Gene
KWMTBOMO14395
Annotation
endonuclease-reverse_transcriptase_[Bombyx_mori]
Location in the cell
Mitochondrial Reliability : 3.323
Sequence
CDS
ATGAGAACCCTTATCTTCTCCATATTCTTGTACGCTTCTGAGACGTGGACCCTCAAGGCCCGTACCCGATCGAGAATTGACGCATTTGAAATGTGGTGTTGGCGCAAAATGTTGGGCGTATCATGGACGGAGCGCCGAACAAATGTGTCAATTCTCGAGGAGCTAAACGTCAAGACCCGTCTATCTACAATCTGTATGCAGCGTCTCCTCAAGTACTTCGGACACGTGGCCCGACAAAATCCTGAGAACTTGGAAAAATTAATGGTTGTAGGTCACGTGGAAGGAAAGAGAAGTCGAGGAAGATTGCCAACCAGGTGGACGGCCATTATCAAGGAGGCCACAAACACCAGCGTCCCGGGCGCCATTAAGCAGGCCGAATGCAGGCAACATTGGAGAAAGCTGGTGCAAAAACTGAACCAAGGTGGTCACGACCCTCAGTAA
Protein
MRTLIFSIFLYASETWTLKARTRSRIDAFEMWCWRKMLGVSWTERRTNVSILEELNVKTRLSTICMQRLLKYFGHVARQNPENLEKLMVVGHVEGKRSRGRLPTRWTAIIKEATNTSVPGAIKQAECRQHWRKLVQKLNQGGHDPQ
Summary
Uniprot
D7F174
A0A3S2L0R7
A0A2H1VNK5
D7F163
A0A2H1V742
D7F161
+ More
A0A2H1VU13 D7F175 D7F162 D7F167 D7F168 D7F172 D5LB39 A0A2W1BGD9 A0A2G8LJK7 A0A2G8K327 A0A2G8KTJ4 A0A2G8JE23 A0A2G8K796 A0A2G8K183 A0A2G8JVA1 A0A2G8JVI4 A0A2G8L6P4 W4YPU3 A0A2W1B3R1 A0A2H1W7G4 H2YWZ6 A0A085LZ21 H2YWZ5 A0A3S3RLW1 A0A3S3NR17 X1D4L4 W5NMM5 A0A085NA37 A0A194PHS9 W4ZJA4 A0A0B7BSZ9 A0A0B7BUR9 A0A0B7BT80 W4Y921 A0A3Q1MCX3 A0A3Q1LT03 A0A2A4J795 A0A0B7BVD9 A0A0B7BVD3 A0A0B7BSB6 W5MM77 A0A2W1BSS7 A0A3Q1M4E7 A0A3Q1LLH3 W5NIE2 J9LZK0 W4XFH1 X1WK45 A0A3S3P648 A0A3S1AZH6 A0A2S2NMQ2 A0A3S1HJZ0 A0A3Q1NB85 D5LB38 O97916 A0A3S0ZE77 A0A3S0ZVZ5 W4YNG7 A0A3Q1MZC9 A0A067QUH9 A0A067RQM5 W5NLK6 A0A3Q1M7G0 F6R5D0 X1XC23 A0A2S2PZ46 A0A3Q1MMS2 A0A2S2QAF2 A0A2S2P9Y7 A0A3Q1M4J9 W4Z5A1 A0A3Q1NE26 A0A2A4JXP9 A0A3Q1MNS1 A0A3Q1MIT2 A0A3Q1MJZ2 A0A2S2QE77 A0A3Q1MXT5 A0A067QXS5 A0A067QJX4 A0A067QZY5 A0A3Q1LGI2
A0A2H1VU13 D7F175 D7F162 D7F167 D7F168 D7F172 D5LB39 A0A2W1BGD9 A0A2G8LJK7 A0A2G8K327 A0A2G8KTJ4 A0A2G8JE23 A0A2G8K796 A0A2G8K183 A0A2G8JVA1 A0A2G8JVI4 A0A2G8L6P4 W4YPU3 A0A2W1B3R1 A0A2H1W7G4 H2YWZ6 A0A085LZ21 H2YWZ5 A0A3S3RLW1 A0A3S3NR17 X1D4L4 W5NMM5 A0A085NA37 A0A194PHS9 W4ZJA4 A0A0B7BSZ9 A0A0B7BUR9 A0A0B7BT80 W4Y921 A0A3Q1MCX3 A0A3Q1LT03 A0A2A4J795 A0A0B7BVD9 A0A0B7BVD3 A0A0B7BSB6 W5MM77 A0A2W1BSS7 A0A3Q1M4E7 A0A3Q1LLH3 W5NIE2 J9LZK0 W4XFH1 X1WK45 A0A3S3P648 A0A3S1AZH6 A0A2S2NMQ2 A0A3S1HJZ0 A0A3Q1NB85 D5LB38 O97916 A0A3S0ZE77 A0A3S0ZVZ5 W4YNG7 A0A3Q1MZC9 A0A067QUH9 A0A067RQM5 W5NLK6 A0A3Q1M7G0 F6R5D0 X1XC23 A0A2S2PZ46 A0A3Q1MMS2 A0A2S2QAF2 A0A2S2P9Y7 A0A3Q1M4J9 W4Z5A1 A0A3Q1NE26 A0A2A4JXP9 A0A3Q1MNS1 A0A3Q1MIT2 A0A3Q1MJZ2 A0A2S2QE77 A0A3Q1MXT5 A0A067QXS5 A0A067QJX4 A0A067QZY5 A0A3Q1LGI2
EMBL
FJ265559
ADI61827.1
RSAL01000289
RVE42931.1
ODYU01003523
SOQ42419.1
+ More
FJ265548 ADI61816.1 ODYU01001007 SOQ36619.1 FJ265546 ADI61814.1 ODYU01004401 SOQ44236.1 FJ265560 ADI61828.1 FJ265547 ADI61815.1 FJ265552 ADI61820.1 FJ265553 ADI61821.1 FJ265557 ADI61825.1 GU815090 ADF18553.1 KZ150203 PZC72267.1 MRZV01000056 PIK60437.1 MRZV01000940 PIK42365.1 MRZV01000378 PIK51329.1 MRZV01002322 PIK33998.1 MRZV01000817 PIK43880.1 MRZV01000994 PIK41715.1 MRZV01001216 PIK39650.1 MRZV01001207 PIK39730.1 MRZV01000196 PIK55924.1 AAGJ04055194 KZ150366 PZC71172.1 ODYU01006805 SOQ48998.1 KL363256 KL367985 KFD50217.1 KFD59415.1 NCKU01008836 RWS01655.1 NCKU01012182 RWS00163.1 BART01031562 GAH15691.1 AHAT01022938 KL363266 KL367526 KFD49675.1 KFD66333.1 KQ459603 KPI92872.1 AAGJ04021158 HACG01049248 HACG01049253 CEK96113.1 CEK96118.1 HACG01049251 HACG01049252 CEK96116.1 CEK96117.1 HACG01049244 HACG01049247 CEK96109.1 CEK96112.1 AAGJ04161766 NWSH01002736 PCG67606.1 HACG01049255 CEK96120.1 HACG01049245 CEK96110.1 HACG01049249 CEK96114.1 AHAT01015286 KZ149985 PZC75696.1 AHAT01009145 ABLF02042206 AAGJ04075245 ABLF02024992 ABLF02024994 ABLF02025002 ABLF02043975 NCKU01004395 RWS06003.1 RQTK01000685 RUS76136.1 GGMR01005851 MBY18470.1 RQTK01000360 RUS81011.1 GU815089 ADF18552.1 AJ132772 CAA10770.1 RQTK01000625 RUS76894.1 RQTK01000209 RUS84333.1 AAGJ04167631 KK852941 KDR13536.1 KK852482 KDR22935.1 AHAT01012943 ABLF02031909 GGMS01001437 MBY70640.1 GGMS01004979 MBY74182.1 GGMR01013640 MBY26259.1 AAGJ04128321 NWSH01000469 PCG76200.1 GGMS01006790 MBY75993.1 KK853131 KDR10935.1 KK853249 KDR09341.1 KK852843 KDR15156.1
FJ265548 ADI61816.1 ODYU01001007 SOQ36619.1 FJ265546 ADI61814.1 ODYU01004401 SOQ44236.1 FJ265560 ADI61828.1 FJ265547 ADI61815.1 FJ265552 ADI61820.1 FJ265553 ADI61821.1 FJ265557 ADI61825.1 GU815090 ADF18553.1 KZ150203 PZC72267.1 MRZV01000056 PIK60437.1 MRZV01000940 PIK42365.1 MRZV01000378 PIK51329.1 MRZV01002322 PIK33998.1 MRZV01000817 PIK43880.1 MRZV01000994 PIK41715.1 MRZV01001216 PIK39650.1 MRZV01001207 PIK39730.1 MRZV01000196 PIK55924.1 AAGJ04055194 KZ150366 PZC71172.1 ODYU01006805 SOQ48998.1 KL363256 KL367985 KFD50217.1 KFD59415.1 NCKU01008836 RWS01655.1 NCKU01012182 RWS00163.1 BART01031562 GAH15691.1 AHAT01022938 KL363266 KL367526 KFD49675.1 KFD66333.1 KQ459603 KPI92872.1 AAGJ04021158 HACG01049248 HACG01049253 CEK96113.1 CEK96118.1 HACG01049251 HACG01049252 CEK96116.1 CEK96117.1 HACG01049244 HACG01049247 CEK96109.1 CEK96112.1 AAGJ04161766 NWSH01002736 PCG67606.1 HACG01049255 CEK96120.1 HACG01049245 CEK96110.1 HACG01049249 CEK96114.1 AHAT01015286 KZ149985 PZC75696.1 AHAT01009145 ABLF02042206 AAGJ04075245 ABLF02024992 ABLF02024994 ABLF02025002 ABLF02043975 NCKU01004395 RWS06003.1 RQTK01000685 RUS76136.1 GGMR01005851 MBY18470.1 RQTK01000360 RUS81011.1 GU815089 ADF18552.1 AJ132772 CAA10770.1 RQTK01000625 RUS76894.1 RQTK01000209 RUS84333.1 AAGJ04167631 KK852941 KDR13536.1 KK852482 KDR22935.1 AHAT01012943 ABLF02031909 GGMS01001437 MBY70640.1 GGMS01004979 MBY74182.1 GGMR01013640 MBY26259.1 AAGJ04128321 NWSH01000469 PCG76200.1 GGMS01006790 MBY75993.1 KK853131 KDR10935.1 KK853249 KDR09341.1 KK852843 KDR15156.1
Proteomes
Pfam
Interpro
Gene 3D
ProteinModelPortal
D7F174
A0A3S2L0R7
A0A2H1VNK5
D7F163
A0A2H1V742
D7F161
+ More
A0A2H1VU13 D7F175 D7F162 D7F167 D7F168 D7F172 D5LB39 A0A2W1BGD9 A0A2G8LJK7 A0A2G8K327 A0A2G8KTJ4 A0A2G8JE23 A0A2G8K796 A0A2G8K183 A0A2G8JVA1 A0A2G8JVI4 A0A2G8L6P4 W4YPU3 A0A2W1B3R1 A0A2H1W7G4 H2YWZ6 A0A085LZ21 H2YWZ5 A0A3S3RLW1 A0A3S3NR17 X1D4L4 W5NMM5 A0A085NA37 A0A194PHS9 W4ZJA4 A0A0B7BSZ9 A0A0B7BUR9 A0A0B7BT80 W4Y921 A0A3Q1MCX3 A0A3Q1LT03 A0A2A4J795 A0A0B7BVD9 A0A0B7BVD3 A0A0B7BSB6 W5MM77 A0A2W1BSS7 A0A3Q1M4E7 A0A3Q1LLH3 W5NIE2 J9LZK0 W4XFH1 X1WK45 A0A3S3P648 A0A3S1AZH6 A0A2S2NMQ2 A0A3S1HJZ0 A0A3Q1NB85 D5LB38 O97916 A0A3S0ZE77 A0A3S0ZVZ5 W4YNG7 A0A3Q1MZC9 A0A067QUH9 A0A067RQM5 W5NLK6 A0A3Q1M7G0 F6R5D0 X1XC23 A0A2S2PZ46 A0A3Q1MMS2 A0A2S2QAF2 A0A2S2P9Y7 A0A3Q1M4J9 W4Z5A1 A0A3Q1NE26 A0A2A4JXP9 A0A3Q1MNS1 A0A3Q1MIT2 A0A3Q1MJZ2 A0A2S2QE77 A0A3Q1MXT5 A0A067QXS5 A0A067QJX4 A0A067QZY5 A0A3Q1LGI2
A0A2H1VU13 D7F175 D7F162 D7F167 D7F168 D7F172 D5LB39 A0A2W1BGD9 A0A2G8LJK7 A0A2G8K327 A0A2G8KTJ4 A0A2G8JE23 A0A2G8K796 A0A2G8K183 A0A2G8JVA1 A0A2G8JVI4 A0A2G8L6P4 W4YPU3 A0A2W1B3R1 A0A2H1W7G4 H2YWZ6 A0A085LZ21 H2YWZ5 A0A3S3RLW1 A0A3S3NR17 X1D4L4 W5NMM5 A0A085NA37 A0A194PHS9 W4ZJA4 A0A0B7BSZ9 A0A0B7BUR9 A0A0B7BT80 W4Y921 A0A3Q1MCX3 A0A3Q1LT03 A0A2A4J795 A0A0B7BVD9 A0A0B7BVD3 A0A0B7BSB6 W5MM77 A0A2W1BSS7 A0A3Q1M4E7 A0A3Q1LLH3 W5NIE2 J9LZK0 W4XFH1 X1WK45 A0A3S3P648 A0A3S1AZH6 A0A2S2NMQ2 A0A3S1HJZ0 A0A3Q1NB85 D5LB38 O97916 A0A3S0ZE77 A0A3S0ZVZ5 W4YNG7 A0A3Q1MZC9 A0A067QUH9 A0A067RQM5 W5NLK6 A0A3Q1M7G0 F6R5D0 X1XC23 A0A2S2PZ46 A0A3Q1MMS2 A0A2S2QAF2 A0A2S2P9Y7 A0A3Q1M4J9 W4Z5A1 A0A3Q1NE26 A0A2A4JXP9 A0A3Q1MNS1 A0A3Q1MIT2 A0A3Q1MJZ2 A0A2S2QE77 A0A3Q1MXT5 A0A067QXS5 A0A067QJX4 A0A067QZY5 A0A3Q1LGI2
Ontologies
Topology
SignalP
Position: 1 - 17,
Likelihood: 0.656338
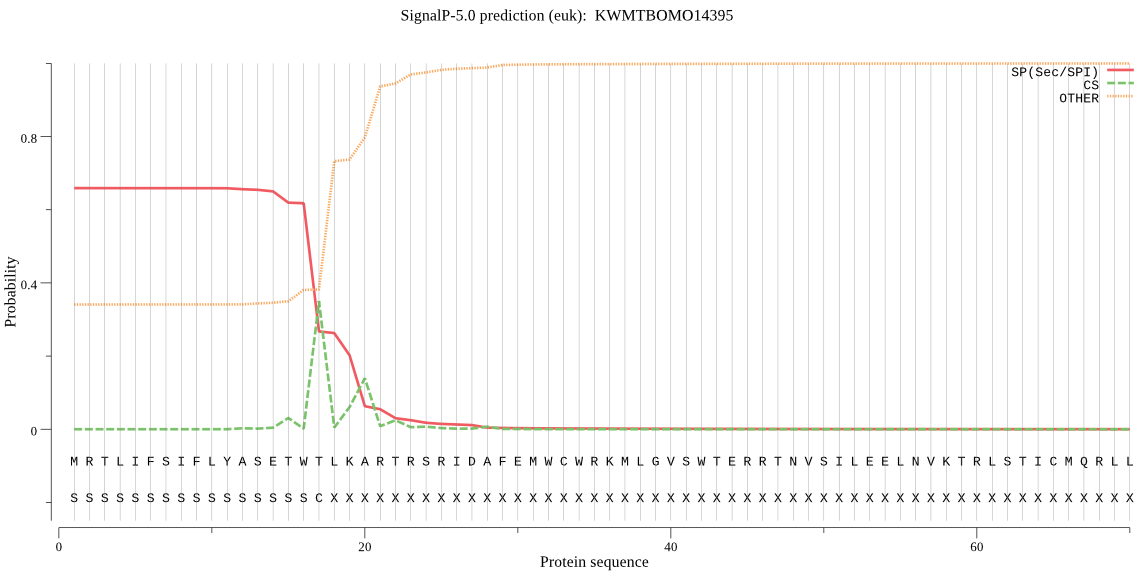
Length:
146
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00437
Exp number, first 60 AAs:
0.00437
Total prob of N-in:
0.03339
outside
1 - 146
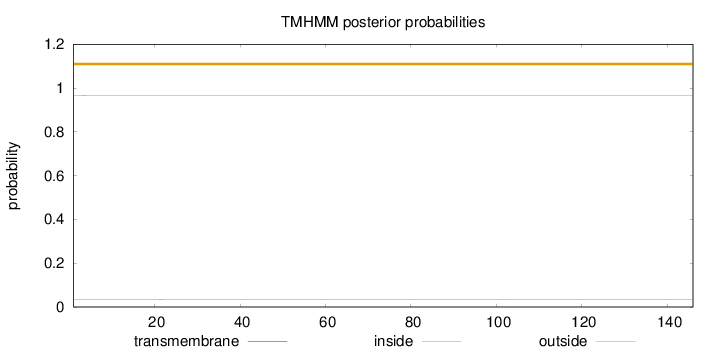
Population Genetic Test Statistics
Pi
278.360231
Theta
181.19229
Tajima's D
1.687308
CLR
0
CSRT
0.826708664566772
Interpretation
Uncertain
