Pre Gene Modal
BGIBMGA004980
Annotation
Serine/threonine-protein_phosphatase_2A_56_kDa_regulatory_subunit_alpha_isoform_[Papilio_xuthus]
Full name
Serine/threonine protein phosphatase 2A regulatory subunit
Location in the cell
Cytoplasmic Reliability : 4.084
Sequence
CDS
ATGTCTTCGGGAACGTTTGTGGACAGGATTGATCCATTTGCTAAAAGATCACTGAAGAAAAAGACTAAGAAGTCTCAGGGTTCCTCTCGTTATCGTAATACGCAAGATGTGGAGCTACAAGCATTGCCATTGTTAAAAGATGTCCCAAGCAGTGAACAAGAGGAATTGTTCATCCGTAAACTCCGACAATGTTGTGTGGCGTTTGATTTCATGGATCCTGTTGCTGATTTGAAAGGCAAAGAGATCAAACGTGCTACACTGAATGAACTTGTTGAGTACATTCATGGTGGCCGCGGTGTACTTACAGAGCCTGTTTATCCAGAAATTATAAAAATGATATCGGCAAATCTGTTCCGTACCCTGCCTCCAAGTGAAAACCCTGATTTTGATCCTGAAGAGGATGATCCGACTCTGGAGGCGTCATGGCCCCATCTTCAACTAGTCTACGAATTCTTCCTGCGCTTCCTTGAGTCAACTGATTTCCAACCTACTATAGGCAAGAAGGTTATCGATCAAAAATTTGTTTTACAGCTTTTAGATTTGTTCGACTCTGAAGATCCGCGCGAGCGCGATTTTCTGAAAACCGTGCTACATCGCATCTATGGAAAATTTTTGGGTCTGCGAGCATTTATACGGAAACAGATCAATAATATATTTTTGAGATTTGTCTATGAAACTGAACACTTCAATGGTGTTGGTGAACTGCTAGAAATATTGGGAAGTATAATAAACGGATTCGCCTTACCGTTGAAGAGTGAGCACAAACAATTTTTGGTGAAGGTGCTACTGCCTCTGCACAAGGTGAAGTGCTTGTCATTATACCACGCTCAGCTTGCCTACTGCGTGGTTCAGTTCCTCGAGAAGGACGCTACGCTCACGGAGCCCGTTGTGCGTGGACTACTCAAGTTTTGGCCGAAGACTTGTTCACAGAAAGAGGTAATGTTTTTGGGCGAGATTGAAGAAATATTGGATGTAATTGAGCCTGCACAATTTGCTAAAATCCAAGAACCACTGTTCCGGCAAATTGCAAAATGTGTTTCCAGTCCTCATTTTCAGGTAGCTGAGAGGGCTTTGTATTTCTGGAATAACGAGTATATAATGTCGTTGATGGAAGAGAACAACCATGTGATCATGCCTATAATGTTCCCAGCGTTGTACCGAATCAGCAAGGAGCACTGGAACCAAACAATCGTGGCCCTCGTCTACAATGTGCTCAAAACATTCATGGAAATGAATTCCAAACTATTTGACGAACTCACAGCTAGCTACAAAGCAGAAAGGCAGAAAGAAAAGAAGCGCGAGAAGGAGCGCGACGAGCTGTGGAAGCGGCTGGGCGAGCTGGAGATCAGCCACAAGCGACAGCAACACGCGGCGCCGCTCGCCAAGTGA
Protein
MSSGTFVDRIDPFAKRSLKKKTKKSQGSSRYRNTQDVELQALPLLKDVPSSEQEELFIRKLRQCCVAFDFMDPVADLKGKEIKRATLNELVEYIHGGRGVLTEPVYPEIIKMISANLFRTLPPSENPDFDPEEDDPTLEASWPHLQLVYEFFLRFLESTDFQPTIGKKVIDQKFVLQLLDLFDSEDPRERDFLKTVLHRIYGKFLGLRAFIRKQINNIFLRFVYETEHFNGVGELLEILGSIINGFALPLKSEHKQFLVKVLLPLHKVKCLSLYHAQLAYCVVQFLEKDATLTEPVVRGLLKFWPKTCSQKEVMFLGEIEEILDVIEPAQFAKIQEPLFRQIAKCVSSPHFQVAERALYFWNNEYIMSLMEENNHVIMPIMFPALYRISKEHWNQTIVALVYNVLKTFMEMNSKLFDELTASYKAERQKEKKREKERDELWKRLGELEISHKRQQHAAPLAK
Summary
Similarity
Belongs to the phosphatase 2A regulatory subunit.
Uniprot
H9J639
A0A194PME8
A0A2A4JL64
S4PEU8
A0A194RQA0
A0A1E1WG11
+ More
A0A139WFD8 A0A1Y1NBD4 A0A1B6GP26 A0A1B6MMP8 E2AI01 V9IL55 A0A026WWL7 E0VNG0 A0A088AB82 A0A195C042 A0A195BH84 A0A151JSW2 A0A158NL58 A0A151XBH2 F4WFV9 A0A195F0S0 A0A1B6DLE0 K7J318 A0A023F4W1 A0A069DU76 A0A224XCA5 A0A0V0G8P0 A0A067RTM9 A0A345AH86 A0A0P4WJC7 A0A1L8DM34 A0A1B0CNW0 A0A182IJW4 A0A182UYY7 A0A182N5K5 A0A182QR69 W5JHD9 A0A182K9V0 A0A182PNM5 A0A182W0G4 A0A182XFJ6 A0A084VCI4 A0A182L7K5 Q7QBB5 A0A182I800 A0A182Y755 A0A2M3Z090 A0A2M4A5C0 A0A2M4A4M5 A0A2M4A4X3 A0A2M4BIY7 A0A2R5LMC4 A0A182FAT2 A0A154P5E2 A0A023FKH3 A0A2M3Z1R9 A0A2M3Z1P3 A0A1E1XUH3 A0A336KXD5 U5EJN6 A0A1Q3EU41 A0A0K8WGZ5 A0A034VG76 T1DE39 A0A0A1WEH5 A0A023GKV0 A0A1E1X271 B4NAQ2 A0A2P2HVP2 A0A0N8ERZ7 A0A182MSC9 A0A1I8PST0 A0A1W4UCB9 A0A0K8WH49 W8BI76 A0A1I8MC24 B3LXJ3 T1PDN9 A0A131XC73 B3P601 Q9VB23 B4PRC8 B4KBA5 A0A224Z8E5 V5HH63 A0A131YLQ1 L7M3M4 A0A0K8TNV7 A0A131Y560 B4JRY7 A0A1B0A3R7 A0A1B0FKS4 A0A1B0BSW6 A0A0K8TPJ1 B4M634 A0A1A9VGB6 A0A1A9WT78 A0A0L0CEE2 A0A3B0JIS5
A0A139WFD8 A0A1Y1NBD4 A0A1B6GP26 A0A1B6MMP8 E2AI01 V9IL55 A0A026WWL7 E0VNG0 A0A088AB82 A0A195C042 A0A195BH84 A0A151JSW2 A0A158NL58 A0A151XBH2 F4WFV9 A0A195F0S0 A0A1B6DLE0 K7J318 A0A023F4W1 A0A069DU76 A0A224XCA5 A0A0V0G8P0 A0A067RTM9 A0A345AH86 A0A0P4WJC7 A0A1L8DM34 A0A1B0CNW0 A0A182IJW4 A0A182UYY7 A0A182N5K5 A0A182QR69 W5JHD9 A0A182K9V0 A0A182PNM5 A0A182W0G4 A0A182XFJ6 A0A084VCI4 A0A182L7K5 Q7QBB5 A0A182I800 A0A182Y755 A0A2M3Z090 A0A2M4A5C0 A0A2M4A4M5 A0A2M4A4X3 A0A2M4BIY7 A0A2R5LMC4 A0A182FAT2 A0A154P5E2 A0A023FKH3 A0A2M3Z1R9 A0A2M3Z1P3 A0A1E1XUH3 A0A336KXD5 U5EJN6 A0A1Q3EU41 A0A0K8WGZ5 A0A034VG76 T1DE39 A0A0A1WEH5 A0A023GKV0 A0A1E1X271 B4NAQ2 A0A2P2HVP2 A0A0N8ERZ7 A0A182MSC9 A0A1I8PST0 A0A1W4UCB9 A0A0K8WH49 W8BI76 A0A1I8MC24 B3LXJ3 T1PDN9 A0A131XC73 B3P601 Q9VB23 B4PRC8 B4KBA5 A0A224Z8E5 V5HH63 A0A131YLQ1 L7M3M4 A0A0K8TNV7 A0A131Y560 B4JRY7 A0A1B0A3R7 A0A1B0FKS4 A0A1B0BSW6 A0A0K8TPJ1 B4M634 A0A1A9VGB6 A0A1A9WT78 A0A0L0CEE2 A0A3B0JIS5
Pubmed
19121390
26354079
23622113
18362917
19820115
28004739
+ More
20798317 24508170 30249741 20566863 21347285 21719571 20075255 25474469 26334808 24845553 20920257 23761445 24438588 20966253 12364791 14747013 17210077 25244985 29209593 25348373 24330624 25830018 28503490 17994087 26999592 24495485 25315136 18057021 28049606 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 17550304 28797301 25765539 26830274 25576852 26369729 29652888 26108605
20798317 24508170 30249741 20566863 21347285 21719571 20075255 25474469 26334808 24845553 20920257 23761445 24438588 20966253 12364791 14747013 17210077 25244985 29209593 25348373 24330624 25830018 28503490 17994087 26999592 24495485 25315136 18057021 28049606 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 26109357 26109356 17550304 28797301 25765539 26830274 25576852 26369729 29652888 26108605
EMBL
BABH01029939
BABH01029940
KQ459604
KPI92290.1
NWSH01001069
PCG72797.1
+ More
GAIX01004332 JAA88228.1 KQ460045 KPJ18201.1 GDQN01005131 JAT85923.1 KQ971352 KYB26642.1 GEZM01007309 JAV95252.1 GECZ01005594 JAS64175.1 GEBQ01002763 JAT37214.1 GL439621 EFN66909.1 JR050232 AEY61206.1 KK107078 QOIP01000011 EZA60452.1 RLU16633.1 DS235339 EEB14916.1 KQ978501 KYM93523.1 KQ976488 KYM83514.1 KQ978530 KYN30310.1 ADTU01019257 ADTU01019258 ADTU01019259 ADTU01019260 KQ982320 KYQ57723.1 GL888128 EGI66726.1 KQ981866 KYN34073.1 GEDC01029357 GEDC01023225 GEDC01020674 GEDC01011875 GEDC01010836 GEDC01007979 GEDC01005386 JAS07941.1 JAS14073.1 JAS16624.1 JAS25423.1 JAS26462.1 JAS29319.1 JAS31912.1 GBBI01002196 JAC16516.1 GBGD01001409 JAC87480.1 GFTR01006381 JAW10045.1 GECL01001739 JAP04385.1 KK852425 KDR24165.1 MF175170 AXF35720.1 GDRN01085051 GDRN01085050 JAI61407.1 GFDF01006551 JAV07533.1 AJWK01021007 AXCN02001161 AXCN02001162 ADMH02001517 ETN62224.1 ATLV01010762 KE524612 KFB35678.1 AAAB01008880 EAA08635.3 APCN01002592 GGFM01001189 MBW21940.1 GGFK01002662 MBW35983.1 GGFK01002445 MBW35766.1 GGFK01002451 MBW35772.1 GGFJ01003874 MBW53015.1 GGLE01006560 MBY10686.1 KQ434809 KZC06340.1 GBBK01002350 JAC22132.1 GGFM01001637 MBW22388.1 GGFM01001674 MBW22425.1 GFAA01000459 JAU02976.1 UFQS01001020 UFQT01001020 SSX08605.1 SSX28521.1 GANO01002171 JAB57700.1 GFDL01016222 JAV18823.1 GDHF01033581 GDHF01001888 JAI18733.1 JAI50426.1 GAKP01017444 GAKP01017443 GAKP01017442 GAKP01017441 JAC41509.1 GALA01001207 JAA93645.1 GBXI01016883 GBXI01014217 GBXI01004701 JAC97408.1 JAD00075.1 JAD09591.1 GBBM01000906 JAC34512.1 GFAC01005870 JAT93318.1 CH964232 EDW80866.1 IACF01000060 LAB65873.1 GDUN01000874 JAN95045.1 AXCM01002872 GDHF01024358 GDHF01010458 GDHF01001833 JAI27956.1 JAI41856.1 JAI50481.1 GAMC01009883 GAMC01009882 GAMC01009881 GAMC01009880 JAB96673.1 CH902617 EDV43887.2 KPU80522.1 KA646809 KA649640 AFP61438.1 GEFH01003327 JAP65254.1 CH954182 EDV53401.1 AF145696 AE014297 AAD38671.1 AAF56720.2 CM000160 EDW98492.1 CH933806 EDW16833.2 KRG02327.1 GFPF01011708 MAA22854.1 GANP01011620 JAB72848.1 GEDV01009082 JAP79475.1 GACK01007291 JAA57743.1 GDAI01001792 JAI15811.1 GEFM01002735 GEGO01000918 JAP73061.1 JAR94486.1 CH916373 EDV94527.1 CCAG010000730 JXJN01019901 GDAI01001793 JAI15810.1 CH940652 EDW59110.1 KRF78683.1 JRES01000501 KNC30616.1 OUUW01000005 SPP80643.1
GAIX01004332 JAA88228.1 KQ460045 KPJ18201.1 GDQN01005131 JAT85923.1 KQ971352 KYB26642.1 GEZM01007309 JAV95252.1 GECZ01005594 JAS64175.1 GEBQ01002763 JAT37214.1 GL439621 EFN66909.1 JR050232 AEY61206.1 KK107078 QOIP01000011 EZA60452.1 RLU16633.1 DS235339 EEB14916.1 KQ978501 KYM93523.1 KQ976488 KYM83514.1 KQ978530 KYN30310.1 ADTU01019257 ADTU01019258 ADTU01019259 ADTU01019260 KQ982320 KYQ57723.1 GL888128 EGI66726.1 KQ981866 KYN34073.1 GEDC01029357 GEDC01023225 GEDC01020674 GEDC01011875 GEDC01010836 GEDC01007979 GEDC01005386 JAS07941.1 JAS14073.1 JAS16624.1 JAS25423.1 JAS26462.1 JAS29319.1 JAS31912.1 GBBI01002196 JAC16516.1 GBGD01001409 JAC87480.1 GFTR01006381 JAW10045.1 GECL01001739 JAP04385.1 KK852425 KDR24165.1 MF175170 AXF35720.1 GDRN01085051 GDRN01085050 JAI61407.1 GFDF01006551 JAV07533.1 AJWK01021007 AXCN02001161 AXCN02001162 ADMH02001517 ETN62224.1 ATLV01010762 KE524612 KFB35678.1 AAAB01008880 EAA08635.3 APCN01002592 GGFM01001189 MBW21940.1 GGFK01002662 MBW35983.1 GGFK01002445 MBW35766.1 GGFK01002451 MBW35772.1 GGFJ01003874 MBW53015.1 GGLE01006560 MBY10686.1 KQ434809 KZC06340.1 GBBK01002350 JAC22132.1 GGFM01001637 MBW22388.1 GGFM01001674 MBW22425.1 GFAA01000459 JAU02976.1 UFQS01001020 UFQT01001020 SSX08605.1 SSX28521.1 GANO01002171 JAB57700.1 GFDL01016222 JAV18823.1 GDHF01033581 GDHF01001888 JAI18733.1 JAI50426.1 GAKP01017444 GAKP01017443 GAKP01017442 GAKP01017441 JAC41509.1 GALA01001207 JAA93645.1 GBXI01016883 GBXI01014217 GBXI01004701 JAC97408.1 JAD00075.1 JAD09591.1 GBBM01000906 JAC34512.1 GFAC01005870 JAT93318.1 CH964232 EDW80866.1 IACF01000060 LAB65873.1 GDUN01000874 JAN95045.1 AXCM01002872 GDHF01024358 GDHF01010458 GDHF01001833 JAI27956.1 JAI41856.1 JAI50481.1 GAMC01009883 GAMC01009882 GAMC01009881 GAMC01009880 JAB96673.1 CH902617 EDV43887.2 KPU80522.1 KA646809 KA649640 AFP61438.1 GEFH01003327 JAP65254.1 CH954182 EDV53401.1 AF145696 AE014297 AAD38671.1 AAF56720.2 CM000160 EDW98492.1 CH933806 EDW16833.2 KRG02327.1 GFPF01011708 MAA22854.1 GANP01011620 JAB72848.1 GEDV01009082 JAP79475.1 GACK01007291 JAA57743.1 GDAI01001792 JAI15811.1 GEFM01002735 GEGO01000918 JAP73061.1 JAR94486.1 CH916373 EDV94527.1 CCAG010000730 JXJN01019901 GDAI01001793 JAI15810.1 CH940652 EDW59110.1 KRF78683.1 JRES01000501 KNC30616.1 OUUW01000005 SPP80643.1
Proteomes
UP000005204
UP000053268
UP000218220
UP000053240
UP000007266
UP000000311
+ More
UP000053097 UP000279307 UP000009046 UP000005203 UP000078542 UP000078540 UP000078492 UP000005205 UP000075809 UP000007755 UP000078541 UP000002358 UP000027135 UP000092461 UP000075880 UP000075903 UP000075884 UP000075886 UP000000673 UP000075881 UP000075885 UP000075920 UP000076407 UP000030765 UP000075882 UP000007062 UP000075840 UP000076408 UP000069272 UP000076502 UP000007798 UP000075883 UP000095300 UP000192221 UP000095301 UP000007801 UP000008711 UP000000803 UP000002282 UP000009192 UP000001070 UP000092445 UP000092444 UP000092460 UP000008792 UP000078200 UP000091820 UP000037069 UP000268350
UP000053097 UP000279307 UP000009046 UP000005203 UP000078542 UP000078540 UP000078492 UP000005205 UP000075809 UP000007755 UP000078541 UP000002358 UP000027135 UP000092461 UP000075880 UP000075903 UP000075884 UP000075886 UP000000673 UP000075881 UP000075885 UP000075920 UP000076407 UP000030765 UP000075882 UP000007062 UP000075840 UP000076408 UP000069272 UP000076502 UP000007798 UP000075883 UP000095300 UP000192221 UP000095301 UP000007801 UP000008711 UP000000803 UP000002282 UP000009192 UP000001070 UP000092445 UP000092444 UP000092460 UP000008792 UP000078200 UP000091820 UP000037069 UP000268350
Interpro
SUPFAM
SSF48371
SSF48371
Gene 3D
ProteinModelPortal
H9J639
A0A194PME8
A0A2A4JL64
S4PEU8
A0A194RQA0
A0A1E1WG11
+ More
A0A139WFD8 A0A1Y1NBD4 A0A1B6GP26 A0A1B6MMP8 E2AI01 V9IL55 A0A026WWL7 E0VNG0 A0A088AB82 A0A195C042 A0A195BH84 A0A151JSW2 A0A158NL58 A0A151XBH2 F4WFV9 A0A195F0S0 A0A1B6DLE0 K7J318 A0A023F4W1 A0A069DU76 A0A224XCA5 A0A0V0G8P0 A0A067RTM9 A0A345AH86 A0A0P4WJC7 A0A1L8DM34 A0A1B0CNW0 A0A182IJW4 A0A182UYY7 A0A182N5K5 A0A182QR69 W5JHD9 A0A182K9V0 A0A182PNM5 A0A182W0G4 A0A182XFJ6 A0A084VCI4 A0A182L7K5 Q7QBB5 A0A182I800 A0A182Y755 A0A2M3Z090 A0A2M4A5C0 A0A2M4A4M5 A0A2M4A4X3 A0A2M4BIY7 A0A2R5LMC4 A0A182FAT2 A0A154P5E2 A0A023FKH3 A0A2M3Z1R9 A0A2M3Z1P3 A0A1E1XUH3 A0A336KXD5 U5EJN6 A0A1Q3EU41 A0A0K8WGZ5 A0A034VG76 T1DE39 A0A0A1WEH5 A0A023GKV0 A0A1E1X271 B4NAQ2 A0A2P2HVP2 A0A0N8ERZ7 A0A182MSC9 A0A1I8PST0 A0A1W4UCB9 A0A0K8WH49 W8BI76 A0A1I8MC24 B3LXJ3 T1PDN9 A0A131XC73 B3P601 Q9VB23 B4PRC8 B4KBA5 A0A224Z8E5 V5HH63 A0A131YLQ1 L7M3M4 A0A0K8TNV7 A0A131Y560 B4JRY7 A0A1B0A3R7 A0A1B0FKS4 A0A1B0BSW6 A0A0K8TPJ1 B4M634 A0A1A9VGB6 A0A1A9WT78 A0A0L0CEE2 A0A3B0JIS5
A0A139WFD8 A0A1Y1NBD4 A0A1B6GP26 A0A1B6MMP8 E2AI01 V9IL55 A0A026WWL7 E0VNG0 A0A088AB82 A0A195C042 A0A195BH84 A0A151JSW2 A0A158NL58 A0A151XBH2 F4WFV9 A0A195F0S0 A0A1B6DLE0 K7J318 A0A023F4W1 A0A069DU76 A0A224XCA5 A0A0V0G8P0 A0A067RTM9 A0A345AH86 A0A0P4WJC7 A0A1L8DM34 A0A1B0CNW0 A0A182IJW4 A0A182UYY7 A0A182N5K5 A0A182QR69 W5JHD9 A0A182K9V0 A0A182PNM5 A0A182W0G4 A0A182XFJ6 A0A084VCI4 A0A182L7K5 Q7QBB5 A0A182I800 A0A182Y755 A0A2M3Z090 A0A2M4A5C0 A0A2M4A4M5 A0A2M4A4X3 A0A2M4BIY7 A0A2R5LMC4 A0A182FAT2 A0A154P5E2 A0A023FKH3 A0A2M3Z1R9 A0A2M3Z1P3 A0A1E1XUH3 A0A336KXD5 U5EJN6 A0A1Q3EU41 A0A0K8WGZ5 A0A034VG76 T1DE39 A0A0A1WEH5 A0A023GKV0 A0A1E1X271 B4NAQ2 A0A2P2HVP2 A0A0N8ERZ7 A0A182MSC9 A0A1I8PST0 A0A1W4UCB9 A0A0K8WH49 W8BI76 A0A1I8MC24 B3LXJ3 T1PDN9 A0A131XC73 B3P601 Q9VB23 B4PRC8 B4KBA5 A0A224Z8E5 V5HH63 A0A131YLQ1 L7M3M4 A0A0K8TNV7 A0A131Y560 B4JRY7 A0A1B0A3R7 A0A1B0FKS4 A0A1B0BSW6 A0A0K8TPJ1 B4M634 A0A1A9VGB6 A0A1A9WT78 A0A0L0CEE2 A0A3B0JIS5
PDB
2NPP
E-value=3.10818e-150,
Score=1364
Ontologies
GO
GO:0007165
GO:0000159
GO:0019888
GO:0072542
GO:0005634
GO:0006470
GO:0031952
GO:0005829
GO:0016021
GO:0007059
GO:0045879
GO:0051898
GO:0045880
GO:0006914
GO:0010888
GO:0001933
GO:0051225
GO:0046627
GO:0008340
GO:0005654
GO:0007623
GO:0005488
GO:0006535
GO:0006391
GO:0031167
GO:0003677
GO:0008408
GO:0005112
GO:0015930
PANTHER
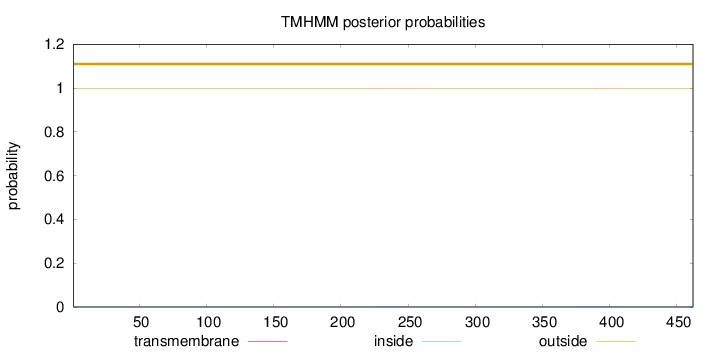
Topology
Length:
462
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.02509
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00345
outside
1 - 462
Population Genetic Test Statistics
Pi
261.20855
Theta
205.022916
Tajima's D
0.932414
CLR
0.008959
CSRT
0.640817959102045
Interpretation
Uncertain
