Gene
KWMTBOMO14328
Annotation
mariner_transposase_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 1.29 Mitochondrial Reliability : 1.245 Nuclear Reliability : 1.513
Sequence
CDS
ATGGAATTGACTCGAGAAAATTCAAGAGCGATGATTTATTATGACTTTCGAAGTGGTTTAACACAAAAACAGTGTGTTGACCGGATGATTTCTGCATTTGGTGATGAAGCCCTATCCAAAACCACAATTTATCGCTGGTTTACTGAGTTTCAACGTGGACGTGTCAAGCTCAGTGATGTTCCCCGTCAAGGTCGTCCAAAAACTGCAGTCACCCAAGAAAACGTTGATGCTGTGCGTAAGCTGATTGAGGAAGATCGACATGTGACATACCGCGAAATTCAGGCAACTTTAGACATTGGCATGAGTCAAATACAAATAATCCTGCATGAACAATTAGGTGTAAAAAAGTTGTTTTACCGATGGATACCGCATTCGCTCTGTGAAGAGCAAAAAGCGGCTCGCGTTACTTGGTGCGTCAGAACTCTCGAAAGATTCCACGCAGGATCCTCAAATGCTGTATACAACATTGTATCAGGTGACGAATCCTGGATATACGCGTACGAACCCGAAACAAAAAACCAGTCACGAGTTTGGGTGTTCGAAAATGAGTTAAAGCCAACAAAAATTGTTCGTTCACGGAGTGTTGCAAAAAAAATGGTGGCCACGTTTATCTCCAAAGCCGGCCATGTTACGACTATTCCTCTTGAGGGACAAAGAACGGTTAATGCAGAATGGTATGCTAGCATTTTTTTGCCACAGGTCATTTCTGAACTCCGTAAAGAGAACTGCAACCGCCGCATCATCCTCCATCACGACAATGCGAGTTCTCACACCGCGCACAGAACAAAAGAGTTTTTAGAGCTAGAAAACATAGAATTATGA
Protein
MELTRENSRAMIYYDFRSGLTQKQCVDRMISAFGDEALSKTTIYRWFTEFQRGRVKLSDVPRQGRPKTAVTQENVDAVRKLIEEDRHVTYREIQATLDIGMSQIQIILHEQLGVKKLFYRWIPHSLCEEQKAARVTWCVRTLERFHAGSSNAVYNIVSGDESWIYAYEPETKNQSRVWVFENELKPTKIVRSRSVAKKMVATFISKAGHVTTIPLEGQRTVNAEWYASIFLPQVISELRKENCNRRIILHHDNASSHTAHRTKEFLELENIEL
Summary
Similarity
Belongs to the ETS family.
Uniprot
Q1HPJ3
A0A069DT60
A0A023F6H6
A0A1L5BY10
A0A1B6JMK6
A0A0V0G5J0
+ More
E2B0A5 A0A034VN11 A0A267ESW3 A0A034VJN3 A0A1I8J9Y1 Q0QXC1 A0A267EV83 A0A1I8GYJ3 Q13539 A0A1I8HUL9 B3CK19 A0A151WWS0 A0A0J7MQ55 A0A1L5BY07 A0A267GHL5 V5GGS5 R4G5I0 A0A2H9T3D9 A0A1L5BY05 B3CK18 A6GV71 A0A2J7Q027 B3CK17 B3CK26 B7ZDI8 B7ZDI6 B3CK24 B7ZDI7 A0A023F6Y7 A0A085NER3 A0A0J7JXI0 A6GV70 A0A224XT40 A0A2J7Q5D2 A6GV69 B3CK27 A6GV72 A0A2J7R1D0 A0A147BDA2 A0A085NES9 A0A1L5BY03 A0A1B6HD45 D2WLV0 A0A069DTF4 A0A069DT95 A0A085MJ32 B3CK30 A0A1L5BY08 A0A023F6P2 R4G4T8 A0A1I8I720 A0A085MGB5 A0A085LSG3 A0A267E0K0 A0A0J7K8P2 A0A267GB40 A0A267FT86 A0A2J7PHE2 A0A085MPC9 A0A0J7NFU3 A0A085NH43 A0A085MM23 B3CK20 A0A2J7PES0 A0A2J7RSZ3 B3CK23 A0A1L5BY09 R4FP54 R4FPS5 A0A085MLI2 A0A2S2PZK5 A0A0J7K193
E2B0A5 A0A034VN11 A0A267ESW3 A0A034VJN3 A0A1I8J9Y1 Q0QXC1 A0A267EV83 A0A1I8GYJ3 Q13539 A0A1I8HUL9 B3CK19 A0A151WWS0 A0A0J7MQ55 A0A1L5BY07 A0A267GHL5 V5GGS5 R4G5I0 A0A2H9T3D9 A0A1L5BY05 B3CK18 A6GV71 A0A2J7Q027 B3CK17 B3CK26 B7ZDI8 B7ZDI6 B3CK24 B7ZDI7 A0A023F6Y7 A0A085NER3 A0A0J7JXI0 A6GV70 A0A224XT40 A0A2J7Q5D2 A6GV69 B3CK27 A6GV72 A0A2J7R1D0 A0A147BDA2 A0A085NES9 A0A1L5BY03 A0A1B6HD45 D2WLV0 A0A069DTF4 A0A069DT95 A0A085MJ32 B3CK30 A0A1L5BY08 A0A023F6P2 R4G4T8 A0A1I8I720 A0A085MGB5 A0A085LSG3 A0A267E0K0 A0A0J7K8P2 A0A267GB40 A0A267FT86 A0A2J7PHE2 A0A085MPC9 A0A0J7NFU3 A0A085NH43 A0A085MM23 B3CK20 A0A2J7PES0 A0A2J7RSZ3 B3CK23 A0A1L5BY09 R4FP54 R4FPS5 A0A085MLI2 A0A2S2PZK5 A0A0J7K193
Pubmed
EMBL
DQ443409
ABF51498.1
GBGD01001983
JAC86906.1
GBBI01001865
JAC16847.1
+ More
KX930998 APL98291.1 GECU01007301 JAT00406.1 GECL01002811 JAP03313.1 GL444487 EFN60885.1 GAKP01015435 JAC43517.1 NIVC01001798 PAA63957.1 GAKP01015436 JAC43516.1 DQ174779 ABA55502.1 NIVC01001731 PAA64652.1 U49974 AAC52011.1 AM906136 CAP20050.1 KQ982684 KYQ52352.1 LBMM01023556 KMQ82700.1 KX931001 APL98294.1 NIVC01000364 PAA84702.1 GALX01005282 JAB63184.1 GAHY01000550 JAA76960.1 NSIT01000395 PJE77748.1 KX930994 APL98287.1 AM906135 CAP20049.1 AM231072 CAJ76986.1 NEVH01020327 PNF21945.1 AM906133 CAP20047.1 AM906145 CAP20059.1 AM231082 CAJ76996.1 AM231079 CAJ76993.1 AM906142 AM906144 CAP20056.1 AM231080 CAJ76994.1 GBBI01001710 JAC17002.1 KL367509 KFD67959.1 LBMM01023216 KMQ82764.1 AM231070 CAJ76984.1 GFTR01005295 JAW11131.1 NEVH01017576 PNF23785.1 AM231069 CAJ76983.1 AM906151 AM906152 CAP20065.1 AM231074 CAJ76988.1 NEVH01008208 PNF34637.1 GEGO01006651 JAR88753.1 KFD67975.1 KX930996 APL98289.1 GECU01035151 JAS72555.1 GQ398105 ADB28039.1 GBGD01001947 JAC86942.1 GBGD01001923 JAC86966.1 KL363189 KFD57228.1 AM906154 CAP20068.1 KX931000 APL98293.1 GBBI01001812 JAC16900.1 GAHY01001080 JAA76430.1 KL363194 KFD56261.1 KL363310 KFD47909.1 NIVC01002911 PAA54417.1 LBMM01011730 KMQ86664.1 NIVC01001708 NIVC01001487 NIVC01000426 NIVC01000107 PAA64861.1 PAA67372.1 PAA83241.1 PAA90594.1 NIVC01000831 PAA76287.1 NEVH01025141 PNF15743.1 KL363182 KFD59075.1 LBMM01005538 KMQ91440.1 KL367501 KFD68789.1 KL363185 KFD58269.1 AM906137 CAP20051.1 NEVH01027114 NEVH01027075 NEVH01026089 NEVH01026087 NEVH01024426 NEVH01021956 NEVH01021925 NEVH01015301 NEVH01013964 NEVH01013194 NEVH01011885 NEVH01009393 NEVH01007824 NEVH01007823 NEVH01006721 NEVH01006578 NEVH01005888 NEVH01005885 NEVH01005277 NEVH01002149 NEVH01001358 PNF13652.1 PNF13785.1 PNF14826.1 PNF14870.1 PNF17130.1 PNF18970.1 PNF19346.1 PNF27270.1 PNF28165.1 PNF28197.1 PNF30137.1 PNF31284.1 PNF33174.1 PNF35196.1 PNF35254.1 PNF37314.1 PNF37715.1 PNF38363.1 PNF38519.1 PNF39139.1 PNF42450.1 PNF42722.1 NEVH01000249 PNF43951.1 AM906141 CAP20055.1 KX930999 APL98292.1 GAHY01001054 JAA76456.1 GAHY01000734 JAA76776.1 KFD58078.1 GGMS01001658 MBY70861.1 LBMM01017492 KMQ84067.1
KX930998 APL98291.1 GECU01007301 JAT00406.1 GECL01002811 JAP03313.1 GL444487 EFN60885.1 GAKP01015435 JAC43517.1 NIVC01001798 PAA63957.1 GAKP01015436 JAC43516.1 DQ174779 ABA55502.1 NIVC01001731 PAA64652.1 U49974 AAC52011.1 AM906136 CAP20050.1 KQ982684 KYQ52352.1 LBMM01023556 KMQ82700.1 KX931001 APL98294.1 NIVC01000364 PAA84702.1 GALX01005282 JAB63184.1 GAHY01000550 JAA76960.1 NSIT01000395 PJE77748.1 KX930994 APL98287.1 AM906135 CAP20049.1 AM231072 CAJ76986.1 NEVH01020327 PNF21945.1 AM906133 CAP20047.1 AM906145 CAP20059.1 AM231082 CAJ76996.1 AM231079 CAJ76993.1 AM906142 AM906144 CAP20056.1 AM231080 CAJ76994.1 GBBI01001710 JAC17002.1 KL367509 KFD67959.1 LBMM01023216 KMQ82764.1 AM231070 CAJ76984.1 GFTR01005295 JAW11131.1 NEVH01017576 PNF23785.1 AM231069 CAJ76983.1 AM906151 AM906152 CAP20065.1 AM231074 CAJ76988.1 NEVH01008208 PNF34637.1 GEGO01006651 JAR88753.1 KFD67975.1 KX930996 APL98289.1 GECU01035151 JAS72555.1 GQ398105 ADB28039.1 GBGD01001947 JAC86942.1 GBGD01001923 JAC86966.1 KL363189 KFD57228.1 AM906154 CAP20068.1 KX931000 APL98293.1 GBBI01001812 JAC16900.1 GAHY01001080 JAA76430.1 KL363194 KFD56261.1 KL363310 KFD47909.1 NIVC01002911 PAA54417.1 LBMM01011730 KMQ86664.1 NIVC01001708 NIVC01001487 NIVC01000426 NIVC01000107 PAA64861.1 PAA67372.1 PAA83241.1 PAA90594.1 NIVC01000831 PAA76287.1 NEVH01025141 PNF15743.1 KL363182 KFD59075.1 LBMM01005538 KMQ91440.1 KL367501 KFD68789.1 KL363185 KFD58269.1 AM906137 CAP20051.1 NEVH01027114 NEVH01027075 NEVH01026089 NEVH01026087 NEVH01024426 NEVH01021956 NEVH01021925 NEVH01015301 NEVH01013964 NEVH01013194 NEVH01011885 NEVH01009393 NEVH01007824 NEVH01007823 NEVH01006721 NEVH01006578 NEVH01005888 NEVH01005885 NEVH01005277 NEVH01002149 NEVH01001358 PNF13652.1 PNF13785.1 PNF14826.1 PNF14870.1 PNF17130.1 PNF18970.1 PNF19346.1 PNF27270.1 PNF28165.1 PNF28197.1 PNF30137.1 PNF31284.1 PNF33174.1 PNF35196.1 PNF35254.1 PNF37314.1 PNF37715.1 PNF38363.1 PNF38519.1 PNF39139.1 PNF42450.1 PNF42722.1 NEVH01000249 PNF43951.1 AM906141 CAP20055.1 KX930999 APL98292.1 GAHY01001054 JAA76456.1 GAHY01000734 JAA76776.1 KFD58078.1 GGMS01001658 MBY70861.1 LBMM01017492 KMQ84067.1
Proteomes
Pfam
Interpro
IPR038717
Tc1-like_DDE_dom
+ More
IPR041426 Mos1_HTH
IPR001888 Transposase_1
IPR038765 Papain-like_cys_pep_sf
IPR036390 WH_DNA-bd_sf
IPR036388 WH-like_DNA-bd_sf
IPR000418 Ets_dom
IPR008271 Ser/Thr_kinase_AS
IPR011009 Kinase-like_dom_sf
IPR017441 Protein_kinase_ATP_BS
IPR000719 Prot_kinase_dom
IPR022165 PKK
IPR013543 Ca/CaM-dep_prot_kinase-assoc
IPR032710 NTF2-like_dom_sf
IPR004018 RPEL_repeat
IPR041426 Mos1_HTH
IPR001888 Transposase_1
IPR038765 Papain-like_cys_pep_sf
IPR036390 WH_DNA-bd_sf
IPR036388 WH-like_DNA-bd_sf
IPR000418 Ets_dom
IPR008271 Ser/Thr_kinase_AS
IPR011009 Kinase-like_dom_sf
IPR017441 Protein_kinase_ATP_BS
IPR000719 Prot_kinase_dom
IPR022165 PKK
IPR013543 Ca/CaM-dep_prot_kinase-assoc
IPR032710 NTF2-like_dom_sf
IPR004018 RPEL_repeat
Gene 3D
ProteinModelPortal
Q1HPJ3
A0A069DT60
A0A023F6H6
A0A1L5BY10
A0A1B6JMK6
A0A0V0G5J0
+ More
E2B0A5 A0A034VN11 A0A267ESW3 A0A034VJN3 A0A1I8J9Y1 Q0QXC1 A0A267EV83 A0A1I8GYJ3 Q13539 A0A1I8HUL9 B3CK19 A0A151WWS0 A0A0J7MQ55 A0A1L5BY07 A0A267GHL5 V5GGS5 R4G5I0 A0A2H9T3D9 A0A1L5BY05 B3CK18 A6GV71 A0A2J7Q027 B3CK17 B3CK26 B7ZDI8 B7ZDI6 B3CK24 B7ZDI7 A0A023F6Y7 A0A085NER3 A0A0J7JXI0 A6GV70 A0A224XT40 A0A2J7Q5D2 A6GV69 B3CK27 A6GV72 A0A2J7R1D0 A0A147BDA2 A0A085NES9 A0A1L5BY03 A0A1B6HD45 D2WLV0 A0A069DTF4 A0A069DT95 A0A085MJ32 B3CK30 A0A1L5BY08 A0A023F6P2 R4G4T8 A0A1I8I720 A0A085MGB5 A0A085LSG3 A0A267E0K0 A0A0J7K8P2 A0A267GB40 A0A267FT86 A0A2J7PHE2 A0A085MPC9 A0A0J7NFU3 A0A085NH43 A0A085MM23 B3CK20 A0A2J7PES0 A0A2J7RSZ3 B3CK23 A0A1L5BY09 R4FP54 R4FPS5 A0A085MLI2 A0A2S2PZK5 A0A0J7K193
E2B0A5 A0A034VN11 A0A267ESW3 A0A034VJN3 A0A1I8J9Y1 Q0QXC1 A0A267EV83 A0A1I8GYJ3 Q13539 A0A1I8HUL9 B3CK19 A0A151WWS0 A0A0J7MQ55 A0A1L5BY07 A0A267GHL5 V5GGS5 R4G5I0 A0A2H9T3D9 A0A1L5BY05 B3CK18 A6GV71 A0A2J7Q027 B3CK17 B3CK26 B7ZDI8 B7ZDI6 B3CK24 B7ZDI7 A0A023F6Y7 A0A085NER3 A0A0J7JXI0 A6GV70 A0A224XT40 A0A2J7Q5D2 A6GV69 B3CK27 A6GV72 A0A2J7R1D0 A0A147BDA2 A0A085NES9 A0A1L5BY03 A0A1B6HD45 D2WLV0 A0A069DTF4 A0A069DT95 A0A085MJ32 B3CK30 A0A1L5BY08 A0A023F6P2 R4G4T8 A0A1I8I720 A0A085MGB5 A0A085LSG3 A0A267E0K0 A0A0J7K8P2 A0A267GB40 A0A267FT86 A0A2J7PHE2 A0A085MPC9 A0A0J7NFU3 A0A085NH43 A0A085MM23 B3CK20 A0A2J7PES0 A0A2J7RSZ3 B3CK23 A0A1L5BY09 R4FP54 R4FPS5 A0A085MLI2 A0A2S2PZK5 A0A0J7K193
PDB
5HOO
E-value=2.77508e-18,
Score=224
Ontologies
GO
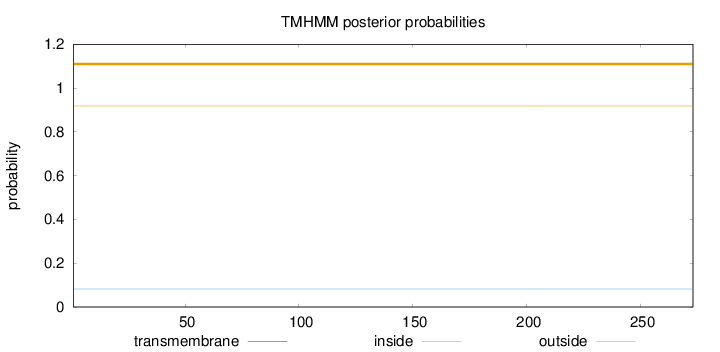
Topology
Subcellular location
Nucleus
Length:
273
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00408000000000001
Exp number, first 60 AAs:
0.00099
Total prob of N-in:
0.08183
outside
1 - 273
Population Genetic Test Statistics
Pi
36.950877
Theta
24.384714
Tajima's D
1.157794
CLR
0
CSRT
0.705414729263537
Interpretation
Uncertain
