Pre Gene Modal
BGIBMGA004993
Annotation
hypothetical_protein_KGM_16296_[Danaus_plexippus]
Full name
Probable pterin-4-alpha-carbinolamine dehydratase
+ More
Pterin-4-alpha-carbinolamine dehydratase
Pterin-4-alpha-carbinolamine dehydratase
Alternative Name
4-alpha-hydroxy-tetrahydropterin dehydratase
Pterin carbinolamine dehydratase
Pterin carbinolamine dehydratase
Location in the cell
Cytoplasmic Reliability : 1.352 Nuclear Reliability : 1.828
Sequence
CDS
ATGGCTGATAAACTTAATCAAGAGGAGCGCACAACTGAGCTCAAACCTCTACTGGAATCTGGTTGGAAAATTCAGAGCAACAGAGACGCAATAGAAAAGGAATTTCAGTTCAAAAATTTTAATGAAGCTTTTGGATTCATGACCAGGGTCGCGCTGCTAGCTGAGAAAATGGATCATCATCCTGAGTGGTTCAATGTATACAACAAGTTGCAAGTGACACTTTCTTCACATGACGTAAATGGATTGAGCAAAAGGGATATCAAAATGGCCAGTTTTATGGATAAAAATTATAATTAA
Protein
MADKLNQEERTTELKPLLESGWKIQSNRDAIEKEFQFKNFNEAFGFMTRVALLAEKMDHHPEWFNVYNKLQVTLSSHDVNGLSKRDIKMASFMDKNYN
Summary
Description
Involved in tetrahydrobiopterin biosynthesis.
Catalytic Activity
(4aS,6R)-4a-hydroxy-L-erythro-5,6,7,8-tetrahydrobiopterin = (6R)-L-erythro-6,7-dihydrobiopterin + H2O
Similarity
Belongs to the tRNA nucleotidyltransferase/poly(A) polymerase family.
Belongs to the pterin-4-alpha-carbinolamine dehydratase family.
Belongs to the pterin-4-alpha-carbinolamine dehydratase family.
Keywords
Lyase
Tetrahydrobiopterin biosynthesis
Feature
chain Probable pterin-4-alpha-carbinolamine dehydratase
Uniprot
A0A2H1W3E9
A0A212FKX8
A0A1E1WEZ9
A0A3S2LMS3
A0A194RLQ8
I4DJ38
+ More
A0A194PG24 S4PDT3 A0A0L7KSR2 D6WSN1 E9J8L2 A0A151IET1 A0A0J7NF19 Q0PXW3 A0A0M9A2A9 E2BIN1 A0A195DMN3 A0A0L7R9Q1 F4W696 A0A1W4XLJ6 A0A2P8XPX6 A0A158NWP0 A0A2J7QUJ4 A0A067RQQ6 A0A1Y1MD96 K7J1K5 A0A154P0E2 A0A1J1IQF5 A0A1W0WCQ8 P0C8L6 A0A087TT03 A0A088A3V1 A0A2H4MCS2 E0W0B4 A0A1B6DMD8 E2ACR2 A0A026WCW7 A0A2A3EKB4 A0A2L2YCR0 A0A367KFP1 A0A0C7B9U7 A0A1D1VAU2 A0A0B2VHE3 A0A151WHQ1 A0A2G4SFG0 A0A1X0R650 A0A0K0G1W5 T1D5J4 A0A182MWS8 H9J652 A0A182RMW7 T1EZ60 A0A0C7CHN1 A0A2C9JM84 N6SYA3 A0A2G4T1X9 A0A1X0RB64 A0A183H5D5 A0A1S4F318 T1J3D0 A0A1D2NAS6 Q17GS3 U4UNN7 A0A1S3H7L4 A0A023EJL2 A0A195AW62 A0A023EJN6 V5IA76 A0A1X0S1K0 A0A367JQZ5 A0A0M3HUM5 F1L9R7 A0A182G557 A0A023EKG5 H3F4J3 A0A1B6J5D6 A0A3M6V4E1 A0A1V9XT71 A0A0N4ZUP2 A0A0P5ZI94 A0A0K8RRF0 A0A0P5YMW9 A0A131Y9F5 S4RZI4 A0A2B4SAN6 A0A367JTB6 A0A0N8EJX8 A0A0R3RW82 A0A023G9I6 A0A023FRB9 A0A023G6T6 G3MMY8 A0A210PVD7 A0A0P6BIV2 A0A1B6LTL2 B7P7V8 E9GGR8 P58249 A0A2R7X2N6 A0A023FG19
A0A194PG24 S4PDT3 A0A0L7KSR2 D6WSN1 E9J8L2 A0A151IET1 A0A0J7NF19 Q0PXW3 A0A0M9A2A9 E2BIN1 A0A195DMN3 A0A0L7R9Q1 F4W696 A0A1W4XLJ6 A0A2P8XPX6 A0A158NWP0 A0A2J7QUJ4 A0A067RQQ6 A0A1Y1MD96 K7J1K5 A0A154P0E2 A0A1J1IQF5 A0A1W0WCQ8 P0C8L6 A0A087TT03 A0A088A3V1 A0A2H4MCS2 E0W0B4 A0A1B6DMD8 E2ACR2 A0A026WCW7 A0A2A3EKB4 A0A2L2YCR0 A0A367KFP1 A0A0C7B9U7 A0A1D1VAU2 A0A0B2VHE3 A0A151WHQ1 A0A2G4SFG0 A0A1X0R650 A0A0K0G1W5 T1D5J4 A0A182MWS8 H9J652 A0A182RMW7 T1EZ60 A0A0C7CHN1 A0A2C9JM84 N6SYA3 A0A2G4T1X9 A0A1X0RB64 A0A183H5D5 A0A1S4F318 T1J3D0 A0A1D2NAS6 Q17GS3 U4UNN7 A0A1S3H7L4 A0A023EJL2 A0A195AW62 A0A023EJN6 V5IA76 A0A1X0S1K0 A0A367JQZ5 A0A0M3HUM5 F1L9R7 A0A182G557 A0A023EKG5 H3F4J3 A0A1B6J5D6 A0A3M6V4E1 A0A1V9XT71 A0A0N4ZUP2 A0A0P5ZI94 A0A0K8RRF0 A0A0P5YMW9 A0A131Y9F5 S4RZI4 A0A2B4SAN6 A0A367JTB6 A0A0N8EJX8 A0A0R3RW82 A0A023G9I6 A0A023FRB9 A0A023G6T6 G3MMY8 A0A210PVD7 A0A0P6BIV2 A0A1B6LTL2 B7P7V8 E9GGR8 P58249 A0A2R7X2N6 A0A023FG19
EC Number
4.2.1.96
Pubmed
22118469
26354079
22651552
23622113
26227816
18362917
+ More
19820115 21282665 20798317 21719571 29403074 21347285 24845553 28004739 20075255 20566863 24508170 30249741 26561354 29674435 27956601 27649274 24330624 19121390 23254933 15562597 23537049 27289101 17510324 24945155 21685128 26483478 18806794 30382153 28327890 22216098 28812685 21292972 11710527
19820115 21282665 20798317 21719571 29403074 21347285 24845553 28004739 20075255 20566863 24508170 30249741 26561354 29674435 27956601 27649274 24330624 19121390 23254933 15562597 23537049 27289101 17510324 24945155 21685128 26483478 18806794 30382153 28327890 22216098 28812685 21292972 11710527
EMBL
ODYU01005792
SOQ47024.1
AGBW02007949
OWR54394.1
GDQN01005490
JAT85564.1
+ More
RSAL01000053 RVE50060.1 KQ460045 KPJ18250.1 AK401306 BAM17928.1 KQ459604 KPI92247.1 GAIX01003521 JAA89039.1 JTDY01006327 KOB66101.1 KQ971354 EFA07130.1 GL769036 EFZ10827.1 KQ977858 KYM99323.1 LBMM01005909 KMQ91105.1 DQ673423 ABG81996.1 KQ435759 KOX75737.1 GL448516 EFN84478.1 KQ980724 KYN14091.1 KQ414621 KOC67600.1 GL887707 EGI70231.1 PYGN01001559 PSN34053.1 ADTU01028278 NEVH01010578 PNF32247.1 KK852478 KDR22975.1 GEZM01037936 JAV81836.1 KQ434787 KZC05311.1 CVRI01000057 CRL02447.1 MTYJ01000133 OQV12970.1 CK326948 KK116615 KFM68242.1 KX898002 ATI15031.1 DS235858 EEB19070.1 GEDC01010471 JAS26827.1 GL438568 EFN68806.1 KK107262 QOIP01000010 EZA53907.1 RLU17706.1 KZ288225 PBC31934.1 IAAA01010135 IAAA01010136 LAA05150.1 PJQL01000029 RCI00999.1 CCYT01000008 CCYT01000333 KV921302 CEG67846.1 CEG82113.1 ORE19922.1 BDGG01000005 GAU98756.1 JPKZ01001747 UYWY01020014 KHN80385.1 VDM40177.1 KQ983112 KYQ47366.1 KZ303875 PHZ07497.1 KV921903 ORE07487.1 GALA01000478 JAA94374.1 AXCM01004309 BABH01030005 BABH01030006 BABH01030007 BABH01030008 AMQM01002737 KB095858 ESO10741.1 CCYT01000424 CEG82969.1 APGK01052220 KB741211 ENN72774.1 KZ303845 PHZ15030.1 KV921879 ORE09118.1 KZ269977 UZAJ01001605 OZC12686.1 VDO33899.1 JH431823 LJIJ01000115 ODN02362.1 CH477256 EAT45864.1 KI207581 ERL95699.1 GAPW01004377 JAC09221.1 KQ976731 KYM76270.1 GAPW01004378 JAC09220.1 GALX01001670 JAB66796.1 KV921339 ORE18089.1 PJQL01000838 RCH92372.1 JI174827 ADY46871.1 JXUM01043026 KQ561347 KXJ78872.1 GAPW01003945 JAC09653.1 GECU01013313 JAS94393.1 RCHS01000120 RMX60775.1 MNPL01004580 OQR76643.1 GDIP01043177 LRGB01000725 JAM60538.1 KZS16236.1 GADI01000609 JAA73199.1 GDIP01056427 JAM47288.1 GEFM01000716 JAP75080.1 LSMT01000140 PFX25880.1 PJQL01000734 RCH93156.1 GDIQ01019444 JAN75293.1 GBBM01005880 JAC29538.1 GBBK01001312 JAC23170.1 GBBM01005879 JAC29539.1 JO843239 AEO34856.1 NEDP02005462 OWF40457.1 GDIP01015202 JAM88513.1 GEBQ01014037 GEBQ01012951 JAT25940.1 JAT27026.1 ABJB010347675 DS653761 EEC02680.1 GL732543 EFX81435.1 AF382144 KK856590 PTY26064.1 GBBK01004814 JAC19668.1
RSAL01000053 RVE50060.1 KQ460045 KPJ18250.1 AK401306 BAM17928.1 KQ459604 KPI92247.1 GAIX01003521 JAA89039.1 JTDY01006327 KOB66101.1 KQ971354 EFA07130.1 GL769036 EFZ10827.1 KQ977858 KYM99323.1 LBMM01005909 KMQ91105.1 DQ673423 ABG81996.1 KQ435759 KOX75737.1 GL448516 EFN84478.1 KQ980724 KYN14091.1 KQ414621 KOC67600.1 GL887707 EGI70231.1 PYGN01001559 PSN34053.1 ADTU01028278 NEVH01010578 PNF32247.1 KK852478 KDR22975.1 GEZM01037936 JAV81836.1 KQ434787 KZC05311.1 CVRI01000057 CRL02447.1 MTYJ01000133 OQV12970.1 CK326948 KK116615 KFM68242.1 KX898002 ATI15031.1 DS235858 EEB19070.1 GEDC01010471 JAS26827.1 GL438568 EFN68806.1 KK107262 QOIP01000010 EZA53907.1 RLU17706.1 KZ288225 PBC31934.1 IAAA01010135 IAAA01010136 LAA05150.1 PJQL01000029 RCI00999.1 CCYT01000008 CCYT01000333 KV921302 CEG67846.1 CEG82113.1 ORE19922.1 BDGG01000005 GAU98756.1 JPKZ01001747 UYWY01020014 KHN80385.1 VDM40177.1 KQ983112 KYQ47366.1 KZ303875 PHZ07497.1 KV921903 ORE07487.1 GALA01000478 JAA94374.1 AXCM01004309 BABH01030005 BABH01030006 BABH01030007 BABH01030008 AMQM01002737 KB095858 ESO10741.1 CCYT01000424 CEG82969.1 APGK01052220 KB741211 ENN72774.1 KZ303845 PHZ15030.1 KV921879 ORE09118.1 KZ269977 UZAJ01001605 OZC12686.1 VDO33899.1 JH431823 LJIJ01000115 ODN02362.1 CH477256 EAT45864.1 KI207581 ERL95699.1 GAPW01004377 JAC09221.1 KQ976731 KYM76270.1 GAPW01004378 JAC09220.1 GALX01001670 JAB66796.1 KV921339 ORE18089.1 PJQL01000838 RCH92372.1 JI174827 ADY46871.1 JXUM01043026 KQ561347 KXJ78872.1 GAPW01003945 JAC09653.1 GECU01013313 JAS94393.1 RCHS01000120 RMX60775.1 MNPL01004580 OQR76643.1 GDIP01043177 LRGB01000725 JAM60538.1 KZS16236.1 GADI01000609 JAA73199.1 GDIP01056427 JAM47288.1 GEFM01000716 JAP75080.1 LSMT01000140 PFX25880.1 PJQL01000734 RCH93156.1 GDIQ01019444 JAN75293.1 GBBM01005880 JAC29538.1 GBBK01001312 JAC23170.1 GBBM01005879 JAC29539.1 JO843239 AEO34856.1 NEDP02005462 OWF40457.1 GDIP01015202 JAM88513.1 GEBQ01014037 GEBQ01012951 JAT25940.1 JAT27026.1 ABJB010347675 DS653761 EEC02680.1 GL732543 EFX81435.1 AF382144 KK856590 PTY26064.1 GBBK01004814 JAC19668.1
Proteomes
UP000007151
UP000283053
UP000053240
UP000053268
UP000037510
UP000007266
+ More
UP000078542 UP000036403 UP000079169 UP000053105 UP000008237 UP000078492 UP000053825 UP000007755 UP000192223 UP000245037 UP000005205 UP000235965 UP000027135 UP000002358 UP000076502 UP000183832 UP000054359 UP000005203 UP000009046 UP000000311 UP000053097 UP000279307 UP000242457 UP000252139 UP000054919 UP000242381 UP000186922 UP000031036 UP000050794 UP000267007 UP000075809 UP000242254 UP000242414 UP000035680 UP000075883 UP000005204 UP000075900 UP000015101 UP000076420 UP000019118 UP000050787 UP000094527 UP000008820 UP000030742 UP000085678 UP000078540 UP000036681 UP000069940 UP000249989 UP000005239 UP000275408 UP000192247 UP000038045 UP000076858 UP000245300 UP000225706 UP000050640 UP000242188 UP000001555 UP000000305
UP000078542 UP000036403 UP000079169 UP000053105 UP000008237 UP000078492 UP000053825 UP000007755 UP000192223 UP000245037 UP000005205 UP000235965 UP000027135 UP000002358 UP000076502 UP000183832 UP000054359 UP000005203 UP000009046 UP000000311 UP000053097 UP000279307 UP000242457 UP000252139 UP000054919 UP000242381 UP000186922 UP000031036 UP000050794 UP000267007 UP000075809 UP000242254 UP000242414 UP000035680 UP000075883 UP000005204 UP000075900 UP000015101 UP000076420 UP000019118 UP000050787 UP000094527 UP000008820 UP000030742 UP000085678 UP000078540 UP000036681 UP000069940 UP000249989 UP000005239 UP000275408 UP000192247 UP000038045 UP000076858 UP000245300 UP000225706 UP000050640 UP000242188 UP000001555 UP000000305
Pfam
Interpro
IPR036428
PCD_sf
+ More
IPR001533 Pterin_deHydtase
IPR036891 Signal_recog_part_SRP54_M_sf
IPR004125 Signal_recog_particle_SRP54_M
IPR002646 PolA_pol_head_dom
IPR020588 RecA_ATP-bd
IPR013632 DNA_recomb/repair_Rad51_C
IPR032828 PolyA_RNA-bd
IPR027417 P-loop_NTPase
IPR009668 RNA_pol-assoc_fac_A49-like
IPR036225 SRP/SRP_N
IPR022941 SRP54
IPR003593 AAA+_ATPase
IPR000897 SRP54_GTPase_dom
IPR042101 SRP54_N_sf
IPR006325 SRP54_euk
IPR013822 Signal_recog_particl_SRP54_hlx
IPR001533 Pterin_deHydtase
IPR036891 Signal_recog_part_SRP54_M_sf
IPR004125 Signal_recog_particle_SRP54_M
IPR002646 PolA_pol_head_dom
IPR020588 RecA_ATP-bd
IPR013632 DNA_recomb/repair_Rad51_C
IPR032828 PolyA_RNA-bd
IPR027417 P-loop_NTPase
IPR009668 RNA_pol-assoc_fac_A49-like
IPR036225 SRP/SRP_N
IPR022941 SRP54
IPR003593 AAA+_ATPase
IPR000897 SRP54_GTPase_dom
IPR042101 SRP54_N_sf
IPR006325 SRP54_euk
IPR013822 Signal_recog_particl_SRP54_hlx
Gene 3D
ProteinModelPortal
A0A2H1W3E9
A0A212FKX8
A0A1E1WEZ9
A0A3S2LMS3
A0A194RLQ8
I4DJ38
+ More
A0A194PG24 S4PDT3 A0A0L7KSR2 D6WSN1 E9J8L2 A0A151IET1 A0A0J7NF19 Q0PXW3 A0A0M9A2A9 E2BIN1 A0A195DMN3 A0A0L7R9Q1 F4W696 A0A1W4XLJ6 A0A2P8XPX6 A0A158NWP0 A0A2J7QUJ4 A0A067RQQ6 A0A1Y1MD96 K7J1K5 A0A154P0E2 A0A1J1IQF5 A0A1W0WCQ8 P0C8L6 A0A087TT03 A0A088A3V1 A0A2H4MCS2 E0W0B4 A0A1B6DMD8 E2ACR2 A0A026WCW7 A0A2A3EKB4 A0A2L2YCR0 A0A367KFP1 A0A0C7B9U7 A0A1D1VAU2 A0A0B2VHE3 A0A151WHQ1 A0A2G4SFG0 A0A1X0R650 A0A0K0G1W5 T1D5J4 A0A182MWS8 H9J652 A0A182RMW7 T1EZ60 A0A0C7CHN1 A0A2C9JM84 N6SYA3 A0A2G4T1X9 A0A1X0RB64 A0A183H5D5 A0A1S4F318 T1J3D0 A0A1D2NAS6 Q17GS3 U4UNN7 A0A1S3H7L4 A0A023EJL2 A0A195AW62 A0A023EJN6 V5IA76 A0A1X0S1K0 A0A367JQZ5 A0A0M3HUM5 F1L9R7 A0A182G557 A0A023EKG5 H3F4J3 A0A1B6J5D6 A0A3M6V4E1 A0A1V9XT71 A0A0N4ZUP2 A0A0P5ZI94 A0A0K8RRF0 A0A0P5YMW9 A0A131Y9F5 S4RZI4 A0A2B4SAN6 A0A367JTB6 A0A0N8EJX8 A0A0R3RW82 A0A023G9I6 A0A023FRB9 A0A023G6T6 G3MMY8 A0A210PVD7 A0A0P6BIV2 A0A1B6LTL2 B7P7V8 E9GGR8 P58249 A0A2R7X2N6 A0A023FG19
A0A194PG24 S4PDT3 A0A0L7KSR2 D6WSN1 E9J8L2 A0A151IET1 A0A0J7NF19 Q0PXW3 A0A0M9A2A9 E2BIN1 A0A195DMN3 A0A0L7R9Q1 F4W696 A0A1W4XLJ6 A0A2P8XPX6 A0A158NWP0 A0A2J7QUJ4 A0A067RQQ6 A0A1Y1MD96 K7J1K5 A0A154P0E2 A0A1J1IQF5 A0A1W0WCQ8 P0C8L6 A0A087TT03 A0A088A3V1 A0A2H4MCS2 E0W0B4 A0A1B6DMD8 E2ACR2 A0A026WCW7 A0A2A3EKB4 A0A2L2YCR0 A0A367KFP1 A0A0C7B9U7 A0A1D1VAU2 A0A0B2VHE3 A0A151WHQ1 A0A2G4SFG0 A0A1X0R650 A0A0K0G1W5 T1D5J4 A0A182MWS8 H9J652 A0A182RMW7 T1EZ60 A0A0C7CHN1 A0A2C9JM84 N6SYA3 A0A2G4T1X9 A0A1X0RB64 A0A183H5D5 A0A1S4F318 T1J3D0 A0A1D2NAS6 Q17GS3 U4UNN7 A0A1S3H7L4 A0A023EJL2 A0A195AW62 A0A023EJN6 V5IA76 A0A1X0S1K0 A0A367JQZ5 A0A0M3HUM5 F1L9R7 A0A182G557 A0A023EKG5 H3F4J3 A0A1B6J5D6 A0A3M6V4E1 A0A1V9XT71 A0A0N4ZUP2 A0A0P5ZI94 A0A0K8RRF0 A0A0P5YMW9 A0A131Y9F5 S4RZI4 A0A2B4SAN6 A0A367JTB6 A0A0N8EJX8 A0A0R3RW82 A0A023G9I6 A0A023FRB9 A0A023G6T6 G3MMY8 A0A210PVD7 A0A0P6BIV2 A0A1B6LTL2 B7P7V8 E9GGR8 P58249 A0A2R7X2N6 A0A023FG19
PDB
4WIL
E-value=8.89756e-29,
Score=309
Ontologies
PATHWAY
GO
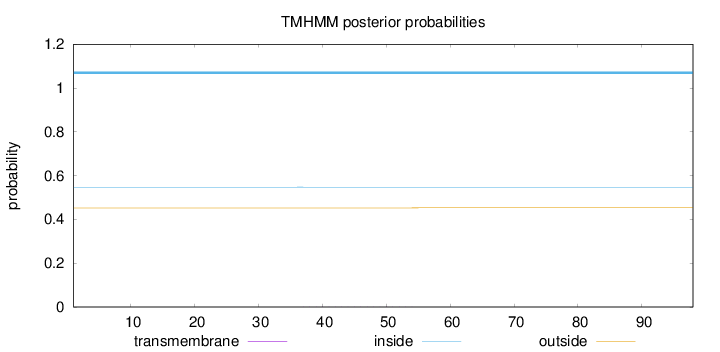
Topology
Length:
98
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00417999999999999
Exp number, first 60 AAs:
0.00399
Total prob of N-in:
0.54738
inside
1 - 98
Population Genetic Test Statistics
Pi
210.368292
Theta
182.416624
Tajima's D
0.398594
CLR
1.938909
CSRT
0.489575521223939
Interpretation
Uncertain
