Gene
KWMTBOMO15851
Pre Gene Modal
BGIBMGA005783
Annotation
PREDICTED:_piggyBac_transposable_element-derived_protein_4-like_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 2.806
Sequence
CDS
ATGTTTAGTCTTGGAGATGAAAACGTGGAAGTGGAAACTTCTCCAGGCCCATCTGGAATTCCACAACGGCCTTCATTGACAGAGCAATTGAAAGTCGCCAAGGTGGCGAGGCTTTTGGATAAAAAGCATGAGAATTTGCTGTCTGCGGACCAAAAAAAGGAGCTGGCCAGGCTGCGCGATGGCAAGCAAGTGGGCAATTTTAAAATGCCAGATGCTCCAGAACCACAAAGAGGAAGAGGAAAGGGAAAGGGGAAGGGCAAAGGAAGGGGCAGTGAAGACTCGCGGATGTGGAGGCCATGCCCACTTTTACCTGGGGTACCAACTGTCCCTCCTCCACGTGTACGTCAGTCGTGGGAATTGCAACAGCAACGCCCGATTGATATCACCGCGGAGCAGGAGGGTGAGGATGATGACATCGAAAGCTATTTTCTTCCAGATATCACCGAGGATGAGGAGATGCGTCCTCCCCTCCTCGAGGATGAGGAGGCCGTTCCTTCTGCAGGGACCCCACCTCAAGCGAAGAAACGCAAGATCCAATTTTCGCCCTCGATGGGGATCATCCAGGCTACTGCATTGCCTCAAAATATCGTGCCAGCAGAGATAGAACTCCTGGGGTTTACACCTTCTTCTACCGGTACTCGAACAGCGCTTGACTCTGGAAGAAGTAATTACAGTATGGATATCGATATAGATATGGAGCGTGACTTCATAGAGCCTGGCGAAGGCAGCGAAGTAGATGAAGGTGATTCGGCGAATATGGTCGGTATGATCGATAATTCTTTGAAACCAAGACGAATCGATGATTTTCCGATGGACTTAGACAGTATTGCTGAACATGAGGTAGTTGAGGTGATACAGGAGCTGGACAATGCTGCCGACGAAGTCCTTCGGAACGTGGTAGGACCGGAAACCGATCTGCTTCATTTTGACTGGCGAGGCGAGCCGGAAAATTTTGAGTCTGTCAGGGAAGCATTTACTGTGCCATCTACTGGACCAACGTTTGATAACGCCGATTTAATTCCCTTAGATGTTTTTTTCAAGATTTGGGATGCAGATATTATTGCTTACATCGTGCGCGAAACCAACCGCTACGGTGCTCAGCTAATAAGGTCGACCATTCCATCCAAGCCAAATTCTAGATTAAGCCGTTGGAAAGACGTCACCAGTGACGAAATCCTACGGTTTTTAGCAGTTTTAATGTTACAGTCACTAGTTATAGATTATGTGGAGCGCGAATATTGGTATGCCGTGATTGAAGAACTCCAAATAGGCAATTTTAAAGAAATAATGACTTATAACCGTTTCATCGTCATCAAACGTTGTCTCCATTTTATAGACAATGCCACTTTGCCCGTGCCGCCTACCAAACTCGATAAAATCATCCCCATCATAGAGCACCTAAACAAAAAATTTAAGTCTTTATACGTACTAGAACAAAATATAGCAATTGACGAGTCACTCCTTCTTTGGAAGGGGCGCCTCTCATTTGCGCAAAAAATTGCTACAAAAAGAGCTCGAGTAGGTATAAAGTCTTACGAGTTATGTGAATCCAGGACTGGATATTTATGGCAGATGGAGGTCTACACGGGCAAAGGGCACGCACATGTGGTACAAGACGGGGAGCCGGAAGAGCGCGGGCAAGAGTCTGACGAGCCGGAGAGTGCCACGGCCCAGATTGTGCTGAACTTAACGAGACCACTTTTTGATAAAGGCCATACATTAATAATGGACAATTTTTATAATGCGCCACTTTTGTCTCGGATCCTAAAAGTTCAGCACAAGACTGATTCGATGGGTACTCTACGACTTAACAGAGAGTTTGTCCCCGAAGCTCTTAAAAAAAAAACAAAAAAAAACATGAAGGAGGGCGAGGTGGCATTCAGCACAACCAAAGACCTCAGTGTGGTTGTGTGGATGGACAAAAATATTGTGGCAATGATTTCTTCTTTCCATCACATAGAAGTCGGCGGCATTGAAAAATACGGTTATTACCGGTACAAACCGCAAGTTGTTCTTGATTATAATTTCTCAATGGGCGGCATCGACCACAAAGATCAAATGCTGTCTGCCTATCCCATTGAGAGATCACGTAACATAATCTGGTACAAAAAATTATTTCGCCGACTTTTAAATGTATCGATGCATAATGCTCTCGTGATGTTCAACCACAACCGCACTCACGCCCTTCGTCACCGTGAATTCCGCCTGCAGTTGGCAAAGCAACTGCTGCAAAGCAGCAGATCAGCAGCACCTTCCCGGTCTTTAGCACCGGCAGCACGACCAGAACCAGCGGCATGCATACGGGACATGCACTTGCCAGGAAAAAATACAAAAAAACAGCGCTGCAAGCTCTGCTACTCTGCCAAAGTGCAGCGCTCGACGGTGTGGCGGTGTACCACCTGCAATGTCAATCTGTGTATCGTAGGATGCTACACCGTATACCATATCAATCTAGCGCAAACGCCCGCCGTTTAG
Protein
MFSLGDENVEVETSPGPSGIPQRPSLTEQLKVAKVARLLDKKHENLLSADQKKELARLRDGKQVGNFKMPDAPEPQRGRGKGKGKGKGRGSEDSRMWRPCPLLPGVPTVPPPRVRQSWELQQQRPIDITAEQEGEDDDIESYFLPDITEDEEMRPPLLEDEEAVPSAGTPPQAKKRKIQFSPSMGIIQATALPQNIVPAEIELLGFTPSSTGTRTALDSGRSNYSMDIDIDMERDFIEPGEGSEVDEGDSANMVGMIDNSLKPRRIDDFPMDLDSIAEHEVVEVIQELDNAADEVLRNVVGPETDLLHFDWRGEPENFESVREAFTVPSTGPTFDNADLIPLDVFFKIWDADIIAYIVRETNRYGAQLIRSTIPSKPNSRLSRWKDVTSDEILRFLAVLMLQSLVIDYVEREYWYAVIEELQIGNFKEIMTYNRFIVIKRCLHFIDNATLPVPPTKLDKIIPIIEHLNKKFKSLYVLEQNIAIDESLLLWKGRLSFAQKIATKRARVGIKSYELCESRTGYLWQMEVYTGKGHAHVVQDGEPEERGQESDEPESATAQIVLNLTRPLFDKGHTLIMDNFYNAPLLSRILKVQHKTDSMGTLRLNREFVPEALKKKTKKNMKEGEVAFSTTKDLSVVVWMDKNIVAMISSFHHIEVGGIEKYGYYRYKPQVVLDYNFSMGGIDHKDQMLSAYPIERSRNIIWYKKLFRRLLNVSMHNALVMFNHNRTHALRHREFRLQLAKQLLQSSRSAAPSRSLAPAARPEPAACIRDMHLPGKNTKKQRCKLCYSAKVQRSTVWRCTTCNVNLCIVGCYTVYHINLAQTPAV
Summary
Uniprot
Pubmed
EMBL
Proteomes
Interpro
SUPFAM
SSF50630
SSF50630
Gene 3D
ProteinModelPortal
Ontologies
GO
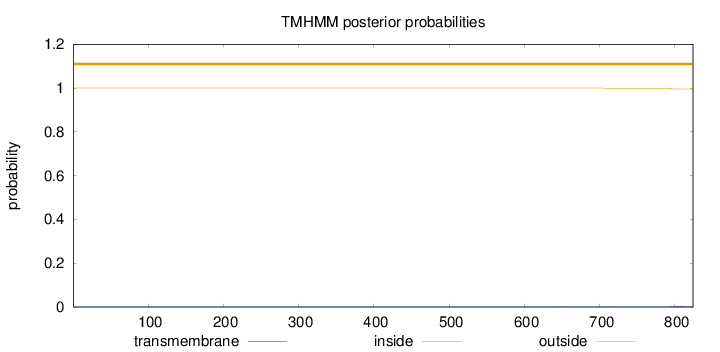
Topology
Length:
824
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.10268
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00006
outside
1 - 824
Population Genetic Test Statistics
Pi
62.438892
Theta
149.776668
Tajima's D
-0.691703
CLR
0.070565
CSRT
0.1974401279936
Interpretation
Uncertain
