Gene
KWMTBOMO15492 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA010763
Annotation
aminopeptidase_N_precursor_[Bombyx_mori]
Full name
Aminopeptidase
Location in the cell
PlasmaMembrane Reliability : 1.328
Sequence
CDS
ATGGCGTGCTTGCATTTTATTTTATTGTTGTCCTGTGCGCTGCTGTCCACGGGCGAATACCTGCTGCCCGGCGATGTGGTGCCGACATTCTACGACGTGCTCCTCATTTACGACGTGGATCCGGCGACGAACTTCAGCTACTTCGGCCGTGTCGACATTAAATTAAACATCCTCAAACCGACCTCAAAGATCGTGCTCCACGCGCAAGGCTTCAGCATACCGGAAGAGGAAGTGACCCTGACAGGGCCCAAGGAGGTGGCAGTTGACAATGTCAAACTGAACGACACGTTCAATTTATTGACACTGTCGCTGAGTCAGCAGTTGGATGAAGGCGACAACTACGTGCTGCGAATACCTTTCTACGGCAATCTGCAGCAAGATCTGGATGGGACCTACATCAGTAAATACGTCGATAAGAAAACAAAGAAAAATGAATATTTGGTGACGACGCAATTCGAGGCGATATCAGCTAGAAAAGGTTTCCCGTGCTTTGACGAACCGATGTACAAGGCGCCCTTCAAGATATCCATCGGCCATCACATCAATTATGTTGCCGTTTCCAACATGCCTCTAGTGTCGACGGATAATTATAATGTCCTGGAGGAAACATGGCCTTGGAAAACTATCGGACAAAAGTTCAATAAAGCCGTTTACGATTTCGTCTGGGACCACTTCGACGTGTCCCTGTCAATGTCAACGTATCTCGTCGCTTACGTCATTTCGAAGTTCACATACGAGGAGAGCCCGCCCGAGCTGTCCGACACTAAGTTCAGGATCTGGGCTAACAAGGACACTATTGACCAGGCCGCCTACGCTAAGTCGATAGCGCCAAAAGTGCTGACACACTTCGAGAAGTGGTTCGACGTGAAATACCCGCTGCCTAAACAGGACATGATCGCCATTCCGGACTTCGCGGCCGGCGCCATGGAGAACTGGGGCCTCATAACCTACAGGGAGACCACCTTGCTCTACGATAAGAAGCACTCCTCGTTCTTGAACAAGGAAAGAGTAGCAGAGGTGATAGCGCACGAGCTGGCCCACCAGTGGTTCGGGAACCTGGTCACGATGAAGTGGTGGTCCGACCTGTGGCTGAACGAGGGGTTCGCGACGTACGCGGCGTCCGTGGGCGTGGCGGCCGTGGAGCCCACGTGGCGCGCGGACAACAACTACGCCGTCGACAACATGCTGGCCGTGCTCAACCTCGACGCCTTGGAGTCTTCGCACCCTGTGTCGGTCACAATAGACGACCCGAAGCGCATCTTCGAGATATTCGATGAGATCTCGTACAAGAAGGGCTCCACACTCATCAGGATGATGGTCATGTTCCTCGGAGAAGACGTCTTCAGGAAAGCCATTAAAAATTACCTCGTGAAGTACACGTTGAAGAACGCCGAACAAGATGACCTGTGGCACGAGCTCACTGAAGAGAGTAAGAGGCAGGGAGGGATCGCGAACAACGTGACCATCAAGGAGATAATGGACACGTGGACCACGCAAACCGGATACCCGATCCTGACCGTCACCAGGGATTATATAGACAAGTCTGTCAGCGTCACTCAGCGGCGGTACTTGTCGCTGGGGGCGCGCGCGGCGTCGCACCGCTCGTGGTGGGTGCCACTGCGCGTGGCGGCCGCGGGCGAGCAGCCCGAGCCCTCGCTGCACTGGCTCGCGACGCACCAGGGCCTGGACGAGCCCTACCGCTTCCCGCACGACGCAGAGCCCACCGACTGGATGCTCTTCAATCCCGATATGATCGCACCTTATCGCGTCAATTATGATGCTCGTAACTGGGAACTTCTTTCGGAGACACTGCTCAGCGACAAGTACGATGAGATCCCAGTACTGGGCAGAGTGCAACTGATATCGGACGCCTTCGCTCTGGCGGCCACCAACAGACTCAACTATTCCACTACACTGATTCTTGCAAGCTACCTCCAGAGAGAGAGGGAGTACCTGCCTCTGTCGACAGCGCTGAGAGGGTTGGCCAAGATTGAGAACCTCTTGAAGCGCAGCGGAGACTACAGCGCCTTCCAGAGCTACATCAGGCATCTGGTGCAGAACTCGTATCGAAGGGCTGGAGGCCTCTCCGTCAAGCGTATCGTTAATGGCGACGATCTGAACAGCGTAAAACTGCAGGTGTTGACCAGCGGCTGGGCCTGTCGCGCGCTGATACCGGGCTGCGAGGAAAACGCCATGGAGCTGTTCCAAATGTGGATGAACACTTCCAATCCGGATGAAAATAATCCGCAAGTTTTCTTTATTTTTTCATCTAAGGACACGATCGATGGGGATCACAAGTCGATGTCTCAATTTAAAACGACCCGCGCGGGGATGAAGATCCCGCTGGACCTGCGGCGCACGGTGTGGTGCGCGGGCGCGGCGCGCGGCGGCGAGCGCGAGTGGCGCTTCCTCACGCTGCGGCTGCAGGCCGCGCGCGTTGCCGCCGCCCGCGACGCCATGCTCGCCGCCATGGCCTGCACCAGGCAGGTCTGGCTGCTCAACCAGTTCCTCGAATGGGCGGTGTCGGAGAACTCCCCGGTGCGCAAGCAGGACGCGGCCGCCGTGGTCGGTAACGTGGCCCGCACGCCGGTCGGCTACTACGTCGCGCGGGAGTTCTTCTACAACCGAATCGACGATCTGTTTAAGACTTTCAACGGTCAGAGCCGCCGTTTCGGCTATATCATCAAGACTCTGCTGGACCAGTTCACGACTCAGAGAGAATTGGACGAGTTTCTCATTTTCCACGACAAGAACCCTAAATATTTCGAGGAGACGAAGCTGGCCGTCGTTCAGGCGATTGAGAAAGCCAAAGTGAACATCGACTGGGTCGACCAAAACAAAGATTTAGTTATGAGTGAATAA
Protein
MACLHFILLLSCALLSTGEYLLPGDVVPTFYDVLLIYDVDPATNFSYFGRVDIKLNILKPTSKIVLHAQGFSIPEEEVTLTGPKEVAVDNVKLNDTFNLLTLSLSQQLDEGDNYVLRIPFYGNLQQDLDGTYISKYVDKKTKKNEYLVTTQFEAISARKGFPCFDEPMYKAPFKISIGHHINYVAVSNMPLVSTDNYNVLEETWPWKTIGQKFNKAVYDFVWDHFDVSLSMSTYLVAYVISKFTYEESPPELSDTKFRIWANKDTIDQAAYAKSIAPKVLTHFEKWFDVKYPLPKQDMIAIPDFAAGAMENWGLITYRETTLLYDKKHSSFLNKERVAEVIAHELAHQWFGNLVTMKWWSDLWLNEGFATYAASVGVAAVEPTWRADNNYAVDNMLAVLNLDALESSHPVSVTIDDPKRIFEIFDEISYKKGSTLIRMMVMFLGEDVFRKAIKNYLVKYTLKNAEQDDLWHELTEESKRQGGIANNVTIKEIMDTWTTQTGYPILTVTRDYIDKSVSVTQRRYLSLGARAASHRSWWVPLRVAAAGEQPEPSLHWLATHQGLDEPYRFPHDAEPTDWMLFNPDMIAPYRVNYDARNWELLSETLLSDKYDEIPVLGRVQLISDAFALAATNRLNYSTTLILASYLQREREYLPLSTALRGLAKIENLLKRSGDYSAFQSYIRHLVQNSYRRAGGLSVKRIVNGDDLNSVKLQVLTSGWACRALIPGCEENAMELFQMWMNTSNPDENNPQVFFIFSSKDTIDGDHKSMSQFKTTRAGMKIPLDLRRTVWCAGAARGGEREWRFLTLRLQAARVAAARDAMLAAMACTRQVWLLNQFLEWAVSENSPVRKQDAAAVVGNVARTPVGYYVAREFFYNRIDDLFKTFNGQSRRFGYIIKTLLDQFTTQRELDEFLIFHDKNPKYFEETKLAVVQAIEKAKVNIDWVDQNKDLVMSE
Summary
Cofactor
Zn(2+)
Similarity
Belongs to the peptidase M1 family.
Feature
chain Aminopeptidase
Uniprot
I3VR83
A0A220K8P2
A0A2W1BT37
A0A212EYJ4
A0A0N1IFL5
A0A194PRA0
+ More
A0A2A4JS21 A0A2H1VHB7 D6WB37 A0A1I9WL50 A0A2L1IQ82 A0A1B6D6B8 A0A1B6HB93 A0A1Y1LUJ7 A0A2H8TQM9 E0VS66 N6TGU3 X1X0P5 A0A0K8V993 A0A0A1X3N6 A0A034VBT7 A0A0K8U897 A0A1J1IVX7 A0A1J1IZ42 W8BKE0 U5EU86 A0A1B6LLZ7 A0A182QC88 Q16MQ9 A0A224XI48 A0A1W4XNR6 A0A182GQP1 A0A182Y4U1 B3MTC9 A0A182FM81 A0A182RM22 B4JYJ5 B4K5Y1 A0A084W9N5 A0A182V1G0 A0A0Q9XBS5 A0A182SPU1 J9KGQ7 A0A182WY28 A0A2K8JME3 Q7Q4T0 A0A1W4U620 A0A336M458 A0A3B0JKK2 W5JLT5 J9JLW8 A0A0Q9WV74 B4NFN7 A0A182VV47 B4QZJ5 Q29BX5 A0A1Q3FSF6 Q9NH67 A0A023F109 B4HZ87 Q0KI00 B4MBV1 B4GPI5 A0A182K329 B3P5I4 A0A0M4F678 B4PQ42 A0A336KKE2 Q6IDH1 B3MTC8 A0A0N7Z8X1 A0A182PL06 A0A069DXR4 A0A0R1E4S0 A0A182N0H6 A0A1W4UH69 A0A224X6V4 A0A1B6GC72 A0A1W4UH28 B4PQ46 B3P5I7 A0A0V0G587 B3P5I5 A0A182TL54 A0A0T6AUS1 A0A0L0BPV0 B4QZJ4 A0A1I8N5N4 A0A0R1E4R4 B4PQ45 A0A0R1E9Q7 B4PQ43 B4QZJ3 A0A1I8PCE0 E2BB79 A0A182HMT5 B0X825 K7INE5
A0A2A4JS21 A0A2H1VHB7 D6WB37 A0A1I9WL50 A0A2L1IQ82 A0A1B6D6B8 A0A1B6HB93 A0A1Y1LUJ7 A0A2H8TQM9 E0VS66 N6TGU3 X1X0P5 A0A0K8V993 A0A0A1X3N6 A0A034VBT7 A0A0K8U897 A0A1J1IVX7 A0A1J1IZ42 W8BKE0 U5EU86 A0A1B6LLZ7 A0A182QC88 Q16MQ9 A0A224XI48 A0A1W4XNR6 A0A182GQP1 A0A182Y4U1 B3MTC9 A0A182FM81 A0A182RM22 B4JYJ5 B4K5Y1 A0A084W9N5 A0A182V1G0 A0A0Q9XBS5 A0A182SPU1 J9KGQ7 A0A182WY28 A0A2K8JME3 Q7Q4T0 A0A1W4U620 A0A336M458 A0A3B0JKK2 W5JLT5 J9JLW8 A0A0Q9WV74 B4NFN7 A0A182VV47 B4QZJ5 Q29BX5 A0A1Q3FSF6 Q9NH67 A0A023F109 B4HZ87 Q0KI00 B4MBV1 B4GPI5 A0A182K329 B3P5I4 A0A0M4F678 B4PQ42 A0A336KKE2 Q6IDH1 B3MTC8 A0A0N7Z8X1 A0A182PL06 A0A069DXR4 A0A0R1E4S0 A0A182N0H6 A0A1W4UH69 A0A224X6V4 A0A1B6GC72 A0A1W4UH28 B4PQ46 B3P5I7 A0A0V0G587 B3P5I5 A0A182TL54 A0A0T6AUS1 A0A0L0BPV0 B4QZJ4 A0A1I8N5N4 A0A0R1E4R4 B4PQ45 A0A0R1E9Q7 B4PQ43 B4QZJ3 A0A1I8PCE0 E2BB79 A0A182HMT5 B0X825 K7INE5
EC Number
3.4.11.-
Pubmed
28630352
28756777
22118469
26354079
18362917
19820115
+ More
27538518 29415259 28004739 20566863 23537049 25830018 25348373 24495485 17510324 26483478 25244985 17994087 24438588 12364791 14747013 17210077 20920257 23761445 15632085 25474469 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 17550304 27129103 26334808 26108605 25315136 20798317 20075255
27538518 29415259 28004739 20566863 23537049 25830018 25348373 24495485 17510324 26483478 25244985 17994087 24438588 12364791 14747013 17210077 20920257 23761445 15632085 25474469 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 17550304 27129103 26334808 26108605 25315136 20798317 20075255
EMBL
JQ061154
AFK85028.1
MF319742
ASJ26456.1
KZ149908
PZC78228.1
+ More
AGBW02011488 OWR46566.1 KQ461003 KPJ10021.1 KQ459601 KPI93655.1 NWSH01000721 PCG74619.1 ODYU01002553 SOQ40207.1 KQ971311 EEZ99336.1 KU932234 APA33870.1 MF741655 AVD96938.1 GEDC01016064 JAS21234.1 GECU01035837 JAS71869.1 GEZM01046358 GEZM01046357 JAV77303.1 GFXV01004127 MBW15932.1 DS235745 EEB16222.1 APGK01027780 APGK01027781 APGK01027782 APGK01027783 APGK01027784 APGK01027785 KB740671 ENN79629.1 ABLF02020109 GDHF01016887 JAI35427.1 GBXI01008343 JAD05949.1 GAKP01019737 JAC39215.1 GDHF01029538 JAI22776.1 CVRI01000060 CRL03858.1 CRL03857.1 GAMC01009092 JAB97463.1 GANO01002430 JAB57441.1 GEBQ01015299 JAT24678.1 AXCN02001411 CH477852 EAT35628.1 GFTR01008306 JAW08120.1 JXUM01080917 KQ563216 KXJ74267.1 CH902623 EDV30519.1 CH916377 EDV90757.1 CH933806 EDW16218.2 ATLV01021848 KE525324 KFB46929.1 KRG01998.1 ABLF02013994 KY031093 ATU82844.1 AAAB01008963 EAA12046.5 UFQS01000417 UFQT01000417 SSX03766.1 SSX24131.1 OUUW01000005 SPP81333.1 ADMH02001160 ETN63860.1 CH964251 KRF99732.1 EDW83104.2 CM000364 EDX14833.1 CM000070 EAL26872.1 GFDL01004639 JAV30406.1 AF231040 AAF34809.1 GBBI01003749 JAC14963.1 CH480819 EDW53344.1 AE014297 BT133378 AAF56884.1 AFH36363.1 CH940656 EDW58572.1 CH479186 EDW39068.1 CH954182 EDV53234.1 CP012526 ALC47157.1 CM000160 EDW98311.1 SSX03765.1 SSX24130.1 BT014634 AAT27258.1 EDV30518.1 GDKW01002163 JAI54432.1 GBGD01000392 JAC88497.1 KRK04205.1 GFTR01008270 JAW08156.1 GECZ01009731 JAS60038.1 EDW98315.1 EDV53237.1 GECL01003570 JAP02554.1 EDV53235.1 LJIG01022766 KRT78825.1 JRES01001565 KNC22036.1 EDX14832.1 KRK04208.1 EDW98314.1 KRK04210.1 EDW98312.1 EDX14831.1 GL446946 EFN87052.1 APCN01004903 DS232472 EDS42294.1
AGBW02011488 OWR46566.1 KQ461003 KPJ10021.1 KQ459601 KPI93655.1 NWSH01000721 PCG74619.1 ODYU01002553 SOQ40207.1 KQ971311 EEZ99336.1 KU932234 APA33870.1 MF741655 AVD96938.1 GEDC01016064 JAS21234.1 GECU01035837 JAS71869.1 GEZM01046358 GEZM01046357 JAV77303.1 GFXV01004127 MBW15932.1 DS235745 EEB16222.1 APGK01027780 APGK01027781 APGK01027782 APGK01027783 APGK01027784 APGK01027785 KB740671 ENN79629.1 ABLF02020109 GDHF01016887 JAI35427.1 GBXI01008343 JAD05949.1 GAKP01019737 JAC39215.1 GDHF01029538 JAI22776.1 CVRI01000060 CRL03858.1 CRL03857.1 GAMC01009092 JAB97463.1 GANO01002430 JAB57441.1 GEBQ01015299 JAT24678.1 AXCN02001411 CH477852 EAT35628.1 GFTR01008306 JAW08120.1 JXUM01080917 KQ563216 KXJ74267.1 CH902623 EDV30519.1 CH916377 EDV90757.1 CH933806 EDW16218.2 ATLV01021848 KE525324 KFB46929.1 KRG01998.1 ABLF02013994 KY031093 ATU82844.1 AAAB01008963 EAA12046.5 UFQS01000417 UFQT01000417 SSX03766.1 SSX24131.1 OUUW01000005 SPP81333.1 ADMH02001160 ETN63860.1 CH964251 KRF99732.1 EDW83104.2 CM000364 EDX14833.1 CM000070 EAL26872.1 GFDL01004639 JAV30406.1 AF231040 AAF34809.1 GBBI01003749 JAC14963.1 CH480819 EDW53344.1 AE014297 BT133378 AAF56884.1 AFH36363.1 CH940656 EDW58572.1 CH479186 EDW39068.1 CH954182 EDV53234.1 CP012526 ALC47157.1 CM000160 EDW98311.1 SSX03765.1 SSX24130.1 BT014634 AAT27258.1 EDV30518.1 GDKW01002163 JAI54432.1 GBGD01000392 JAC88497.1 KRK04205.1 GFTR01008270 JAW08156.1 GECZ01009731 JAS60038.1 EDW98315.1 EDV53237.1 GECL01003570 JAP02554.1 EDV53235.1 LJIG01022766 KRT78825.1 JRES01001565 KNC22036.1 EDX14832.1 KRK04208.1 EDW98314.1 KRK04210.1 EDW98312.1 EDX14831.1 GL446946 EFN87052.1 APCN01004903 DS232472 EDS42294.1
Proteomes
UP000007151
UP000053240
UP000053268
UP000218220
UP000007266
UP000009046
+ More
UP000019118 UP000007819 UP000183832 UP000075886 UP000008820 UP000192223 UP000069940 UP000249989 UP000076408 UP000007801 UP000069272 UP000075900 UP000001070 UP000009192 UP000030765 UP000075903 UP000075901 UP000076407 UP000007062 UP000192221 UP000268350 UP000000673 UP000007798 UP000075920 UP000000304 UP000001819 UP000001292 UP000000803 UP000008792 UP000008744 UP000075881 UP000008711 UP000092553 UP000002282 UP000075885 UP000075884 UP000075902 UP000037069 UP000095301 UP000095300 UP000008237 UP000075840 UP000002320 UP000002358
UP000019118 UP000007819 UP000183832 UP000075886 UP000008820 UP000192223 UP000069940 UP000249989 UP000076408 UP000007801 UP000069272 UP000075900 UP000001070 UP000009192 UP000030765 UP000075903 UP000075901 UP000076407 UP000007062 UP000192221 UP000268350 UP000000673 UP000007798 UP000075920 UP000000304 UP000001819 UP000001292 UP000000803 UP000008792 UP000008744 UP000075881 UP000008711 UP000092553 UP000002282 UP000075885 UP000075884 UP000075902 UP000037069 UP000095301 UP000095300 UP000008237 UP000075840 UP000002320 UP000002358
Interpro
Gene 3D
CDD
ProteinModelPortal
I3VR83
A0A220K8P2
A0A2W1BT37
A0A212EYJ4
A0A0N1IFL5
A0A194PRA0
+ More
A0A2A4JS21 A0A2H1VHB7 D6WB37 A0A1I9WL50 A0A2L1IQ82 A0A1B6D6B8 A0A1B6HB93 A0A1Y1LUJ7 A0A2H8TQM9 E0VS66 N6TGU3 X1X0P5 A0A0K8V993 A0A0A1X3N6 A0A034VBT7 A0A0K8U897 A0A1J1IVX7 A0A1J1IZ42 W8BKE0 U5EU86 A0A1B6LLZ7 A0A182QC88 Q16MQ9 A0A224XI48 A0A1W4XNR6 A0A182GQP1 A0A182Y4U1 B3MTC9 A0A182FM81 A0A182RM22 B4JYJ5 B4K5Y1 A0A084W9N5 A0A182V1G0 A0A0Q9XBS5 A0A182SPU1 J9KGQ7 A0A182WY28 A0A2K8JME3 Q7Q4T0 A0A1W4U620 A0A336M458 A0A3B0JKK2 W5JLT5 J9JLW8 A0A0Q9WV74 B4NFN7 A0A182VV47 B4QZJ5 Q29BX5 A0A1Q3FSF6 Q9NH67 A0A023F109 B4HZ87 Q0KI00 B4MBV1 B4GPI5 A0A182K329 B3P5I4 A0A0M4F678 B4PQ42 A0A336KKE2 Q6IDH1 B3MTC8 A0A0N7Z8X1 A0A182PL06 A0A069DXR4 A0A0R1E4S0 A0A182N0H6 A0A1W4UH69 A0A224X6V4 A0A1B6GC72 A0A1W4UH28 B4PQ46 B3P5I7 A0A0V0G587 B3P5I5 A0A182TL54 A0A0T6AUS1 A0A0L0BPV0 B4QZJ4 A0A1I8N5N4 A0A0R1E4R4 B4PQ45 A0A0R1E9Q7 B4PQ43 B4QZJ3 A0A1I8PCE0 E2BB79 A0A182HMT5 B0X825 K7INE5
A0A2A4JS21 A0A2H1VHB7 D6WB37 A0A1I9WL50 A0A2L1IQ82 A0A1B6D6B8 A0A1B6HB93 A0A1Y1LUJ7 A0A2H8TQM9 E0VS66 N6TGU3 X1X0P5 A0A0K8V993 A0A0A1X3N6 A0A034VBT7 A0A0K8U897 A0A1J1IVX7 A0A1J1IZ42 W8BKE0 U5EU86 A0A1B6LLZ7 A0A182QC88 Q16MQ9 A0A224XI48 A0A1W4XNR6 A0A182GQP1 A0A182Y4U1 B3MTC9 A0A182FM81 A0A182RM22 B4JYJ5 B4K5Y1 A0A084W9N5 A0A182V1G0 A0A0Q9XBS5 A0A182SPU1 J9KGQ7 A0A182WY28 A0A2K8JME3 Q7Q4T0 A0A1W4U620 A0A336M458 A0A3B0JKK2 W5JLT5 J9JLW8 A0A0Q9WV74 B4NFN7 A0A182VV47 B4QZJ5 Q29BX5 A0A1Q3FSF6 Q9NH67 A0A023F109 B4HZ87 Q0KI00 B4MBV1 B4GPI5 A0A182K329 B3P5I4 A0A0M4F678 B4PQ42 A0A336KKE2 Q6IDH1 B3MTC8 A0A0N7Z8X1 A0A182PL06 A0A069DXR4 A0A0R1E4S0 A0A182N0H6 A0A1W4UH69 A0A224X6V4 A0A1B6GC72 A0A1W4UH28 B4PQ46 B3P5I7 A0A0V0G587 B3P5I5 A0A182TL54 A0A0T6AUS1 A0A0L0BPV0 B4QZJ4 A0A1I8N5N4 A0A0R1E4R4 B4PQ45 A0A0R1E9Q7 B4PQ43 B4QZJ3 A0A1I8PCE0 E2BB79 A0A182HMT5 B0X825 K7INE5
PDB
6BV4
E-value=4.81329e-136,
Score=1245
Ontologies
PATHWAY
GO
PANTHER
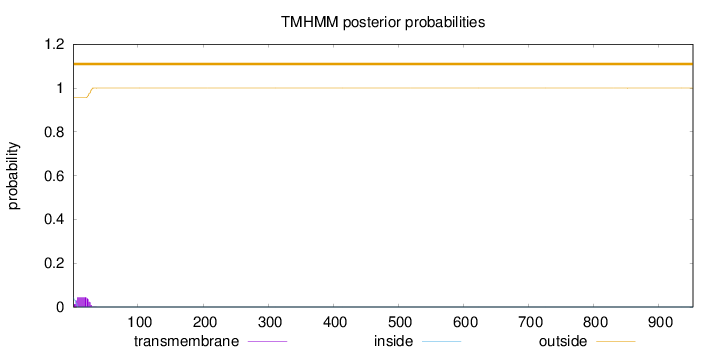
Topology
SignalP
Position: 1 - 18,
Likelihood: 0.977212

Length:
953
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.961269999999999
Exp number, first 60 AAs:
0.95679
Total prob of N-in:
0.04433
outside
1 - 953
Population Genetic Test Statistics
Pi
220.398907
Theta
160.095999
Tajima's D
-0.844946
CLR
0.26506
CSRT
0.165741712914354
Interpretation
Uncertain
