Gene
KWMTBOMO14831
Annotation
PREDICTED:_uncharacterized_protein_LOC106138993_[Amyelois_transitella]
Location in the cell
Nuclear Reliability : 3.397
Sequence
CDS
ATGCCTCGGAAGGGTGCTAAAGAAGACGAAGAATGCTTTGCTGAACCTGCTCCACAAATTTCCTTCTGCGAATTAGAAAAATCTATGACTGCATTTTCTGGTGACGATGCATACGGCGTGGATACCTTTATAAAGGATTTTGAAGATATCGCTAATCTCATGCAATGGACGGGCATTGAAAAACTGATCTACGCAAAAAGGCTTTTGAAGGGGACAGCAAAATTGTTCCTGAGATCTTTGACTAGTACGACTACATGGGAACTCTTGAAAACGGGATTAAAAGAGGAATTTGGACTACATCTGAACAGCGCAGCCATTCACAAGACCATATCAAATAGGAAGATGAAACCAGGTGAAACCTACCAACAATATTTTTTGCAACTCAAAGAACTAGCAGTACTCGGAAATATTGAAGATGACGCCCTAATGGAATACATCATTGATGGCATCCCTGACAATGAAATGAATAAAACTATACTATATGGGGCTTCCAACATTAAAGAATTTAGGAAAAAACTTGATTTATATGCTGAAATAAAGAAGAAATGTCGAACGTCAAACAAAGCTTATAATCAATCAACAACATCCAAGCCTACAACATCTGGATGGAAGAAACCATTACCTAAGAAGAGATGTTTCAACTGCGGAGACCTAGATCATGAATCTTCAGGCTGCTCTAAAGGAATAAAGTGCTTCCGTTGTAATGACTTTGGTCATAAGTCAACTGACTGTCCCAAAAATATAAAGAAGACTCTAGAAATAAAATGTAACAAGGAGAATGATAGCCAAGCACCCTACAAAGAAGTTTTGATCAATAATGTTATGGTAAAATCACTTATTGACACTGGCAGTGATGTGAACCTAATGAATGAATCTACATTTACTCAAATACATGGCGACACTTGTAACTACCATCCTGATTTAGTACAAAGCCTGACAGGCATTGGTGACTGTGAGGTACATACAAGAGGCTCATGTACTCCAAGAATTGAGATTGATGGTTGCCAATGTGAAGCAACGTTCTATATAATCGAAGATGACAAAATACCAGTTGATGTAATAATAGGGAACCCAATATTACAGGACATAGAACTCAGTTTTAAGTCTGGTGTTATTTCTGCTAAAAGAATTTTACAACTCACAATGTTAGAGGATCAGGCTGAGGAACCAATGATTATAGGAAATTTCAAATATAAAGATCAGATTGAGAAAATACTGACAGAATATGACCCTGGAAAATGTTGTAAATAA
Protein
MPRKGAKEDEECFAEPAPQISFCELEKSMTAFSGDDAYGVDTFIKDFEDIANLMQWTGIEKLIYAKRLLKGTAKLFLRSLTSTTTWELLKTGLKEEFGLHLNSAAIHKTISNRKMKPGETYQQYFLQLKELAVLGNIEDDALMEYIIDGIPDNEMNKTILYGASNIKEFRKKLDLYAEIKKKCRTSNKAYNQSTTSKPTTSGWKKPLPKKRCFNCGDLDHESSGCSKGIKCFRCNDFGHKSTDCPKNIKKTLEIKCNKENDSQAPYKEVLINNVMVKSLIDTGSDVNLMNESTFTQIHGDTCNYHPDLVQSLTGIGDCEVHTRGSCTPRIEIDGCQCEATFYIIEDDKIPVDVIIGNPILQDIELSFKSGVISAKRILQLTMLEDQAEEPMIIGNFKYKDQIEKILTEYDPGKCCK
Summary
Uniprot
A0A0J7K9X8
A0A023F013
A0A0J7KQL1
A0A023EZG2
A0A023EYH9
J9JZ77
+ More
A0A023EYF0 W8BB66 A0A0K8URI1 A0A0A1XB16 A0A023EY26 X1X261 J9JNX9 A0A0A1X9Z0 A0A3L8DTV3 W8APB2 W8C8Q1 A0A0A1XFE9 A0A212FCW5 J9JQ65 A0A151IWN5 A0A0K8U2D9 J9HJJ8 A0A1B0CKJ5 A0A1L8DIN4 A0A139WKB5 A0A0K8V149 A0A1B0CR74 A0A2M3Z2X0 A0A151IKW6 W8B0V9 A0A034VAM6 A0A2M3Z3Z7 A0A1W7R6L8 A0A224XHR1 A0A0C9Q2I4 A0A0J7K0N3 A0A0E4G8D8 A0A069DWV3 A0A1S4HEU1 A0A2M4BF33 A0A2M4BF27 E9IMZ6 A0A2M4AEU2 W8C3C0 A0A151J262 A0A182W804 A0A2M4BFM2 A0A2M4BFF1 A0A2M4BDJ8 A0A151II89 A0A2M4BF88 A0A2A4JDT3 A0A2M4BDL5 Q24262 A0A232EEC1 A0A139WB30 A0A0A9YCQ1 A0A0J7K890 A0A2M4BN67
A0A023EYF0 W8BB66 A0A0K8URI1 A0A0A1XB16 A0A023EY26 X1X261 J9JNX9 A0A0A1X9Z0 A0A3L8DTV3 W8APB2 W8C8Q1 A0A0A1XFE9 A0A212FCW5 J9JQ65 A0A151IWN5 A0A0K8U2D9 J9HJJ8 A0A1B0CKJ5 A0A1L8DIN4 A0A139WKB5 A0A0K8V149 A0A1B0CR74 A0A2M3Z2X0 A0A151IKW6 W8B0V9 A0A034VAM6 A0A2M3Z3Z7 A0A1W7R6L8 A0A224XHR1 A0A0C9Q2I4 A0A0J7K0N3 A0A0E4G8D8 A0A069DWV3 A0A1S4HEU1 A0A2M4BF33 A0A2M4BF27 E9IMZ6 A0A2M4AEU2 W8C3C0 A0A151J262 A0A182W804 A0A2M4BFM2 A0A2M4BFF1 A0A2M4BDJ8 A0A151II89 A0A2M4BF88 A0A2A4JDT3 A0A2M4BDL5 Q24262 A0A232EEC1 A0A139WB30 A0A0A9YCQ1 A0A0J7K890 A0A2M4BN67
Pubmed
EMBL
LBMM01010976
KMQ87102.1
GBBI01004613
JAC14099.1
LBMM01004258
KMQ92652.1
+ More
GBBI01004551 JAC14161.1 GBBI01004550 JAC14162.1 ABLF02014660 ABLF02042774 GBBI01004549 JAC14163.1 GAMC01016174 JAB90381.1 GDHF01023027 JAI29287.1 GBXI01005773 JAD08519.1 GBBI01004477 JAC14235.1 ABLF02005127 ABLF02005129 ABLF02007401 ABLF02011059 GBXI01006173 JAD08119.1 QOIP01000004 RLU23736.1 GAMC01020207 JAB86348.1 GAMC01007326 JAB99229.1 GBXI01004632 JAD09660.1 AGBW02009151 OWR51557.1 ABLF02005846 KQ980852 KYN12192.1 GDHF01031400 JAI20914.1 CH478509 EJY58114.1 AJWK01016333 GFDF01007768 JAV06316.1 KQ971327 KYB28419.1 GDHF01019733 JAI32581.1 AJWK01024440 GGFM01002109 MBW22860.1 KQ977161 KYN05238.1 GAMC01015907 JAB90648.1 GAKP01019383 JAC39569.1 GGFM01002486 MBW23237.1 GEHC01000930 JAV46715.1 GFTR01008753 JAW07673.1 GBYB01008203 JAG77970.1 LBMM01018094 KMQ83877.1 HACL01000135 CFW94429.1 GBGD01000341 JAC88548.1 AAAB01008984 GGFJ01002513 MBW51654.1 GGFJ01002514 MBW51655.1 GL764261 EFZ18057.1 GGFK01005990 MBW39311.1 GAMC01002792 JAC03764.1 KQ980436 KYN16061.1 GGFJ01002620 MBW51761.1 GGFJ01002621 MBW51762.1 GGFJ01001974 MBW51115.1 KQ977505 KYN02276.1 GGFJ01002579 MBW51720.1 NWSH01001951 PCG69590.1 GGFJ01001952 MBW51093.1 Z27119 CAA81643.1 NNAY01005459 OXU16662.1 KQ971376 KYB25116.1 GBHO01014721 JAG28883.1 LBMM01011782 KMQ86633.1 GGFJ01005354 MBW54495.1
GBBI01004551 JAC14161.1 GBBI01004550 JAC14162.1 ABLF02014660 ABLF02042774 GBBI01004549 JAC14163.1 GAMC01016174 JAB90381.1 GDHF01023027 JAI29287.1 GBXI01005773 JAD08519.1 GBBI01004477 JAC14235.1 ABLF02005127 ABLF02005129 ABLF02007401 ABLF02011059 GBXI01006173 JAD08119.1 QOIP01000004 RLU23736.1 GAMC01020207 JAB86348.1 GAMC01007326 JAB99229.1 GBXI01004632 JAD09660.1 AGBW02009151 OWR51557.1 ABLF02005846 KQ980852 KYN12192.1 GDHF01031400 JAI20914.1 CH478509 EJY58114.1 AJWK01016333 GFDF01007768 JAV06316.1 KQ971327 KYB28419.1 GDHF01019733 JAI32581.1 AJWK01024440 GGFM01002109 MBW22860.1 KQ977161 KYN05238.1 GAMC01015907 JAB90648.1 GAKP01019383 JAC39569.1 GGFM01002486 MBW23237.1 GEHC01000930 JAV46715.1 GFTR01008753 JAW07673.1 GBYB01008203 JAG77970.1 LBMM01018094 KMQ83877.1 HACL01000135 CFW94429.1 GBGD01000341 JAC88548.1 AAAB01008984 GGFJ01002513 MBW51654.1 GGFJ01002514 MBW51655.1 GL764261 EFZ18057.1 GGFK01005990 MBW39311.1 GAMC01002792 JAC03764.1 KQ980436 KYN16061.1 GGFJ01002620 MBW51761.1 GGFJ01002621 MBW51762.1 GGFJ01001974 MBW51115.1 KQ977505 KYN02276.1 GGFJ01002579 MBW51720.1 NWSH01001951 PCG69590.1 GGFJ01001952 MBW51093.1 Z27119 CAA81643.1 NNAY01005459 OXU16662.1 KQ971376 KYB25116.1 GBHO01014721 JAG28883.1 LBMM01011782 KMQ86633.1 GGFJ01005354 MBW54495.1
Proteomes
PRIDE
Pfam
Interpro
IPR001878
Znf_CCHC
+ More
IPR012337 RNaseH-like_sf
IPR036875 Znf_CCHC_sf
IPR036361 SAP_dom_sf
IPR001969 Aspartic_peptidase_AS
IPR001995 Peptidase_A2_cat
IPR041577 RT_RNaseH_2
IPR001584 Integrase_cat-core
IPR000477 RT_dom
IPR003034 SAP_dom
IPR036397 RNaseH_sf
IPR041588 Integrase_H2C2
IPR021109 Peptidase_aspartic_dom_sf
IPR041373 RT_RNaseH
IPR005162 Retrotrans_gag_dom
IPR002401 Cyt_P450_E_grp-I
IPR001128 Cyt_P450
IPR017972 Cyt_P450_CS
IPR036396 Cyt_P450_sf
IPR018061 Retropepsins
IPR025836 Zn_knuckle_CX2CX4HX4C
IPR019103 Peptidase_aspartic_DDI1-type
IPR034145 RP_RTVL-H-like
IPR021896 Transposase_37
IPR012337 RNaseH-like_sf
IPR036875 Znf_CCHC_sf
IPR036361 SAP_dom_sf
IPR001969 Aspartic_peptidase_AS
IPR001995 Peptidase_A2_cat
IPR041577 RT_RNaseH_2
IPR001584 Integrase_cat-core
IPR000477 RT_dom
IPR003034 SAP_dom
IPR036397 RNaseH_sf
IPR041588 Integrase_H2C2
IPR021109 Peptidase_aspartic_dom_sf
IPR041373 RT_RNaseH
IPR005162 Retrotrans_gag_dom
IPR002401 Cyt_P450_E_grp-I
IPR001128 Cyt_P450
IPR017972 Cyt_P450_CS
IPR036396 Cyt_P450_sf
IPR018061 Retropepsins
IPR025836 Zn_knuckle_CX2CX4HX4C
IPR019103 Peptidase_aspartic_DDI1-type
IPR034145 RP_RTVL-H-like
IPR021896 Transposase_37
SUPFAM
Gene 3D
ProteinModelPortal
A0A0J7K9X8
A0A023F013
A0A0J7KQL1
A0A023EZG2
A0A023EYH9
J9JZ77
+ More
A0A023EYF0 W8BB66 A0A0K8URI1 A0A0A1XB16 A0A023EY26 X1X261 J9JNX9 A0A0A1X9Z0 A0A3L8DTV3 W8APB2 W8C8Q1 A0A0A1XFE9 A0A212FCW5 J9JQ65 A0A151IWN5 A0A0K8U2D9 J9HJJ8 A0A1B0CKJ5 A0A1L8DIN4 A0A139WKB5 A0A0K8V149 A0A1B0CR74 A0A2M3Z2X0 A0A151IKW6 W8B0V9 A0A034VAM6 A0A2M3Z3Z7 A0A1W7R6L8 A0A224XHR1 A0A0C9Q2I4 A0A0J7K0N3 A0A0E4G8D8 A0A069DWV3 A0A1S4HEU1 A0A2M4BF33 A0A2M4BF27 E9IMZ6 A0A2M4AEU2 W8C3C0 A0A151J262 A0A182W804 A0A2M4BFM2 A0A2M4BFF1 A0A2M4BDJ8 A0A151II89 A0A2M4BF88 A0A2A4JDT3 A0A2M4BDL5 Q24262 A0A232EEC1 A0A139WB30 A0A0A9YCQ1 A0A0J7K890 A0A2M4BN67
A0A023EYF0 W8BB66 A0A0K8URI1 A0A0A1XB16 A0A023EY26 X1X261 J9JNX9 A0A0A1X9Z0 A0A3L8DTV3 W8APB2 W8C8Q1 A0A0A1XFE9 A0A212FCW5 J9JQ65 A0A151IWN5 A0A0K8U2D9 J9HJJ8 A0A1B0CKJ5 A0A1L8DIN4 A0A139WKB5 A0A0K8V149 A0A1B0CR74 A0A2M3Z2X0 A0A151IKW6 W8B0V9 A0A034VAM6 A0A2M3Z3Z7 A0A1W7R6L8 A0A224XHR1 A0A0C9Q2I4 A0A0J7K0N3 A0A0E4G8D8 A0A069DWV3 A0A1S4HEU1 A0A2M4BF33 A0A2M4BF27 E9IMZ6 A0A2M4AEU2 W8C3C0 A0A151J262 A0A182W804 A0A2M4BFM2 A0A2M4BFF1 A0A2M4BDJ8 A0A151II89 A0A2M4BF88 A0A2A4JDT3 A0A2M4BDL5 Q24262 A0A232EEC1 A0A139WB30 A0A0A9YCQ1 A0A0J7K890 A0A2M4BN67
PDB
2LLI
E-value=0.0038442,
Score=95
Ontologies
KEGG
GO
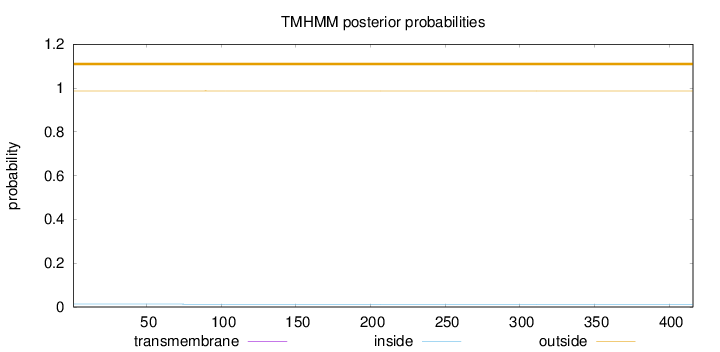
Topology
Length:
416
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00159
Exp number, first 60 AAs:
6e-05
Total prob of N-in:
0.01321
outside
1 - 416
Population Genetic Test Statistics
Pi
32.435705
Theta
21.656573
Tajima's D
3.614564
CLR
41.722675
CSRT
0.995400229988501
Interpretation
Possibly Balancing Selection
