Gene
KWMTBOMO14381
Pre Gene Modal
BGIBMGA013308
Annotation
PREDICTED:_irregular_chiasm_C-roughest_protein-like_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 1.112 Nuclear Reliability : 1.729
Sequence
CDS
ATGTTGTTTATCTTTTTTGTTAGTGTCAGTGCCAGTGTCAGTGCGTTTGGGGAGCAACGGTTCGCGCTAGAACCTCAAGATCAGACCGCCATATTAGGTTCCAGAGTGATCCTTCCGTGCAGAGTTGAGGAGAAGGCTGGGCAACTTCAGTGGACTAAGGATGACTTTGGACTCGGTACACACCGCCATCTGGCCGGATACGAGCGCTACAAAATGATTGGCAGCGACGAAGAAGGCGACTACTCGTTGGACATCAGAGAAGTTACGCTGGATGACGACGCAGCATATCAATGCCAAGTTAGCTCTGGACCGAGGGGTGAGCCAGCAATTCGGTCAAGGTACGCCCGACTGACGGTGCTGGTACCACCAGATCCTCCAAAGATACTTCGGGGGCCCGTCATTCCAGCTATAGAAGACAGAGAGCTGGATTTGGAGTGTGTTTCAGTAGGAGGAAAACCTCCAGCGGAGATAACGTGGGTCGACAATGACGGTGGAGTTTTGACTCAAGGTGTCACTTATACCGTGGAACCGATGGGTGATGGCAGAAGATTTACGGCGAGGTCTACCCTCAGAATACGTCCCAGACGCCAACACCACAATCAGACTTTTACTTGCCAAGCGCAGAACACTGCAGACAGGGCGTATAGATCTGTCAACGTCAAATTACAAGTCCAATACGCGCCTAAGGTCCGTCTCTTCATGAAATCCGGGTCAATAAAAGGGAGAATACAAGAAGGGGAAACATTAATAATAGGGTGCCAATCGAATGCCAACCCTAGTAATGTATCTTACAAATGGTACATAAACAATGATGAAGCAATTGGAGAAGTGAAAAATGAATTGGTAAGTATTCAATGA
Protein
MLFIFFVSVSASVSAFGEQRFALEPQDQTAILGSRVILPCRVEEKAGQLQWTKDDFGLGTHRHLAGYERYKMIGSDEEGDYSLDIREVTLDDDAAYQCQVSSGPRGEPAIRSRYARLTVLVPPDPPKILRGPVIPAIEDRELDLECVSVGGKPPAEITWVDNDGGVLTQGVTYTVEPMGDGRRFTARSTLRIRPRRQHHNQTFTCQAQNTADRAYRSVNVKLQVQYAPKVRLFMKSGSIKGRIQEGETLIIGCQSNANPSNVSYKWYINNDEAIGEVKNELVSIQ
Summary
Uniprot
A0A2A4K797
A0A2W1BS56
A0A194QH44
A0A2W1BXG4
A0A2A4K6N5
A0A212EGX2
+ More
A0A0N1IFK1 A0A182TDV2 A0A0T6BEN3 A0A139WL53 A0A139WKZ8 D6WHF2 A0A0L0C118 A0A182KLH1 Q8MSN7 A0A182WTP6 C4IXX7 A0A1Y1MDY1 A0A1Y1MGV5 A0A084VGX9 N6UM56 U4UA94 A0A182KDT0 Q17HA9 B0WI75 A0A182SHI5 A0A182XXZ4 A0A1I8M630 Q29HP2 A0A0R3P510 A0A1I8PVS3 B3N1W4 A0A182XXZ3 B4GYL5 A0A3B0K780 A0A182LX93 Q17HB1 A0A1A9XTM1 A0A182NVF9 A0A182RNF1 A0A1B0FNC9 A0A1A9V0N2 A0A182IAD7 B4IA24 A0A182UYI5 A0A1W4WYL8 A0A1A9ZLV7 A0A1B0B3Y8 A0A1W4WYV5 A0A1B6GC68 A0A182RCJ2 Q7QEY7 A0A182W5C0 A0A182NVG0 A0A182U923 A0A1A9XDY6 A0A1B0B8K6 A0A182GTF8 A0A1B0CQR5 Q7QEY8 A0A1A9WGC4 A0A0A1XF74 A0A182PBZ6 B4NDY0 A0A182QTQ6 A0A182VPE7 A0A084VGV9 B3NTZ7 A0A1A9V0M1 B4Q0T5 A0A182W5B9 W8BE24 A0A0K8VDK1 Q9W4T9 O97174 Q9N9Y9 A0A182JDB9 A0A1B0A8F0 A0A1I8NE46 A0A3B0K2L0 A0A1I8NE48 A0A182R0F3 A0A336LIT5 A0A1I8PJC8 A0A182J1W2 A0A1W4VSX2 A0A182GAK6 A0A034WIU0 A0A182GHE0 A0A182PL60 B5DM92 A0A0K8WLP0 A0A0K8UAG8 A0A0R3NZM2 A0A182SR59 A0A1A9WL55 W8AUB9 A0A0M4EUN0 B4L899 B4NDX8
A0A0N1IFK1 A0A182TDV2 A0A0T6BEN3 A0A139WL53 A0A139WKZ8 D6WHF2 A0A0L0C118 A0A182KLH1 Q8MSN7 A0A182WTP6 C4IXX7 A0A1Y1MDY1 A0A1Y1MGV5 A0A084VGX9 N6UM56 U4UA94 A0A182KDT0 Q17HA9 B0WI75 A0A182SHI5 A0A182XXZ4 A0A1I8M630 Q29HP2 A0A0R3P510 A0A1I8PVS3 B3N1W4 A0A182XXZ3 B4GYL5 A0A3B0K780 A0A182LX93 Q17HB1 A0A1A9XTM1 A0A182NVF9 A0A182RNF1 A0A1B0FNC9 A0A1A9V0N2 A0A182IAD7 B4IA24 A0A182UYI5 A0A1W4WYL8 A0A1A9ZLV7 A0A1B0B3Y8 A0A1W4WYV5 A0A1B6GC68 A0A182RCJ2 Q7QEY7 A0A182W5C0 A0A182NVG0 A0A182U923 A0A1A9XDY6 A0A1B0B8K6 A0A182GTF8 A0A1B0CQR5 Q7QEY8 A0A1A9WGC4 A0A0A1XF74 A0A182PBZ6 B4NDY0 A0A182QTQ6 A0A182VPE7 A0A084VGV9 B3NTZ7 A0A1A9V0M1 B4Q0T5 A0A182W5B9 W8BE24 A0A0K8VDK1 Q9W4T9 O97174 Q9N9Y9 A0A182JDB9 A0A1B0A8F0 A0A1I8NE46 A0A3B0K2L0 A0A1I8NE48 A0A182R0F3 A0A336LIT5 A0A1I8PJC8 A0A182J1W2 A0A1W4VSX2 A0A182GAK6 A0A034WIU0 A0A182GHE0 A0A182PL60 B5DM92 A0A0K8WLP0 A0A0K8UAG8 A0A0R3NZM2 A0A182SR59 A0A1A9WL55 W8AUB9 A0A0M4EUN0 B4L899 B4NDX8
Pubmed
28756777
26354079
22118469
18362917
19820115
26108605
+ More
20966253 28004739 24438588 23537049 17510324 25244985 25315136 15632085 17994087 12364791 26483478 14747013 17210077 25830018 17550304 24495485 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 24485456 26109357 26109356 10943839 11684659 25348373
20966253 28004739 24438588 23537049 17510324 25244985 25315136 15632085 17994087 12364791 26483478 14747013 17210077 25830018 17550304 24495485 10731132 12537568 12537572 12537573 12537574 16110336 17569856 17569867 24485456 26109357 26109356 10943839 11684659 25348373
EMBL
NWSH01000084
PCG79784.1
KZ149957
PZC76455.1
KQ458859
KPJ04832.1
+ More
PZC76453.1 NWSH01000121 PCG79332.1 AGBW02014999 OWR40735.1 KQ460254 KPJ16356.1 LJIG01001327 KRT85613.1 KQ971321 KYB28680.1 KYB28679.1 EFA00099.1 JRES01001043 KNC25985.1 AY118686 AAM50546.1 BT083408 ACQ83322.1 GEZM01036660 GEZM01036657 GEZM01036656 JAV82535.1 GEZM01036659 GEZM01036658 GEZM01036655 JAV82537.1 ATLV01013115 KE524840 KFB37223.1 APGK01018010 APGK01018011 APGK01018012 APGK01018013 APGK01018014 APGK01018015 KB740064 ENN81761.1 KB632165 ERL89278.1 CH477251 EAT46033.1 DS231944 EDS28290.1 CH379064 EAL31716.3 KRT06480.1 CH902663 EDV30005.1 CH479197 EDW27871.1 OUUW01000015 SPP88532.1 AXCM01000249 EAT46031.1 CCAG010019211 APCN01001155 CH480825 EDW44055.1 JXJN01008112 GECZ01009876 JAS59893.1 AAAB01008846 EAA06355.4 JXJN01009949 JXUM01018125 KQ560503 KXJ82106.1 AJWK01023927 EAA06357.5 GBXI01005154 JAD09138.1 CH964239 EDW81949.1 AXCN02002018 ATLV01013099 KFB37203.1 CH954180 EDV45705.1 CM000162 EDX01302.2 GAMC01011307 GAMC01011306 GAMC01011305 JAB95248.1 GDHF01015366 JAI36948.1 AE014298 AAF45847.2 AAN09598.1 AFH07219.1 AFH07220.1 AFH07221.1 AFH07222.1 AGB95042.1 AL035436 CAB36503.1 AF196553 AJ289882 AAF86308.1 CAB96574.2 SPP88525.1 AXCN02002013 UFQT01000015 SSX17766.1 JXUM01051192 JXUM01051193 JXUM01051194 JXUM01051195 KQ561678 KXJ77827.1 GAKP01004353 JAC54599.1 JXUM01063713 KQ562264 KXJ76278.1 EDY72610.2 GDHF01000554 JAI51760.1 GDHF01032907 GDHF01028726 GDHF01010672 JAI19407.1 JAI23588.1 JAI41642.1 KRT06478.1 GAMC01016710 GAMC01016708 JAB89847.1 CP012528 ALC48822.1 CH933814 EDW05674.2 EDW81947.2
PZC76453.1 NWSH01000121 PCG79332.1 AGBW02014999 OWR40735.1 KQ460254 KPJ16356.1 LJIG01001327 KRT85613.1 KQ971321 KYB28680.1 KYB28679.1 EFA00099.1 JRES01001043 KNC25985.1 AY118686 AAM50546.1 BT083408 ACQ83322.1 GEZM01036660 GEZM01036657 GEZM01036656 JAV82535.1 GEZM01036659 GEZM01036658 GEZM01036655 JAV82537.1 ATLV01013115 KE524840 KFB37223.1 APGK01018010 APGK01018011 APGK01018012 APGK01018013 APGK01018014 APGK01018015 KB740064 ENN81761.1 KB632165 ERL89278.1 CH477251 EAT46033.1 DS231944 EDS28290.1 CH379064 EAL31716.3 KRT06480.1 CH902663 EDV30005.1 CH479197 EDW27871.1 OUUW01000015 SPP88532.1 AXCM01000249 EAT46031.1 CCAG010019211 APCN01001155 CH480825 EDW44055.1 JXJN01008112 GECZ01009876 JAS59893.1 AAAB01008846 EAA06355.4 JXJN01009949 JXUM01018125 KQ560503 KXJ82106.1 AJWK01023927 EAA06357.5 GBXI01005154 JAD09138.1 CH964239 EDW81949.1 AXCN02002018 ATLV01013099 KFB37203.1 CH954180 EDV45705.1 CM000162 EDX01302.2 GAMC01011307 GAMC01011306 GAMC01011305 JAB95248.1 GDHF01015366 JAI36948.1 AE014298 AAF45847.2 AAN09598.1 AFH07219.1 AFH07220.1 AFH07221.1 AFH07222.1 AGB95042.1 AL035436 CAB36503.1 AF196553 AJ289882 AAF86308.1 CAB96574.2 SPP88525.1 AXCN02002013 UFQT01000015 SSX17766.1 JXUM01051192 JXUM01051193 JXUM01051194 JXUM01051195 KQ561678 KXJ77827.1 GAKP01004353 JAC54599.1 JXUM01063713 KQ562264 KXJ76278.1 EDY72610.2 GDHF01000554 JAI51760.1 GDHF01032907 GDHF01028726 GDHF01010672 JAI19407.1 JAI23588.1 JAI41642.1 KRT06478.1 GAMC01016710 GAMC01016708 JAB89847.1 CP012528 ALC48822.1 CH933814 EDW05674.2 EDW81947.2
Proteomes
UP000218220
UP000053268
UP000007151
UP000053240
UP000075902
UP000007266
+ More
UP000037069 UP000075882 UP000076407 UP000030765 UP000019118 UP000030742 UP000075881 UP000008820 UP000002320 UP000075901 UP000076408 UP000095301 UP000001819 UP000095300 UP000007801 UP000008744 UP000268350 UP000075883 UP000092443 UP000075884 UP000075900 UP000092444 UP000078200 UP000075840 UP000001292 UP000075903 UP000192223 UP000092445 UP000092460 UP000007062 UP000075920 UP000069940 UP000249989 UP000092461 UP000091820 UP000075885 UP000007798 UP000075886 UP000008711 UP000002282 UP000000803 UP000075880 UP000192221 UP000092553 UP000009192
UP000037069 UP000075882 UP000076407 UP000030765 UP000019118 UP000030742 UP000075881 UP000008820 UP000002320 UP000075901 UP000076408 UP000095301 UP000001819 UP000095300 UP000007801 UP000008744 UP000268350 UP000075883 UP000092443 UP000075884 UP000075900 UP000092444 UP000078200 UP000075840 UP000001292 UP000075903 UP000192223 UP000092445 UP000092460 UP000007062 UP000075920 UP000069940 UP000249989 UP000092461 UP000091820 UP000075885 UP000007798 UP000075886 UP000008711 UP000002282 UP000000803 UP000075880 UP000192221 UP000092553 UP000009192
Interpro
SUPFAM
SSF48726
SSF48726
Gene 3D
ProteinModelPortal
A0A2A4K797
A0A2W1BS56
A0A194QH44
A0A2W1BXG4
A0A2A4K6N5
A0A212EGX2
+ More
A0A0N1IFK1 A0A182TDV2 A0A0T6BEN3 A0A139WL53 A0A139WKZ8 D6WHF2 A0A0L0C118 A0A182KLH1 Q8MSN7 A0A182WTP6 C4IXX7 A0A1Y1MDY1 A0A1Y1MGV5 A0A084VGX9 N6UM56 U4UA94 A0A182KDT0 Q17HA9 B0WI75 A0A182SHI5 A0A182XXZ4 A0A1I8M630 Q29HP2 A0A0R3P510 A0A1I8PVS3 B3N1W4 A0A182XXZ3 B4GYL5 A0A3B0K780 A0A182LX93 Q17HB1 A0A1A9XTM1 A0A182NVF9 A0A182RNF1 A0A1B0FNC9 A0A1A9V0N2 A0A182IAD7 B4IA24 A0A182UYI5 A0A1W4WYL8 A0A1A9ZLV7 A0A1B0B3Y8 A0A1W4WYV5 A0A1B6GC68 A0A182RCJ2 Q7QEY7 A0A182W5C0 A0A182NVG0 A0A182U923 A0A1A9XDY6 A0A1B0B8K6 A0A182GTF8 A0A1B0CQR5 Q7QEY8 A0A1A9WGC4 A0A0A1XF74 A0A182PBZ6 B4NDY0 A0A182QTQ6 A0A182VPE7 A0A084VGV9 B3NTZ7 A0A1A9V0M1 B4Q0T5 A0A182W5B9 W8BE24 A0A0K8VDK1 Q9W4T9 O97174 Q9N9Y9 A0A182JDB9 A0A1B0A8F0 A0A1I8NE46 A0A3B0K2L0 A0A1I8NE48 A0A182R0F3 A0A336LIT5 A0A1I8PJC8 A0A182J1W2 A0A1W4VSX2 A0A182GAK6 A0A034WIU0 A0A182GHE0 A0A182PL60 B5DM92 A0A0K8WLP0 A0A0K8UAG8 A0A0R3NZM2 A0A182SR59 A0A1A9WL55 W8AUB9 A0A0M4EUN0 B4L899 B4NDX8
A0A0N1IFK1 A0A182TDV2 A0A0T6BEN3 A0A139WL53 A0A139WKZ8 D6WHF2 A0A0L0C118 A0A182KLH1 Q8MSN7 A0A182WTP6 C4IXX7 A0A1Y1MDY1 A0A1Y1MGV5 A0A084VGX9 N6UM56 U4UA94 A0A182KDT0 Q17HA9 B0WI75 A0A182SHI5 A0A182XXZ4 A0A1I8M630 Q29HP2 A0A0R3P510 A0A1I8PVS3 B3N1W4 A0A182XXZ3 B4GYL5 A0A3B0K780 A0A182LX93 Q17HB1 A0A1A9XTM1 A0A182NVF9 A0A182RNF1 A0A1B0FNC9 A0A1A9V0N2 A0A182IAD7 B4IA24 A0A182UYI5 A0A1W4WYL8 A0A1A9ZLV7 A0A1B0B3Y8 A0A1W4WYV5 A0A1B6GC68 A0A182RCJ2 Q7QEY7 A0A182W5C0 A0A182NVG0 A0A182U923 A0A1A9XDY6 A0A1B0B8K6 A0A182GTF8 A0A1B0CQR5 Q7QEY8 A0A1A9WGC4 A0A0A1XF74 A0A182PBZ6 B4NDY0 A0A182QTQ6 A0A182VPE7 A0A084VGV9 B3NTZ7 A0A1A9V0M1 B4Q0T5 A0A182W5B9 W8BE24 A0A0K8VDK1 Q9W4T9 O97174 Q9N9Y9 A0A182JDB9 A0A1B0A8F0 A0A1I8NE46 A0A3B0K2L0 A0A1I8NE48 A0A182R0F3 A0A336LIT5 A0A1I8PJC8 A0A182J1W2 A0A1W4VSX2 A0A182GAK6 A0A034WIU0 A0A182GHE0 A0A182PL60 B5DM92 A0A0K8WLP0 A0A0K8UAG8 A0A0R3NZM2 A0A182SR59 A0A1A9WL55 W8AUB9 A0A0M4EUN0 B4L899 B4NDX8
PDB
4OF8
E-value=6.18773e-83,
Score=781
Ontologies
GO
GO:0016021
GO:0005917
GO:1901739
GO:0097206
GO:0007157
GO:0036059
GO:0050839
GO:0005912
GO:0001745
GO:0005886
GO:0007156
GO:0036058
GO:0007520
GO:0009986
GO:0007523
GO:0016202
GO:0061321
GO:0030176
GO:2001256
GO:0042802
GO:0005913
GO:0030165
GO:0016324
GO:0005515
GO:0005262
GO:0005219
GO:0006816
GO:0005507
GO:0004713
GO:0016491
GO:0015930
Topology
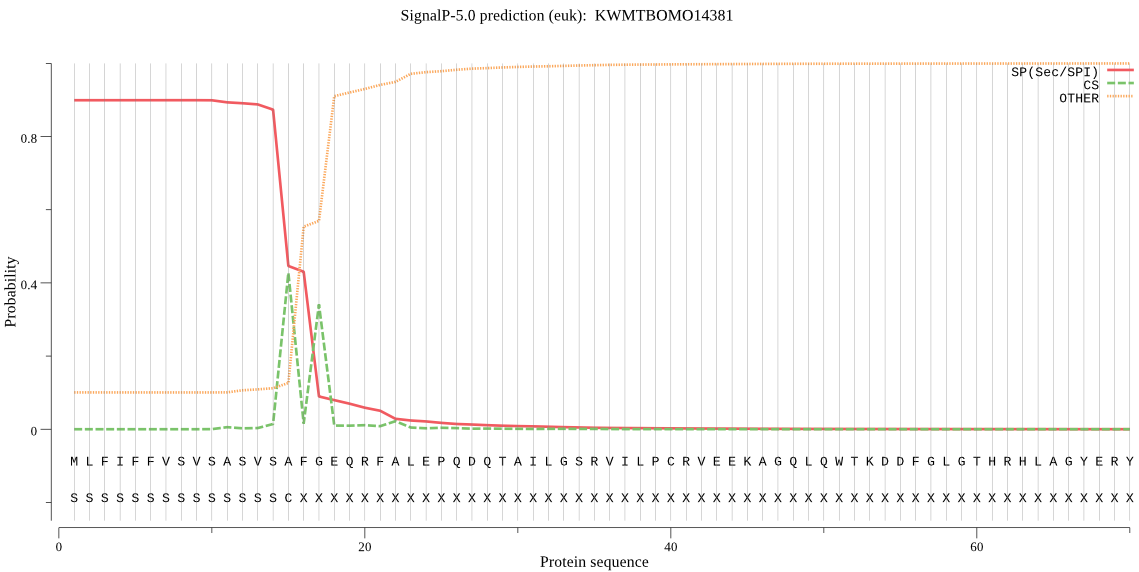
SignalP
Position: 1 - 15,
Likelihood: 0.898563
Length:
285
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.02539
Exp number, first 60 AAs:
0.02539
Total prob of N-in:
0.00386
outside
1 - 285

Population Genetic Test Statistics
Pi
258.478592
Theta
176.870521
Tajima's D
1.490358
CLR
0.169872
CSRT
0.784210789460527
Interpretation
Uncertain
