGATTGATTGAATCACCAAAATGTTGGAATGAAACACTTGACGAATTTGCGAAAGCTAACGGATTCCAAAGATCAGAATATGATTACTGTTTGTATTACAAAGAAGATTGCTGGATGCTGATTTACGTGGACGATATCCTAATCATTGGAAACGGGGAAGATATTATTAAAGCACTTAAGGAGAAGTTTAACGCAAAATATATGGGCGAAATTAAGAACTTTCTCGGAATGGAAGTTAAAAGAAATGAAGATACCTTAGAACTGAAACAAACTAAAATGATTGAAAAGATATTGACTAAATTCAAAATGGATGAATGTAACGGAATTAAAACTCCCATGGAAAGAAATTACAATATAGAAATAAAGGACAATACAGAAGTTTTGGACGTACCGTATCGAGAACTAATAGGAAGTCTAATTTATCTTACTGTAACAACAAGACCGGATTTAACATTTGCAGTGTCATATCTGAGCAGATTTTTAGATAAACCGAACGAAGAATTATGGAAAGCTGGAAAAAGAGTAGTACGTTATTTGAAAGAAACAAAAGAAAAAGGATTATTATTTAAAAAGACAAATGACAATAATTTATACGGATTTTCTGATGCCGATTGGGCTGGAGATAGAAGCGACAGAAAAAGTGTTAGCGGTGGTATTATCCTACAAGGGCAAAACCCCATTGCTTGGTTTTCAAAGAAGCAACAATGCGTAACTTTATCGTCTGCTGAAAGCGAGTATGTAGCCGCCGCATCATTAGCACAGGAAATTGTTAATTTAAAAGGCATATGTAAGCACCTAGGTTGTGATGTAAATGCCGTGCTATTTGTTGATAACAAAAGTGCGATATCAATGGCCAAGAATTACGAGAACTCTAAGCGTACAAAACACATTGATATACGGTTTCATTATATAAAAGACTTGGTTGTCAATAAGGTGATCAGCATAGAACATATTTGTACCAATGAAAATTTAGCGGATGTTATGACTAAAAGTTTGAGCACAGACAAATTTTATAAATGCATTGATAAGTTCTTGTGCTAA
Protein
MRYDNESNLQQYFLKFEGILSELAAINVNVDVEDQICYLLSSLPKIYDQLITSIETMASEKQLNLEFVKARLLDMETKLKMDRNSESYENNAFITCHSCGKRGHKSFECRRSGRGQGPSRGQSLTRGWSYPRSRGRSFRGNSSSRHTTYQHTSAAGRDDKEIGFIASDIIANVGECGNIELIIDSGCTEHLINEKYENYMTDITYLDKHVKIFVANGQCITTSKRGKLKVKYQNTTINIDALIVKDVSYNLLSVKKIVEAGFNINFNKYNVKIFNKNTFLRGTVQGKLYKLQAELCKSEVCNVTDATDLWHKHLGHLNRKSLNILNLPVSNRICSPCMEGKATRSPFTPIPKPRTKMTAEMLHTDIAGPIRHQGPNGEKYFMTIIDDFSHFCVVYPLKSKSEATERLMNYVLRIENETGNKVKRIRCDNGGEYTANSLKRFCINRGISLEYTLPYTPQLNGISERMIRTLCDKTRTLLAETNLPKHLWCEAIQCAVYQLNRSPSWAINFKTPSEIFFGKNDLSRLRVFGSKAWATIVPKPDKFAKRAKETRMVGYGTVGYRLWDPEEDKIVISRDVRFDESDIKFKNNEEVKQVRYIQDKEEEINKEKQDIVEQEDEFKDAEMNTDQTEEAIIRPKRTINPPAYLKDYETNIENINETNIVICLSAHAPETFEEAIKEPEWKNAINKELDALQRLETWEEQEITDGRKLIDTKWIFKQKEDGTKKARLVARGFQQKKKDIFDSVYAPVARHSTIRMILSKAVQDDLKIIQLDIPTAFLNGKLKSDVFIKIPQGLEIKNNNTALKLKRALYGLIESPKCWNETLDEFAKANGFQRSEYDYCLYYKEDCWMLIYVDDILIIGNGEDIIKALKEKFNAKYMGEIKNFLGMEVKRNEDTLELKQTKMIEKILTKFKMDECNGIKTPMERNYNIEIKDNTEVLDVPYRELIGSLIYLTVTTRPDLTFAVSYLSRFLDKPNEELWKAGKRVVRYLKETKEKGLLFKKTNDNNLYGFSDADWAGDRSDRKSVSGGIILQGQNPIAWFSKKQQCVTLSSAESEYVAAASLAQEIVNLKGICKHLGCDVNAVLFVDNKSAISMAKNYENSKRTKHIDIRFHYIKDLVVNKVISIEHICTNENLADVMTKSLSTDKFYKCIDKFLC
Summary
Uniprot
O96968
A0A146L6N5
A0A0A9XG27
A0A0A9Z0I6
A0A146KTS1
A0A1B6DFK3
+ More
A0A0A9WLY1 A0A146LV28 A0A224XA26 A0A0T6AT79 A0A182GA29 A0A182GSW5 A0A1Y1L115 J9KJ72 A0A0J7K6U3 A0A182G1G7 A0A182H4X3 A0A1W7R6I5 A0A023F102 A0A1Y1MQF0 A0A151S6I6 A0A151QZS0 A0A1W7R6C3 A0A151RAL1 A0A034VP82 A0A151RCT9 A0A1W7R685 A0A1Y1MJE9 A0A182GGE6 A0A2R6RYD4 A0A2N9I6N4 A0A068B703 A0A0A9W6B0 A0A087SVE4 A0A1B6C2D3 Q9ZRJ0 A0A2N9ETC8 A0A151QNH6 A0A151T355 A0A2N9H4V1 A0A2N9FQJ1 A0A0A9ZBW6 A0A2N9FZ01 A0A2N9IGN4 A0A151R6U8 A0A2R6QHR4 A0A2N9GA80 A0A2K3N1R3 B8YLY4 A0A2N9HKJ4 A0A1Y1LCN3 B8YLY5 B8YLY6 H6U1F0 A0A2N9EE21 A0A151THM1 A0A151R2G6 A0A151TKT1 A0A2N9I9F7 B8YLY3 B8YLY7 A0A2N9I8Q5 A0A151TNK0 A0A151SDY6 A0A151RJN4 A0A1W7R6A3 A0A151UBW1 A0A2N9IGN3 A0A151UC72 A0A251VAZ6 A0A151UCJ8 H6U1E9 A0A2N9H6B2 A0A0A9W6J9 A0A0A9WDS7 A0A0J7K6W1 B1N668 A0A2Z6N0M9 A0A2B4R8Q4 A0A2N9FPD9 A0A2N9H0T6 T1PKV2 A0A151TVS9 A0A251V331 A0A224XJ98 A0A2K3NXK0 A0A2N9FMR2 A0A2N9FWL6 Q6L3N8 A0A2N9GLP2 A0A1Y1NGN0 Q967L5 A0A2N9I0Q5 A0A2N9ISP1
A0A0A9WLY1 A0A146LV28 A0A224XA26 A0A0T6AT79 A0A182GA29 A0A182GSW5 A0A1Y1L115 J9KJ72 A0A0J7K6U3 A0A182G1G7 A0A182H4X3 A0A1W7R6I5 A0A023F102 A0A1Y1MQF0 A0A151S6I6 A0A151QZS0 A0A1W7R6C3 A0A151RAL1 A0A034VP82 A0A151RCT9 A0A1W7R685 A0A1Y1MJE9 A0A182GGE6 A0A2R6RYD4 A0A2N9I6N4 A0A068B703 A0A0A9W6B0 A0A087SVE4 A0A1B6C2D3 Q9ZRJ0 A0A2N9ETC8 A0A151QNH6 A0A151T355 A0A2N9H4V1 A0A2N9FQJ1 A0A0A9ZBW6 A0A2N9FZ01 A0A2N9IGN4 A0A151R6U8 A0A2R6QHR4 A0A2N9GA80 A0A2K3N1R3 B8YLY4 A0A2N9HKJ4 A0A1Y1LCN3 B8YLY5 B8YLY6 H6U1F0 A0A2N9EE21 A0A151THM1 A0A151R2G6 A0A151TKT1 A0A2N9I9F7 B8YLY3 B8YLY7 A0A2N9I8Q5 A0A151TNK0 A0A151SDY6 A0A151RJN4 A0A1W7R6A3 A0A151UBW1 A0A2N9IGN3 A0A151UC72 A0A251VAZ6 A0A151UCJ8 H6U1E9 A0A2N9H6B2 A0A0A9W6J9 A0A0A9WDS7 A0A0J7K6W1 B1N668 A0A2Z6N0M9 A0A2B4R8Q4 A0A2N9FPD9 A0A2N9H0T6 T1PKV2 A0A151TVS9 A0A251V331 A0A224XJ98 A0A2K3NXK0 A0A2N9FMR2 A0A2N9FWL6 Q6L3N8 A0A2N9GLP2 A0A1Y1NGN0 Q967L5 A0A2N9I0Q5 A0A2N9ISP1
Pubmed
EMBL
AB014676
BAA74713.1
GDHC01015802
JAQ02827.1
GBHO01027564
JAG16040.1
+ More
GBHO01004832 JAG38772.1 GDHC01019310 JAP99318.1 GEDC01012841 GEDC01000503 JAS24457.1 JAS36795.1 GBHO01035173 JAG08431.1 GDHC01007058 JAQ11571.1 GFTR01008662 JAW07764.1 LJIG01022881 KRT78250.1 JXUM01155348 KQ572178 KXJ68066.1 JXUM01085595 KQ563513 KXJ73717.1 GEZM01072005 JAV65635.1 ABLF02041793 LBMM01012492 KMQ86193.1 JXUM01136904 KQ568499 KXJ68982.1 JXUM01025857 KQ560736 KXJ81076.1 GEHC01000878 JAV46767.1 GBBI01004033 JAC14679.1 GEZM01028916 JAV86196.1 KQ483456 KYP50361.1 KQ484324 KYP35753.1 GEHC01000928 JAV46717.1 KQ483890 KYP39660.1 GAKP01015342 JAC43610.1 KQ483843 KYP40337.1 GEHC01000966 JAV46679.1 GEZM01032204 JAV84770.1 JXUM01011189 KQ560325 KXJ83029.1 NKQK01000002 PSS35035.1 OIVN01004879 SPD19723.1 KF740825 AIC77183.1 GBHO01040663 JAG02941.1 KK112142 KFM56833.1 GEDC01029908 JAS07390.1 D83003 BAA11674.1 OIVN01000557 SPC82046.1 KQ485623 KYP31853.1 CM003610 KYP61490.1 OIVN01002854 SPD06945.1 OIVN01001052 SPC89229.1 GBHO01002761 JAG40843.1 OIVN01001309 SPC92453.1 OIVN01005657 SPD23465.1 KQ484038 KYP38095.1 NKQK01000016 PSS08126.1 OIVN01001666 SPC96375.1 ASHM01015075 PNX96991.1 FJ544853 ACL97384.1 OIVN01003569 SPD12194.1 GEZM01063091 JAV69316.1 FJ544854 ACL97385.1 FJ544856 ACL97386.1 JN402333 AFB73912.1 OIVN01000036 SPC73022.1 CM003608 KYP66486.1 KQ484186 KYP36645.1 CM003607 KYP67664.1 OIVN01005059 SPD20680.1 FJ544851 ACL97383.1 FJ544857 ACL97387.1 OIVN01005024 SPD20514.1 CM003606 KYP68607.1 KQ483417 KYP53032.1 KQ483702 KYP42748.1 GEHC01000959 JAV46686.1 CM003603 KYP76810.1 OIVN01005591 SPD23209.1 KYP76811.1 CM007892 OTG31811.1 KYP77007.1 JN402332 AFB73911.1 OIVN01002904 SPD07378.1 GBHO01040573 JAG03031.1 GBHO01040574 JAG03030.1 LBMM01012645 KMQ86082.1 EF094939 EF094940 ABO36622.1 ABO36636.1 DF973299 GAU24969.1 LSMT01001145 PFX12880.1 OIVN01001022 SPC88819.1 OIVN01002680 SPD05612.1 KA648745 AFP63374.1 CM003605 KYP71150.1 CM007893 OTG30017.1 GFTR01008327 JAW08099.1 ASHM01002084 PNY07765.1 OIVN01000991 OIVN01001835 SPC88370.1 SPC98114.1 OIVN01001534 SPC95017.1 AC149291 AAT38758.1 OIVN01002055 SPD00174.1 GEZM01005617 GEZM01005616 GEZM01005615 GEZM01005614 JAV96040.1 AY009101 AAK12626.1 OIVN01004867 SPD19656.1 OIVN01006233 SPD28466.1
GBHO01004832 JAG38772.1 GDHC01019310 JAP99318.1 GEDC01012841 GEDC01000503 JAS24457.1 JAS36795.1 GBHO01035173 JAG08431.1 GDHC01007058 JAQ11571.1 GFTR01008662 JAW07764.1 LJIG01022881 KRT78250.1 JXUM01155348 KQ572178 KXJ68066.1 JXUM01085595 KQ563513 KXJ73717.1 GEZM01072005 JAV65635.1 ABLF02041793 LBMM01012492 KMQ86193.1 JXUM01136904 KQ568499 KXJ68982.1 JXUM01025857 KQ560736 KXJ81076.1 GEHC01000878 JAV46767.1 GBBI01004033 JAC14679.1 GEZM01028916 JAV86196.1 KQ483456 KYP50361.1 KQ484324 KYP35753.1 GEHC01000928 JAV46717.1 KQ483890 KYP39660.1 GAKP01015342 JAC43610.1 KQ483843 KYP40337.1 GEHC01000966 JAV46679.1 GEZM01032204 JAV84770.1 JXUM01011189 KQ560325 KXJ83029.1 NKQK01000002 PSS35035.1 OIVN01004879 SPD19723.1 KF740825 AIC77183.1 GBHO01040663 JAG02941.1 KK112142 KFM56833.1 GEDC01029908 JAS07390.1 D83003 BAA11674.1 OIVN01000557 SPC82046.1 KQ485623 KYP31853.1 CM003610 KYP61490.1 OIVN01002854 SPD06945.1 OIVN01001052 SPC89229.1 GBHO01002761 JAG40843.1 OIVN01001309 SPC92453.1 OIVN01005657 SPD23465.1 KQ484038 KYP38095.1 NKQK01000016 PSS08126.1 OIVN01001666 SPC96375.1 ASHM01015075 PNX96991.1 FJ544853 ACL97384.1 OIVN01003569 SPD12194.1 GEZM01063091 JAV69316.1 FJ544854 ACL97385.1 FJ544856 ACL97386.1 JN402333 AFB73912.1 OIVN01000036 SPC73022.1 CM003608 KYP66486.1 KQ484186 KYP36645.1 CM003607 KYP67664.1 OIVN01005059 SPD20680.1 FJ544851 ACL97383.1 FJ544857 ACL97387.1 OIVN01005024 SPD20514.1 CM003606 KYP68607.1 KQ483417 KYP53032.1 KQ483702 KYP42748.1 GEHC01000959 JAV46686.1 CM003603 KYP76810.1 OIVN01005591 SPD23209.1 KYP76811.1 CM007892 OTG31811.1 KYP77007.1 JN402332 AFB73911.1 OIVN01002904 SPD07378.1 GBHO01040573 JAG03031.1 GBHO01040574 JAG03030.1 LBMM01012645 KMQ86082.1 EF094939 EF094940 ABO36622.1 ABO36636.1 DF973299 GAU24969.1 LSMT01001145 PFX12880.1 OIVN01001022 SPC88819.1 OIVN01002680 SPD05612.1 KA648745 AFP63374.1 CM003605 KYP71150.1 CM007893 OTG30017.1 GFTR01008327 JAW08099.1 ASHM01002084 PNY07765.1 OIVN01000991 OIVN01001835 SPC88370.1 SPC98114.1 OIVN01001534 SPC95017.1 AC149291 AAT38758.1 OIVN01002055 SPD00174.1 GEZM01005617 GEZM01005616 GEZM01005615 GEZM01005614 JAV96040.1 AY009101 AAK12626.1 OIVN01004867 SPD19656.1 OIVN01006233 SPD28466.1
Proteomes
PRIDE
Pfam
Interpro
IPR036397
RNaseH_sf
+ More
IPR039537 Retrotran_Ty1/copia-like
IPR001584 Integrase_cat-core
IPR013103 RVT_2
IPR012337 RNaseH-like_sf
IPR001878 Znf_CCHC
IPR036875 Znf_CCHC_sf
IPR021109 Peptidase_aspartic_dom_sf
IPR025724 GAG-pre-integrase_dom
IPR025314 DUF4219
IPR038717 Tc1-like_DDE_dom
IPR032675 LRR_dom_sf
IPR036576 WRKY_dom_sf
IPR001611 Leu-rich_rpt
IPR003657 WRKY_dom
IPR039537 Retrotran_Ty1/copia-like
IPR001584 Integrase_cat-core
IPR013103 RVT_2
IPR012337 RNaseH-like_sf
IPR001878 Znf_CCHC
IPR036875 Znf_CCHC_sf
IPR021109 Peptidase_aspartic_dom_sf
IPR025724 GAG-pre-integrase_dom
IPR025314 DUF4219
IPR038717 Tc1-like_DDE_dom
IPR032675 LRR_dom_sf
IPR036576 WRKY_dom_sf
IPR001611 Leu-rich_rpt
IPR003657 WRKY_dom
Gene 3D
ProteinModelPortal
O96968
A0A146L6N5
A0A0A9XG27
A0A0A9Z0I6
A0A146KTS1
A0A1B6DFK3
+ More
A0A0A9WLY1 A0A146LV28 A0A224XA26 A0A0T6AT79 A0A182GA29 A0A182GSW5 A0A1Y1L115 J9KJ72 A0A0J7K6U3 A0A182G1G7 A0A182H4X3 A0A1W7R6I5 A0A023F102 A0A1Y1MQF0 A0A151S6I6 A0A151QZS0 A0A1W7R6C3 A0A151RAL1 A0A034VP82 A0A151RCT9 A0A1W7R685 A0A1Y1MJE9 A0A182GGE6 A0A2R6RYD4 A0A2N9I6N4 A0A068B703 A0A0A9W6B0 A0A087SVE4 A0A1B6C2D3 Q9ZRJ0 A0A2N9ETC8 A0A151QNH6 A0A151T355 A0A2N9H4V1 A0A2N9FQJ1 A0A0A9ZBW6 A0A2N9FZ01 A0A2N9IGN4 A0A151R6U8 A0A2R6QHR4 A0A2N9GA80 A0A2K3N1R3 B8YLY4 A0A2N9HKJ4 A0A1Y1LCN3 B8YLY5 B8YLY6 H6U1F0 A0A2N9EE21 A0A151THM1 A0A151R2G6 A0A151TKT1 A0A2N9I9F7 B8YLY3 B8YLY7 A0A2N9I8Q5 A0A151TNK0 A0A151SDY6 A0A151RJN4 A0A1W7R6A3 A0A151UBW1 A0A2N9IGN3 A0A151UC72 A0A251VAZ6 A0A151UCJ8 H6U1E9 A0A2N9H6B2 A0A0A9W6J9 A0A0A9WDS7 A0A0J7K6W1 B1N668 A0A2Z6N0M9 A0A2B4R8Q4 A0A2N9FPD9 A0A2N9H0T6 T1PKV2 A0A151TVS9 A0A251V331 A0A224XJ98 A0A2K3NXK0 A0A2N9FMR2 A0A2N9FWL6 Q6L3N8 A0A2N9GLP2 A0A1Y1NGN0 Q967L5 A0A2N9I0Q5 A0A2N9ISP1
A0A0A9WLY1 A0A146LV28 A0A224XA26 A0A0T6AT79 A0A182GA29 A0A182GSW5 A0A1Y1L115 J9KJ72 A0A0J7K6U3 A0A182G1G7 A0A182H4X3 A0A1W7R6I5 A0A023F102 A0A1Y1MQF0 A0A151S6I6 A0A151QZS0 A0A1W7R6C3 A0A151RAL1 A0A034VP82 A0A151RCT9 A0A1W7R685 A0A1Y1MJE9 A0A182GGE6 A0A2R6RYD4 A0A2N9I6N4 A0A068B703 A0A0A9W6B0 A0A087SVE4 A0A1B6C2D3 Q9ZRJ0 A0A2N9ETC8 A0A151QNH6 A0A151T355 A0A2N9H4V1 A0A2N9FQJ1 A0A0A9ZBW6 A0A2N9FZ01 A0A2N9IGN4 A0A151R6U8 A0A2R6QHR4 A0A2N9GA80 A0A2K3N1R3 B8YLY4 A0A2N9HKJ4 A0A1Y1LCN3 B8YLY5 B8YLY6 H6U1F0 A0A2N9EE21 A0A151THM1 A0A151R2G6 A0A151TKT1 A0A2N9I9F7 B8YLY3 B8YLY7 A0A2N9I8Q5 A0A151TNK0 A0A151SDY6 A0A151RJN4 A0A1W7R6A3 A0A151UBW1 A0A2N9IGN3 A0A151UC72 A0A251VAZ6 A0A151UCJ8 H6U1E9 A0A2N9H6B2 A0A0A9W6J9 A0A0A9WDS7 A0A0J7K6W1 B1N668 A0A2Z6N0M9 A0A2B4R8Q4 A0A2N9FPD9 A0A2N9H0T6 T1PKV2 A0A151TVS9 A0A251V331 A0A224XJ98 A0A2K3NXK0 A0A2N9FMR2 A0A2N9FWL6 Q6L3N8 A0A2N9GLP2 A0A1Y1NGN0 Q967L5 A0A2N9I0Q5 A0A2N9ISP1
PDB
3S3O
E-value=1.68932e-08,
Score=146
Ontologies
GO
PANTHER
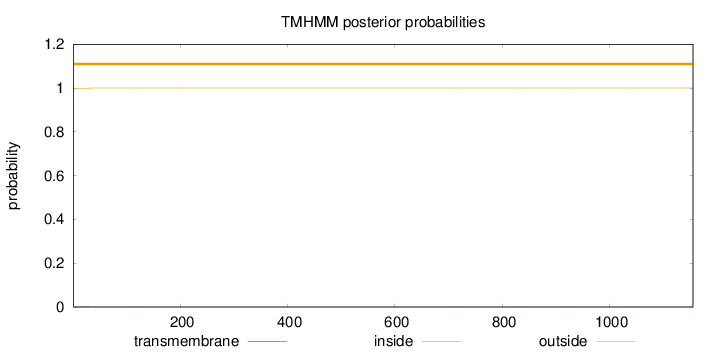
Topology
Subcellular location
Nucleus
Length:
1156
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.02152
Exp number, first 60 AAs:
0.01981
Total prob of N-in:
0.00107
outside
1 - 1156
Population Genetic Test Statistics
Pi
490.659689
Theta
219.433936
Tajima's D
3.898245
CLR
0
CSRT
0.998250087495625
Interpretation
Uncertain
