Gene
KWMTBOMO14507
Annotation
PREDICTED:_uncharacterized_protein_LOC106720812_[Papilio_machaon]
Location in the cell
Nuclear Reliability : 2.129
Sequence
CDS
ATGATCACCCGCAGTAAAAGTAAAATCATGCCGACAAACCCTGCGACTAGTACTGGTTTGCAAGTTAATATTATACAACCTAGTACTTCCCAACATGTAGACAGCATTGCTCCGAGTGAGACGTCTAAGTCAAAATCATCACGCTCGTCATCCACAATTAAGGCTCAAAAGGCCGCCGTAGAGGCCCTCGCCCTGCGCCGACGTCTTGAGCACGAGCAGGAGCTGGCTGAACGTGAGAGGCAACGTGAAAGTCGACTGGCAGCACTCCGTCAGCAGGTAGAGGAGGGCGAACTGAAGGCCGAACTTGCCGCTCTCGACGCGGAAGCCGCGGCAAGTCACAGCAGTCGCAGTGGTTCTGCCAGGGTAAGCGTAGAAAGAACAGAAAGATGGGTACAAAGTTTAGAACAGCCAGTTTGCGTTGGCGAAAATACAGTTAATAGAGGCATTCCCGCAGTAGGGCGAGAGCAAGTAGTGGTTCCGCCTCCCGGCCACCAGGCACCTCTGCCGGCCACTGGGTCAACTGCATCCCCTGTTAACCGTTTTGTTCATCAGAAACAACTTTTAAACGTCACTAAAGCGTCCGAGGCGGCCGTACTTACGCGACCGCTCATTTCCACGGAGGTTGATCAGAGTAGTCTGCTTCAGGCTGTTGCTGACACTGCAGCTGCGGCCAAGGCGCTTGCTACTTCGCACCGTCATAAACATAAGATCGAATTGTTTCCTTTCGACGGAGACGCCTTAACATGGCTACATTTCAAACGTATTTTTGATTTAACCAAAAACAATTTTTCAAATATTGAAAATATTAGTCGGTTACAAAATGCACTTCGCGGCGCGGCCCGAGATGCAGTCGCGTCACTGTTGCTGGCTACAGACAACCCGAACGACATTATAAGAGCTCTGGAGGAGAACTTCGCTCGCCCGGAAATTATAGTATTTAAGGAAGTCTCCGCTCTTAAAAATCTACCACGTTTGGGAAATGACTTGAAAGAGTTAATTACGATAACGAATAAGGTACGTAATTCTATATCTTCTTTTAAAATACTGAATCAAATTGAGTACTTGCATTCTCCAGAACTATTTCACGCCGTCTTAGGAAAATTAAACCCGGTTACTAAAGTAAGATGGACGGACTTTGCTATGCAAGATACTACAGGGCGTCCTAAATTAGAAATTATGTCCGATTTTTTGAGGCGCGAGTTAGATCAACTTATTCGGTTCGGTTTATCACCGGAATCTTGCTCGATCCAACATTCACCACAAAACACTCATCTAAACAAACCGTATAAACGGGAAAATATTCACATAGTTACACACTCTAAACTAAATTCTAAGGCGGCGTCTACACAAAAGCTTGAATGCGCGTTTTGCAAAAAAGATCACAATATTAAGTCATGTTCTGAGTTCAAAGCCTTATCTGTTGATGAAAGGTGGGTCTGGCTGCGTGAGGTAAAAGCGTGTTATAAATGTCTTAGGCGTAGTTCACATCAGTGGAAAACATGTAAAGTTAACCCTTGTGGTGTTGATGGGTGTCTAATACGTCATCACCCGTTAATGCATGGTAAAGAACCTGCCTCTTCTGTCAATACTATGTTCGACACGCAATCCACCGAAAATGATCATACGACTAAGCACCACGAACCTGCATCTGTCGAAAAGGTAATAGTTTCTACCACTTCTACAGGTGTTCAAGAACCCAGCTCCATCTTAACTCTGTGCCCGCGTGTATTTCTGAAAATCTTACCCGTTACTGTCTCTGGTCCCTCAGGAAGTTGCAATGTTTACGCACTTCTCGATGACGGTAGCACTGCTACGCTCATTGACTCAAGTATTGCTACTCAAATCGGTGCGACAGGGCCTCACCAGAAAATAACGGTTAATGGTATAGGCGGTCTCAGTAGAAACGCTACGATATCGTATGTAGATTTTCACATTAAGGGTAGGCACACACGAGACACATTTTTAGTGAAAAATGCGAGAGCTATGGAGTCTTTGTCTCTGCGTCCTCAAACGATTAGGCAGGAAACTATTGCTTCTTTCGCACATTTAGCGAATTTACCTTGCAACGTAACTTATGAAGATGCTACACCAACAGTTTTAATTGGTGCAGAGCATTGGCATTTGTCAATCAGCAGCGAGGTACGGTGTGGTGGTAAGAATGAACCCGTTGCCTGTTTAACTGCCTTAGGCTGGGTGCTTTATGGAATCGCGTCCAGTAAAACTAAATCAGTGGAGTTTGTTAATCACGGTGTGTGTATCGAACCTTCCGATGAGGAGAAATTAGACTCTTTTATTAGGGAACAATACAAAATTGATTCGTTAGGTATTTCTAAAAAAGAAACTCTACATTCTAAGTCAGATAAACGTGCAGTCGAGATTTTAGAAAAAACTGCTAGACGTCTCCCCACTTCTGGTCGATTTGAGGTGGGTTTGCCGTGGCGTGACGATATACAACGCATTCCCGACAGCTACCCTCAAGCCATGTCACGATTTTTAAGTCTAGAAAGGCGCATGGCTAAAGATGCTGACTTTGCAAAAGCATATAACGTGTTCATTTCAAACATGATCGCTAAAGGCTATGCAGAAGAATGTGCTCCCGAGTCATACTACGCCAACCACATTAAAGATAAACAATCTTCTTTTATGAGGCTCTATTTACCCCATTTTGGCGTGTATCACCCTCAGAAACGTAAGCTTCGTGTAGTTCACGACGCCGCAGCCACGAATGCGGGCGTCAGCCTAAACTCTTTGTTGCTACCTGGTCCAGACCTGCTTCAGTCCTTGTTAGGTATTTTACTTCGGTTTTGCGAGGGCCGTATTGCTCTCACAGCTGATATCCGGGAAATGTTCCCTCAAGTAAGGATCAGAGAGCAAGATAGGGACGCCCTCCGGTTTCTCTGGAGGTCAAGCAGAGACCAACCTATAAAGGAGTTTAGGATGACAGCCATCATTTTCGGTGCGTGCTGCAGTCCATTTATAGCGCAATTTATTAAAAATAAAAACGCTCGTGAGCACGAAAGTTTATTTCCTAAAGCAGCTCATGCTATTTTGTACAATCATTATATGGACGATTATATTGACTCTCTCGATGACATTCAAGAGGCAGCACAACTGGCGGCAGATATTGTTGCCGTTCACAATGATGCCTACTTCGAGATGCGGGGCTGGATTTCCAACGATCCATCAGCGCTAAAATTTATTCCTGATGATCTTAGGGCCACTCAACCGTCGTTGGAAATCAACCTTGGCAGCTGCTCTGCAAACAATATTAGAGCGTTGGGAGTCGAGAGCTTTTCCATAGAATTTCTTCATATCCTGAACTCGAAGCCTATTCCTCCGAACAGTCGTCTCAGAAAACTGTCACCCGCTTTGGGAGAGGACAAATTACTGCGGTTAGCCGGAAGAATTAAAGCCGCAGAAGGCATTGACCCTGACACACGCTTCCCCATCCTTCTTGACGGTCGTCACCCAATTATCTTACCTACTGCCTTGAACGATCACGATTTGGTCAGCCGTTCCGAATGGAGGAAAGCATTGAGGCTAGCGGATCATTTTTGGTCGCGTTGGATGCGCGAGGTTCTTCCGACCCTACAACCACGTGAGCAGTCAAGAGGCAACCGCAGTTCAACAGCTTTTCAAGAGGGTGACGTAGTAATTATTGCTGACTTCAACTTGCCCCGCGGAATATGGCCACGAGGACGTGTTGCGAAAGTGTACCCTGGGAAAGATGGCGTGATAAGGGTAGTGGACGTTGCAACAGCAGGGGGAATGCTACGCCGACCCGTACGAAAGCTGGTTAGGCTCTCCACCTAA
Protein
MITRSKSKIMPTNPATSTGLQVNIIQPSTSQHVDSIAPSETSKSKSSRSSSTIKAQKAAVEALALRRRLEHEQELAERERQRESRLAALRQQVEEGELKAELAALDAEAAASHSSRSGSARVSVERTERWVQSLEQPVCVGENTVNRGIPAVGREQVVVPPPGHQAPLPATGSTASPVNRFVHQKQLLNVTKASEAAVLTRPLISTEVDQSSLLQAVADTAAAAKALATSHRHKHKIELFPFDGDALTWLHFKRIFDLTKNNFSNIENISRLQNALRGAARDAVASLLLATDNPNDIIRALEENFARPEIIVFKEVSALKNLPRLGNDLKELITITNKVRNSISSFKILNQIEYLHSPELFHAVLGKLNPVTKVRWTDFAMQDTTGRPKLEIMSDFLRRELDQLIRFGLSPESCSIQHSPQNTHLNKPYKRENIHIVTHSKLNSKAASTQKLECAFCKKDHNIKSCSEFKALSVDERWVWLREVKACYKCLRRSSHQWKTCKVNPCGVDGCLIRHHPLMHGKEPASSVNTMFDTQSTENDHTTKHHEPASVEKVIVSTTSTGVQEPSSILTLCPRVFLKILPVTVSGPSGSCNVYALLDDGSTATLIDSSIATQIGATGPHQKITVNGIGGLSRNATISYVDFHIKGRHTRDTFLVKNARAMESLSLRPQTIRQETIASFAHLANLPCNVTYEDATPTVLIGAEHWHLSISSEVRCGGKNEPVACLTALGWVLYGIASSKTKSVEFVNHGVCIEPSDEEKLDSFIREQYKIDSLGISKKETLHSKSDKRAVEILEKTARRLPTSGRFEVGLPWRDDIQRIPDSYPQAMSRFLSLERRMAKDADFAKAYNVFISNMIAKGYAEECAPESYYANHIKDKQSSFMRLYLPHFGVYHPQKRKLRVVHDAAATNAGVSLNSLLLPGPDLLQSLLGILLRFCEGRIALTADIREMFPQVRIREQDRDALRFLWRSSRDQPIKEFRMTAIIFGACCSPFIAQFIKNKNAREHESLFPKAAHAILYNHYMDDYIDSLDDIQEAAQLAADIVAVHNDAYFEMRGWISNDPSALKFIPDDLRATQPSLEINLGSCSANNIRALGVESFSIEFLHILNSKPIPPNSRLRKLSPALGEDKLLRLAGRIKAAEGIDPDTRFPILLDGRHPIILPTALNDHDLVSRSEWRKALRLADHFWSRWMREVLPTLQPREQSRGNRSSTAFQEGDVVIIADFNLPRGIWPRGRVAKVYPGKDGVIRVVDVATAGGMLRRPVRKLVRLST
Summary
Uniprot
Pubmed
EMBL
Proteomes
Pfam
Interpro
IPR008042
Retrotrans_Pao
+ More
IPR005312 DUF1759
IPR040676 DUF5641
IPR036397 RNaseH_sf
IPR001584 Integrase_cat-core
IPR012337 RNaseH-like_sf
IPR001878 Znf_CCHC
IPR006094 Oxid_FAD_bind_N
IPR016166 FAD-bd_PCMH
IPR016164 FAD-linked_Oxase-like_C
IPR041588 Integrase_H2C2
IPR016170 Cytok_DH_C_sf
IPR015213 Cholesterol_OX_subst-bd
IPR036318 FAD-bd_PCMH-like_sf
IPR016169 FAD-bd_PCMH_sub2
IPR016167 FAD-bd_PCMH_sub1
IPR008737 Peptidase_asp_put
IPR002999 Tudor
IPR019786 Zinc_finger_PHD-type_CS
IPR019787 Znf_PHD-finger
IPR013083 Znf_RING/FYVE/PHD
IPR001965 Znf_PHD
IPR011011 Znf_FYVE_PHD
IPR005312 DUF1759
IPR040676 DUF5641
IPR036397 RNaseH_sf
IPR001584 Integrase_cat-core
IPR012337 RNaseH-like_sf
IPR001878 Znf_CCHC
IPR006094 Oxid_FAD_bind_N
IPR016166 FAD-bd_PCMH
IPR016164 FAD-linked_Oxase-like_C
IPR041588 Integrase_H2C2
IPR016170 Cytok_DH_C_sf
IPR015213 Cholesterol_OX_subst-bd
IPR036318 FAD-bd_PCMH-like_sf
IPR016169 FAD-bd_PCMH_sub2
IPR016167 FAD-bd_PCMH_sub1
IPR008737 Peptidase_asp_put
IPR002999 Tudor
IPR019786 Zinc_finger_PHD-type_CS
IPR019787 Znf_PHD-finger
IPR013083 Znf_RING/FYVE/PHD
IPR001965 Znf_PHD
IPR011011 Znf_FYVE_PHD
ProteinModelPortal
Ontologies
GO
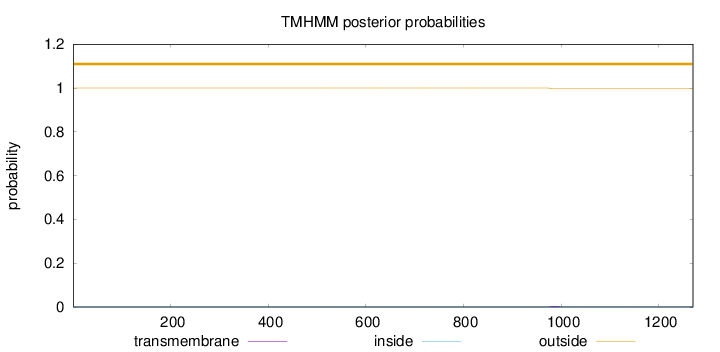
Topology
Length:
1270
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.06906
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00002
outside
1 - 1270
Population Genetic Test Statistics
Pi
444.716092
Theta
215.423581
Tajima's D
3.985758
CLR
0.593893
CSRT
0.99910004499775
Interpretation
Uncertain
