Gene
KWMTBOMO15513
Annotation
PREDICTED:_uncharacterized_protein_LOC101747137_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 1.779 Mitochondrial Reliability : 2.23
Sequence
CDS
ATGCACACGTTCACCGATGCTAGCGAGAAGGCCTACGCGTGTGCAGTCTATTGGCGTCAGAAGACAGACGAGAGCACCTACCGCGTCACGCTTCTAGCAGGGAAAGCCAGGGTGACCCCACTGAGACCAGTCTCTATCCCAAGGCTGGAGCTGCAAGCTGCGCTGCTGGGGACAAGGATGGCGCAGGCGATAGCGGACGAATTGGACATCGCAGTCGGTGTAAGGACGTACTGGACTGAGTCCAGTACAGTCCTGACATGGATAAAGACCGACCCACGCACGTTCAAACCTTTCGTCGCGCACCGACTCGCCGAGATAGAAGAGTCAACGAAGCCCCAAGAATGGCGATGGGTGCCTGGCTCGCAGAACCCAGCAGACGACGCAACTAGAGAGGCGCCGGCGGATTTCGACCACACGCATCGGTGGTTTAATGGACCCGAGTTCCTGACGTGGGATGAATCGCGCTGGCCCAAGCCACGAACATTCAAGCAAGAACCGTCTGGAGAGGAGAAAGAGGCCTTCCTGGTCGCCACGGCGAGGACCGCTGACGCTCGATCCACGCCCGATCCGCACAGATTCTCGAGCTGGTCAAAAGCATGGCTGCCGCTTGATCCGCCCCACTTGAAGAAGGCGGAGAAGATCCTACTGAGAAGCAGCCAAGGAGAGAGCTTCGGTGAAAAAGATCCCGAACGTCACCCCAAGTTGCGACGGCTCGACGTCGTGATGGAAGATGGCCTGTTGCGCCTGCGCGGACGTATCGACGCAGCGCAATACATAGATGCAGGCTGCAAGCGACCGATCGTGCTGGACGGGAAGCACGTGATAGCGAGATTATTGATCAAACACTATCACGAGGCGTTCCAGCACGATGCCACAACTCAGTGTGCCTATAAAGCACCTGACCCGCTCGATGGAGATCAAAAATGA
Protein
MHTFTDASEKAYACAVYWRQKTDESTYRVTLLAGKARVTPLRPVSIPRLELQAALLGTRMAQAIADELDIAVGVRTYWTESSTVLTWIKTDPRTFKPFVAHRLAEIEESTKPQEWRWVPGSQNPADDATREAPADFDHTHRWFNGPEFLTWDESRWPKPRTFKQEPSGEEKEAFLVATARTADARSTPDPHRFSSWSKAWLPLDPPHLKKAEKILLRSSQGESFGEKDPERHPKLRRLDVVMEDGLLRLRGRIDAAQYIDAGCKRPIVLDGKHVIARLLIKHYHEAFQHDATTQCAYKAPDPLDGDQK
Summary
Uniprot
Q2MGA5
A0A2A4IRQ3
Q9NDN0
Q9NDM9
A0A3S2N5J7
K7JA41
+ More
A0A226D351 A0A0N1IGH3 A0A226DH22 A0A226DM06 A0A226EKF8 A0A226DGY5 A0A226E2P7 A0A2M4BC51 A0A2M4BC71 A0A226EFR1 A0A2M4B8T5 A0A2M4B8I4 A0A0L7LE26 A0A2M4CVY8 A0A1W7R6A0 A0A182HCB4 A0A1W7R6H4 A0A182HEF9 W8AJC8 A0A226DTU1 W8AYL5 W8B882 A0A1Y1LL65 A0A226D1L8 A0A1Y1LN76 A0A2B4SHN3 A0A182H4A2 Q76C95 Q76C94 A0A146P0F7 A0A226DIJ1 A0A0N5E779 A0A226EEI4 A0A2M4CKZ0 A0A2M4CKI9 A0A182HZZ0 A0A182RI97 A0A182NZW6 A0A1W7R681 A0A182GL35 A0A182WUY2 A0A2M4B9D0 A0A2M4B9E1 A0A2M4B9W9 A0A146T9T9 A0A2B4RNP8 A0A2G8KVJ2 A0A0K3CPX7 A0A2B4RJC8 A0A0N5E6S2 A0A2M4B8T6 A0A0S7EI80 A0A2M4B963 A0A0S7ELN0 A0A355BAS0 A0A085MSJ2 A0A0V0WU59 A0A1W7R6F7 A0A2B4R5Y7 W8AD81 A0A182G158 A0A182HAG0 A0A146Q8H2 A0A2M4B8V2 A0A0V1LJ36 A0A182G1E0 A0A146PPL2 A0A0V0RN93 A0A226EDU9 W4Y2R6 A0A146S3Y9 A0A0J7MNY6 A0A0V1D8V5 W4XEJ9 A0A0V1AQ67 W4XEJ7 A0A0V1AQ61 A0A0V0T6P1 A0A0V1DBM4 A0A0V0V8P2 A0A2H1VMV3 A0A0V1ECM7 A0A1A8MUQ1 J9KLD6 A0A0V1GWL8 A0A085N9I5 A0A1A8V6P1 A0A0N5DT76 A0A0V1GZ07
A0A226D351 A0A0N1IGH3 A0A226DH22 A0A226DM06 A0A226EKF8 A0A226DGY5 A0A226E2P7 A0A2M4BC51 A0A2M4BC71 A0A226EFR1 A0A2M4B8T5 A0A2M4B8I4 A0A0L7LE26 A0A2M4CVY8 A0A1W7R6A0 A0A182HCB4 A0A1W7R6H4 A0A182HEF9 W8AJC8 A0A226DTU1 W8AYL5 W8B882 A0A1Y1LL65 A0A226D1L8 A0A1Y1LN76 A0A2B4SHN3 A0A182H4A2 Q76C95 Q76C94 A0A146P0F7 A0A226DIJ1 A0A0N5E779 A0A226EEI4 A0A2M4CKZ0 A0A2M4CKI9 A0A182HZZ0 A0A182RI97 A0A182NZW6 A0A1W7R681 A0A182GL35 A0A182WUY2 A0A2M4B9D0 A0A2M4B9E1 A0A2M4B9W9 A0A146T9T9 A0A2B4RNP8 A0A2G8KVJ2 A0A0K3CPX7 A0A2B4RJC8 A0A0N5E6S2 A0A2M4B8T6 A0A0S7EI80 A0A2M4B963 A0A0S7ELN0 A0A355BAS0 A0A085MSJ2 A0A0V0WU59 A0A1W7R6F7 A0A2B4R5Y7 W8AD81 A0A182G158 A0A182HAG0 A0A146Q8H2 A0A2M4B8V2 A0A0V1LJ36 A0A182G1E0 A0A146PPL2 A0A0V0RN93 A0A226EDU9 W4Y2R6 A0A146S3Y9 A0A0J7MNY6 A0A0V1D8V5 W4XEJ9 A0A0V1AQ67 W4XEJ7 A0A0V1AQ61 A0A0V0T6P1 A0A0V1DBM4 A0A0V0V8P2 A0A2H1VMV3 A0A0V1ECM7 A0A1A8MUQ1 J9KLD6 A0A0V1GWL8 A0A085N9I5 A0A1A8V6P1 A0A0N5DT76 A0A0V1GZ07
Pubmed
EMBL
AF530470
AAQ09229.1
NWSH01008670
PCG62455.1
AB042118
BAA95569.1
+ More
AB042119 BAA95570.1 RSAL01000375 RVE42106.1 AAZX01023278 LNIX01000039 OXA39294.1 KQ459986 KPJ18861.1 LNIX01000019 OXA44855.1 LNIX01000015 OXA46575.1 LNIX01000003 OXA57597.1 OXA44812.1 LNIX01000007 OXA52015.1 GGFJ01001462 MBW50603.1 GGFJ01001476 MBW50617.1 LNIX01000004 OXA56382.1 GGFJ01000302 MBW49443.1 GGFJ01000218 MBW49359.1 JTDY01001530 KOB73634.1 GGFL01005193 MBW69371.1 GEHC01000947 JAV46698.1 JXUM01127084 KQ567147 KXJ69602.1 GEHC01000914 JAV46731.1 JXUM01005879 KQ560203 KXJ83865.1 GAMC01017930 JAB88625.1 LNIX01000012 OXA48117.1 GAMC01020594 JAB85961.1 GAMC01020598 GAMC01020596 GAMC01020592 JAB85959.1 GEZM01052587 JAV74399.1 LNIX01000041 OXA39119.1 GEZM01052586 JAV74401.1 LSMT01000075 PFX28896.1 JXUM01109099 KQ565290 KXJ71123.1 AB110069 BAD01589.1 BAD01590.1 GCES01148879 JAQ37443.1 LNIX01000018 OXA45362.1 OXA55241.1 GGFL01001842 MBW66020.1 GGFL01001682 MBW65860.1 APCN01002024 GEHC01000967 JAV46678.1 JXUM01013493 KQ560378 KXJ82704.1 GGFJ01000471 MBW49612.1 GGFJ01000470 MBW49611.1 GGFJ01000487 MBW49628.1 GCES01096631 JAQ89691.1 LSMT01000392 PFX18796.1 MRZV01000348 PIK51982.1 LN877231 CTR11689.1 LSMT01000554 PFX16342.1 GGFJ01000306 MBW49447.1 GBYX01477451 JAO04236.1 GGFJ01000432 MBW49573.1 GBYX01477452 JAO04235.1 DNYX01000180 HBK83090.1 KL367687 KFD60188.1 JYDK01000056 KRX79229.1 GEHC01000945 JAV46700.1 LSMT01001506 PFX12229.1 GAMC01020600 JAB85955.1 JXUM01136032 KQ568367 KXJ69026.1 JXUM01031097 KQ560909 KXJ80382.1 GCES01133947 JAQ52375.1 GGFJ01000333 MBW49474.1 JYDW01000048 KRZ59054.1 JXUM01038372 KQ561162 KXJ79503.1 GCES01140786 JAQ45536.1 JYDL01000119 KRX15895.1 OXA55408.1 AAGJ04043389 GCES01111356 JAQ74966.1 LBMM01026421 KMQ82315.1 JYDI01000031 KRY57459.1 AAGJ04115911 JYDH01000309 KRY26933.1 KRY26929.1 JYDJ01000536 KRX34640.1 JYDI01000018 KRY58742.1 JYDN01000069 KRX59701.1 ODYU01003174 SOQ41584.1 JYDR01000056 KRY71589.1 HAEF01019456 SBR60615.1 ABLF02013940 ABLF02036313 ABLF02066938 ABLF02066950 JYDP01000218 KRZ02721.1 KL367527 KFD66131.1 HAEJ01014775 SBS55232.1 JYDS01000532 KRZ03382.1
AB042119 BAA95570.1 RSAL01000375 RVE42106.1 AAZX01023278 LNIX01000039 OXA39294.1 KQ459986 KPJ18861.1 LNIX01000019 OXA44855.1 LNIX01000015 OXA46575.1 LNIX01000003 OXA57597.1 OXA44812.1 LNIX01000007 OXA52015.1 GGFJ01001462 MBW50603.1 GGFJ01001476 MBW50617.1 LNIX01000004 OXA56382.1 GGFJ01000302 MBW49443.1 GGFJ01000218 MBW49359.1 JTDY01001530 KOB73634.1 GGFL01005193 MBW69371.1 GEHC01000947 JAV46698.1 JXUM01127084 KQ567147 KXJ69602.1 GEHC01000914 JAV46731.1 JXUM01005879 KQ560203 KXJ83865.1 GAMC01017930 JAB88625.1 LNIX01000012 OXA48117.1 GAMC01020594 JAB85961.1 GAMC01020598 GAMC01020596 GAMC01020592 JAB85959.1 GEZM01052587 JAV74399.1 LNIX01000041 OXA39119.1 GEZM01052586 JAV74401.1 LSMT01000075 PFX28896.1 JXUM01109099 KQ565290 KXJ71123.1 AB110069 BAD01589.1 BAD01590.1 GCES01148879 JAQ37443.1 LNIX01000018 OXA45362.1 OXA55241.1 GGFL01001842 MBW66020.1 GGFL01001682 MBW65860.1 APCN01002024 GEHC01000967 JAV46678.1 JXUM01013493 KQ560378 KXJ82704.1 GGFJ01000471 MBW49612.1 GGFJ01000470 MBW49611.1 GGFJ01000487 MBW49628.1 GCES01096631 JAQ89691.1 LSMT01000392 PFX18796.1 MRZV01000348 PIK51982.1 LN877231 CTR11689.1 LSMT01000554 PFX16342.1 GGFJ01000306 MBW49447.1 GBYX01477451 JAO04236.1 GGFJ01000432 MBW49573.1 GBYX01477452 JAO04235.1 DNYX01000180 HBK83090.1 KL367687 KFD60188.1 JYDK01000056 KRX79229.1 GEHC01000945 JAV46700.1 LSMT01001506 PFX12229.1 GAMC01020600 JAB85955.1 JXUM01136032 KQ568367 KXJ69026.1 JXUM01031097 KQ560909 KXJ80382.1 GCES01133947 JAQ52375.1 GGFJ01000333 MBW49474.1 JYDW01000048 KRZ59054.1 JXUM01038372 KQ561162 KXJ79503.1 GCES01140786 JAQ45536.1 JYDL01000119 KRX15895.1 OXA55408.1 AAGJ04043389 GCES01111356 JAQ74966.1 LBMM01026421 KMQ82315.1 JYDI01000031 KRY57459.1 AAGJ04115911 JYDH01000309 KRY26933.1 KRY26929.1 JYDJ01000536 KRX34640.1 JYDI01000018 KRY58742.1 JYDN01000069 KRX59701.1 ODYU01003174 SOQ41584.1 JYDR01000056 KRY71589.1 HAEF01019456 SBR60615.1 ABLF02013940 ABLF02036313 ABLF02066938 ABLF02066950 JYDP01000218 KRZ02721.1 KL367527 KFD66131.1 HAEJ01014775 SBS55232.1 JYDS01000532 KRZ03382.1
Proteomes
UP000218220
UP000283053
UP000002358
UP000198287
UP000053240
UP000037510
+ More
UP000069940 UP000249989 UP000225706 UP000046395 UP000075840 UP000075900 UP000075885 UP000076407 UP000230750 UP000054673 UP000054721 UP000054630 UP000007110 UP000036403 UP000054653 UP000054776 UP000055048 UP000054681 UP000054632 UP000007819 UP000055024 UP000054805
UP000069940 UP000249989 UP000225706 UP000046395 UP000075840 UP000075900 UP000075885 UP000076407 UP000230750 UP000054673 UP000054721 UP000054630 UP000007110 UP000036403 UP000054653 UP000054776 UP000055048 UP000054681 UP000054632 UP000007819 UP000055024 UP000054805
Pfam
Interpro
IPR040676
DUF5641
+ More
IPR036397 RNaseH_sf
IPR005312 DUF1759
IPR001584 Integrase_cat-core
IPR008042 Retrotrans_Pao
IPR012337 RNaseH-like_sf
IPR041588 Integrase_H2C2
IPR001878 Znf_CCHC
IPR008737 Peptidase_asp_put
IPR001995 Peptidase_A2_cat
IPR006094 Oxid_FAD_bind_N
IPR016166 FAD-bd_PCMH
IPR016164 FAD-linked_Oxase-like_C
IPR016170 Cytok_DH_C_sf
IPR015213 Cholesterol_OX_subst-bd
IPR036318 FAD-bd_PCMH-like_sf
IPR016169 FAD-bd_PCMH_sub2
IPR016167 FAD-bd_PCMH_sub1
IPR019787 Znf_PHD-finger
IPR001965 Znf_PHD
IPR013083 Znf_RING/FYVE/PHD
IPR011011 Znf_FYVE_PHD
IPR019786 Zinc_finger_PHD-type_CS
IPR000253 FHA_dom
IPR000477 RT_dom
IPR036875 Znf_CCHC_sf
IPR001969 Aspartic_peptidase_AS
IPR021109 Peptidase_aspartic_dom_sf
IPR002999 Tudor
IPR036397 RNaseH_sf
IPR005312 DUF1759
IPR001584 Integrase_cat-core
IPR008042 Retrotrans_Pao
IPR012337 RNaseH-like_sf
IPR041588 Integrase_H2C2
IPR001878 Znf_CCHC
IPR008737 Peptidase_asp_put
IPR001995 Peptidase_A2_cat
IPR006094 Oxid_FAD_bind_N
IPR016166 FAD-bd_PCMH
IPR016164 FAD-linked_Oxase-like_C
IPR016170 Cytok_DH_C_sf
IPR015213 Cholesterol_OX_subst-bd
IPR036318 FAD-bd_PCMH-like_sf
IPR016169 FAD-bd_PCMH_sub2
IPR016167 FAD-bd_PCMH_sub1
IPR019787 Znf_PHD-finger
IPR001965 Znf_PHD
IPR013083 Znf_RING/FYVE/PHD
IPR011011 Znf_FYVE_PHD
IPR019786 Zinc_finger_PHD-type_CS
IPR000253 FHA_dom
IPR000477 RT_dom
IPR036875 Znf_CCHC_sf
IPR001969 Aspartic_peptidase_AS
IPR021109 Peptidase_aspartic_dom_sf
IPR002999 Tudor
SUPFAM
ProteinModelPortal
Q2MGA5
A0A2A4IRQ3
Q9NDN0
Q9NDM9
A0A3S2N5J7
K7JA41
+ More
A0A226D351 A0A0N1IGH3 A0A226DH22 A0A226DM06 A0A226EKF8 A0A226DGY5 A0A226E2P7 A0A2M4BC51 A0A2M4BC71 A0A226EFR1 A0A2M4B8T5 A0A2M4B8I4 A0A0L7LE26 A0A2M4CVY8 A0A1W7R6A0 A0A182HCB4 A0A1W7R6H4 A0A182HEF9 W8AJC8 A0A226DTU1 W8AYL5 W8B882 A0A1Y1LL65 A0A226D1L8 A0A1Y1LN76 A0A2B4SHN3 A0A182H4A2 Q76C95 Q76C94 A0A146P0F7 A0A226DIJ1 A0A0N5E779 A0A226EEI4 A0A2M4CKZ0 A0A2M4CKI9 A0A182HZZ0 A0A182RI97 A0A182NZW6 A0A1W7R681 A0A182GL35 A0A182WUY2 A0A2M4B9D0 A0A2M4B9E1 A0A2M4B9W9 A0A146T9T9 A0A2B4RNP8 A0A2G8KVJ2 A0A0K3CPX7 A0A2B4RJC8 A0A0N5E6S2 A0A2M4B8T6 A0A0S7EI80 A0A2M4B963 A0A0S7ELN0 A0A355BAS0 A0A085MSJ2 A0A0V0WU59 A0A1W7R6F7 A0A2B4R5Y7 W8AD81 A0A182G158 A0A182HAG0 A0A146Q8H2 A0A2M4B8V2 A0A0V1LJ36 A0A182G1E0 A0A146PPL2 A0A0V0RN93 A0A226EDU9 W4Y2R6 A0A146S3Y9 A0A0J7MNY6 A0A0V1D8V5 W4XEJ9 A0A0V1AQ67 W4XEJ7 A0A0V1AQ61 A0A0V0T6P1 A0A0V1DBM4 A0A0V0V8P2 A0A2H1VMV3 A0A0V1ECM7 A0A1A8MUQ1 J9KLD6 A0A0V1GWL8 A0A085N9I5 A0A1A8V6P1 A0A0N5DT76 A0A0V1GZ07
A0A226D351 A0A0N1IGH3 A0A226DH22 A0A226DM06 A0A226EKF8 A0A226DGY5 A0A226E2P7 A0A2M4BC51 A0A2M4BC71 A0A226EFR1 A0A2M4B8T5 A0A2M4B8I4 A0A0L7LE26 A0A2M4CVY8 A0A1W7R6A0 A0A182HCB4 A0A1W7R6H4 A0A182HEF9 W8AJC8 A0A226DTU1 W8AYL5 W8B882 A0A1Y1LL65 A0A226D1L8 A0A1Y1LN76 A0A2B4SHN3 A0A182H4A2 Q76C95 Q76C94 A0A146P0F7 A0A226DIJ1 A0A0N5E779 A0A226EEI4 A0A2M4CKZ0 A0A2M4CKI9 A0A182HZZ0 A0A182RI97 A0A182NZW6 A0A1W7R681 A0A182GL35 A0A182WUY2 A0A2M4B9D0 A0A2M4B9E1 A0A2M4B9W9 A0A146T9T9 A0A2B4RNP8 A0A2G8KVJ2 A0A0K3CPX7 A0A2B4RJC8 A0A0N5E6S2 A0A2M4B8T6 A0A0S7EI80 A0A2M4B963 A0A0S7ELN0 A0A355BAS0 A0A085MSJ2 A0A0V0WU59 A0A1W7R6F7 A0A2B4R5Y7 W8AD81 A0A182G158 A0A182HAG0 A0A146Q8H2 A0A2M4B8V2 A0A0V1LJ36 A0A182G1E0 A0A146PPL2 A0A0V0RN93 A0A226EDU9 W4Y2R6 A0A146S3Y9 A0A0J7MNY6 A0A0V1D8V5 W4XEJ9 A0A0V1AQ67 W4XEJ7 A0A0V1AQ61 A0A0V0T6P1 A0A0V1DBM4 A0A0V0V8P2 A0A2H1VMV3 A0A0V1ECM7 A0A1A8MUQ1 J9KLD6 A0A0V1GWL8 A0A085N9I5 A0A1A8V6P1 A0A0N5DT76 A0A0V1GZ07
Ontologies
GO
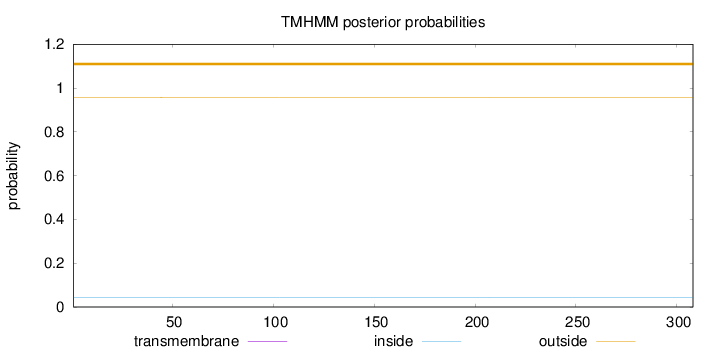
Topology
Length:
308
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00129
Exp number, first 60 AAs:
0.00057
Total prob of N-in:
0.04240
outside
1 - 308
Population Genetic Test Statistics
Pi
304.796096
Theta
175.611446
Tajima's D
2.060706
CLR
0.387904
CSRT
0.892405379731013
Interpretation
Uncertain
