Gene
KWMTBOMO13839
Annotation
PREDICTED:_uncharacterized_protein_LOC105841996_[Bombyx_mori]
Location in the cell
Extracellular Reliability : 1.968 Nuclear Reliability : 1.326
Sequence
CDS
ATGGCCCTGTACGGTGCTCCCGTGTGGTCGCCCGCACTCTCCGCGAGAAACGCGGCTCTGCTGCATAGAGCTCAGCGGGCGCTTGCGGTGAGGGTCATCAGGGGGTACCGCACCATCTCTCAAGATGTGGCCTGCGCTCTCGCCGGATCCCTACCCTGGGACCTCGAGGCCGAAATCTTGGCTGCGGTATATCGGCGTAAGGCGCAGGCCTTGAGTCGGGGGCGGTCGCCCGGCCCGTCGGCTGTCGGTCGGTGGAAGCGTGCTGCGCGTCATGTAGCATACGCTAAGTGGAGAGAGCGATTGCTGAAAGAACTTGGCCAAACTTCGGCCACCCGCCGGCGCACCCTCGAGGCCCTGGTGCCCGTGCTGGAGGCATGGTCGGATCGGCGTCACGGCGTACTCTCCTTTCACCTTGCGCAGGTCCTCTCGGGACATGGCTGTTTCGGGAGGTACCTGTGGAAGGTCTGCAGGAGAGAGCCACACCCGGGCTGCCACCAGTGCGGGCACCCGGACGATGACGCGCAGCACGCGCTCGAAACGTGCCCGCGTTGGGAGCACTCGCGGCGAGATCTCGTCGCGGTGCTGGGGAGAAACCTCTCCCTGCGGGCTGTCGTCGCCCGCATGTTAGAGGATCAGGGCTCGTGGGAGGCCATGGCCAGGTTTTGTGACCTGGTAATGGCGTCCCGTGAGGCTGAGGAGCGTGAAAGGGAGTGGGATCCCTCTTCAGCACCTCAGCGCAGAAGGCGGGATGGCCGGCGCGGGCGTGAAGCTCCCGCTCAGCCCCCGTAG
Protein
MALYGAPVWSPALSARNAALLHRAQRALAVRVIRGYRTISQDVACALAGSLPWDLEAEILAAVYRRKAQALSRGRSPGPSAVGRWKRAARHVAYAKWRERLLKELGQTSATRRRTLEALVPVLEAWSDRRHGVLSFHLAQVLSGHGCFGRYLWKVCRREPHPGCHQCGHPDDDAQHALETCPRWEHSRRDLVAVLGRNLSLRAVVARMLEDQGSWEAMARFCDLVMASREAEEREREWDPSSAPQRRRRDGRRGREAPAQPP
Summary
Uniprot
O01419
A0A2W1BDU5
A0A3S2N998
Q8MY33
Q8MY31
Q8MY27
+ More
Q8MY35 Q5KTM7 A0A0J7KCI1 A0A2W1B333 A0A2W1BSQ5 A0A2W1C3J5 Q868Q4 A0A0J7N1D9 A0A2H1WNZ5 A0A3S2NGT9 A0A0J7MZM4 A0A3L8DDV9 A0A3L8DDS1 A0A0J7N7H9 A0A0J7K2M2 A0A0J7KJB9 A0A3S2LNH8 A0A3L8D2G3 A0A0J7KCQ3 A0A0J7MY07 A0A194R5D3 A0A0J7KS67 A0A0J7KLT9 J9KV04 A0A0J7NEG9 A0A2S2PFB9 A0A0J7KPZ3 A0A0J7KIY5 A0A0K8VFR3 A0A0J7KIY3 A0A0J7K3R6 A0A0J7KIW1 J9KLZ3 J9KPP8 X1X6F1 A0A0J7KFG9 A0A2H8TIT6 A0A142LX37 A0A0K8TSE5 A0A0J7K840 J9LJ80 A0A2S2NS54 A0A0J7K6Z4 A0A0A9Z5Y1 J9KM84 A0A2S2PDK9 A0A0J7KJH4 J9KRD0 V5I8F3 A0A0J7NAB3 A0A0J7JZ13 A0A0J7KMU1 A0A0J7JZF7 X1WZ64 A0A0J7KJG6 A0A2S2PYG9 A0A0J7KYD2 J9K6F0 A0A0J7K3F1 A0A3Q0JIB3 A0A0Q9WRU0 J9LDW8 A0A023EPW0 K7JP42 T1DI88 A0A0J7NMN5 A0A023EQT4 J9KJG8 D7EIB0 A0A2H8TR36 J9KZ24 A0A1W7R6C5 X1WJY4 J9LAC1 A0A023EQZ1 A0A139WNC9
Q8MY35 Q5KTM7 A0A0J7KCI1 A0A2W1B333 A0A2W1BSQ5 A0A2W1C3J5 Q868Q4 A0A0J7N1D9 A0A2H1WNZ5 A0A3S2NGT9 A0A0J7MZM4 A0A3L8DDV9 A0A3L8DDS1 A0A0J7N7H9 A0A0J7K2M2 A0A0J7KJB9 A0A3S2LNH8 A0A3L8D2G3 A0A0J7KCQ3 A0A0J7MY07 A0A194R5D3 A0A0J7KS67 A0A0J7KLT9 J9KV04 A0A0J7NEG9 A0A2S2PFB9 A0A0J7KPZ3 A0A0J7KIY5 A0A0K8VFR3 A0A0J7KIY3 A0A0J7K3R6 A0A0J7KIW1 J9KLZ3 J9KPP8 X1X6F1 A0A0J7KFG9 A0A2H8TIT6 A0A142LX37 A0A0K8TSE5 A0A0J7K840 J9LJ80 A0A2S2NS54 A0A0J7K6Z4 A0A0A9Z5Y1 J9KM84 A0A2S2PDK9 A0A0J7KJH4 J9KRD0 V5I8F3 A0A0J7NAB3 A0A0J7JZ13 A0A0J7KMU1 A0A0J7JZF7 X1WZ64 A0A0J7KJG6 A0A2S2PYG9 A0A0J7KYD2 J9K6F0 A0A0J7K3F1 A0A3Q0JIB3 A0A0Q9WRU0 J9LDW8 A0A023EPW0 K7JP42 T1DI88 A0A0J7NMN5 A0A023EQT4 J9KJG8 D7EIB0 A0A2H8TR36 J9KZ24 A0A1W7R6C5 X1WJY4 J9LAC1 A0A023EQZ1 A0A139WNC9
Pubmed
EMBL
D85594
BAA19776.1
KZ150308
PZC71477.1
RSAL01000481
RVE41577.1
+ More
AB078930 BAC06454.1 AB078931 BAC06456.1 AB078935 BAC06462.1 AB078929 BAC06452.1 AB126050 BAD86650.1 LBMM01009501 KMQ88083.1 KZ150386 PZC71062.1 KZ149950 PZC76665.1 KZ149896 PZC78733.1 AB090825 BAC57926.1 LBMM01012027 KMQ86470.1 ODYU01009945 SOQ54696.1 RSAL01000109 RVE47213.1 LBMM01012957 LBMM01012956 KMQ85900.1 KMQ85901.1 QOIP01000010 RLU18098.1 QOIP01000009 RLU18544.1 LBMM01008861 KMQ88580.1 LBMM01016102 KMQ84557.1 LBMM01006538 KMQ90523.1 RSAL01003504 RVE40273.1 QOIP01000031 RLU14697.1 LBMM01009494 KMQ88097.1 LBMM01014132 KMQ85305.1 KQ460685 KPJ12887.1 LBMM01003679 KMQ93242.1 LBMM01005733 KMQ91252.1 ABLF02004674 ABLF02004675 ABLF02004676 ABLF02011637 LBMM01006110 KMQ90925.1 GGMR01015518 MBY28137.1 LBMM01004505 KMQ92346.1 LBMM01006954 KMQ90166.1 GDHF01014602 JAI37712.1 LBMM01006909 KMQ90209.1 LBMM01015018 KMQ84954.1 LBMM01006725 KMQ90373.1 ABLF02023401 ABLF02029423 ABLF02060278 ABLF02009991 ABLF02009992 ABLF02054952 ABLF02016340 LBMM01008349 KMQ88951.1 GFXV01002242 MBW14047.1 KU543679 AMS38363.1 GDAI01000748 JAI16855.1 LBMM01011877 KMQ86578.1 ABLF02015311 ABLF02015320 ABLF02034009 ABLF02043484 GGMR01006977 MBY19596.1 LBMM01012595 KMQ86127.1 GBHO01002982 JAG40622.1 ABLF02018942 ABLF02041885 GGMR01014918 MBY27537.1 LBMM01006657 KMQ90417.1 ABLF02026677 ABLF02032994 ABLF02059021 ABLF02064578 GALX01004785 JAB63681.1 LBMM01007638 KMQ89555.1 LBMM01019839 KMQ83438.1 LBMM01005216 KMQ91703.1 LBMM01019838 KMQ83439.1 ABLF02011183 ABLF02041884 LBMM01006671 KMQ90407.1 GGMS01001211 MBY70414.1 LBMM01002052 KMQ95336.1 ABLF02012448 ABLF02012450 ABLF02012455 ABLF02012460 LBMM01015025 KMQ84953.1 CH963857 KRF98317.1 ABLF02016435 ABLF02016436 ABLF02064751 GAPW01002021 JAC11577.1 AAZX01012143 AAZX01012194 GALA01001066 JAA93786.1 LBMM01003245 KMQ93755.1 GAPW01002123 JAC11475.1 ABLF02041886 AB593322 BAK38641.1 GFXV01003903 MBW15708.1 ABLF02006132 GEHC01000964 JAV46681.1 ABLF02011238 ABLF02017307 GAPW01002043 JAC11555.1 KQ971310 KYB29500.1
AB078930 BAC06454.1 AB078931 BAC06456.1 AB078935 BAC06462.1 AB078929 BAC06452.1 AB126050 BAD86650.1 LBMM01009501 KMQ88083.1 KZ150386 PZC71062.1 KZ149950 PZC76665.1 KZ149896 PZC78733.1 AB090825 BAC57926.1 LBMM01012027 KMQ86470.1 ODYU01009945 SOQ54696.1 RSAL01000109 RVE47213.1 LBMM01012957 LBMM01012956 KMQ85900.1 KMQ85901.1 QOIP01000010 RLU18098.1 QOIP01000009 RLU18544.1 LBMM01008861 KMQ88580.1 LBMM01016102 KMQ84557.1 LBMM01006538 KMQ90523.1 RSAL01003504 RVE40273.1 QOIP01000031 RLU14697.1 LBMM01009494 KMQ88097.1 LBMM01014132 KMQ85305.1 KQ460685 KPJ12887.1 LBMM01003679 KMQ93242.1 LBMM01005733 KMQ91252.1 ABLF02004674 ABLF02004675 ABLF02004676 ABLF02011637 LBMM01006110 KMQ90925.1 GGMR01015518 MBY28137.1 LBMM01004505 KMQ92346.1 LBMM01006954 KMQ90166.1 GDHF01014602 JAI37712.1 LBMM01006909 KMQ90209.1 LBMM01015018 KMQ84954.1 LBMM01006725 KMQ90373.1 ABLF02023401 ABLF02029423 ABLF02060278 ABLF02009991 ABLF02009992 ABLF02054952 ABLF02016340 LBMM01008349 KMQ88951.1 GFXV01002242 MBW14047.1 KU543679 AMS38363.1 GDAI01000748 JAI16855.1 LBMM01011877 KMQ86578.1 ABLF02015311 ABLF02015320 ABLF02034009 ABLF02043484 GGMR01006977 MBY19596.1 LBMM01012595 KMQ86127.1 GBHO01002982 JAG40622.1 ABLF02018942 ABLF02041885 GGMR01014918 MBY27537.1 LBMM01006657 KMQ90417.1 ABLF02026677 ABLF02032994 ABLF02059021 ABLF02064578 GALX01004785 JAB63681.1 LBMM01007638 KMQ89555.1 LBMM01019839 KMQ83438.1 LBMM01005216 KMQ91703.1 LBMM01019838 KMQ83439.1 ABLF02011183 ABLF02041884 LBMM01006671 KMQ90407.1 GGMS01001211 MBY70414.1 LBMM01002052 KMQ95336.1 ABLF02012448 ABLF02012450 ABLF02012455 ABLF02012460 LBMM01015025 KMQ84953.1 CH963857 KRF98317.1 ABLF02016435 ABLF02016436 ABLF02064751 GAPW01002021 JAC11577.1 AAZX01012143 AAZX01012194 GALA01001066 JAA93786.1 LBMM01003245 KMQ93755.1 GAPW01002123 JAC11475.1 ABLF02041886 AB593322 BAK38641.1 GFXV01003903 MBW15708.1 ABLF02006132 GEHC01000964 JAV46681.1 ABLF02011238 ABLF02017307 GAPW01002043 JAC11555.1 KQ971310 KYB29500.1
Proteomes
Pfam
Interpro
IPR000477
RT_dom
+ More
IPR005135 Endo/exonuclease/phosphatase
IPR036691 Endo/exonu/phosph_ase_sf
IPR012934 Znf_AD
IPR013087 Znf_C2H2_type
IPR036236 Znf_C2H2_sf
IPR002123 Plipid/glycerol_acylTrfase
IPR024079 MetalloPept_cat_dom_sf
IPR001590 Peptidase_M12B
IPR001878 Znf_CCHC
IPR036875 Znf_CCHC_sf
IPR029526 PGBD
IPR037220 BED_dom
IPR012337 RNaseH-like_sf
IPR008906 HATC_C_dom
IPR005135 Endo/exonuclease/phosphatase
IPR036691 Endo/exonu/phosph_ase_sf
IPR012934 Znf_AD
IPR013087 Znf_C2H2_type
IPR036236 Znf_C2H2_sf
IPR002123 Plipid/glycerol_acylTrfase
IPR024079 MetalloPept_cat_dom_sf
IPR001590 Peptidase_M12B
IPR001878 Znf_CCHC
IPR036875 Znf_CCHC_sf
IPR029526 PGBD
IPR037220 BED_dom
IPR012337 RNaseH-like_sf
IPR008906 HATC_C_dom
SUPFAM
Gene 3D
ProteinModelPortal
O01419
A0A2W1BDU5
A0A3S2N998
Q8MY33
Q8MY31
Q8MY27
+ More
Q8MY35 Q5KTM7 A0A0J7KCI1 A0A2W1B333 A0A2W1BSQ5 A0A2W1C3J5 Q868Q4 A0A0J7N1D9 A0A2H1WNZ5 A0A3S2NGT9 A0A0J7MZM4 A0A3L8DDV9 A0A3L8DDS1 A0A0J7N7H9 A0A0J7K2M2 A0A0J7KJB9 A0A3S2LNH8 A0A3L8D2G3 A0A0J7KCQ3 A0A0J7MY07 A0A194R5D3 A0A0J7KS67 A0A0J7KLT9 J9KV04 A0A0J7NEG9 A0A2S2PFB9 A0A0J7KPZ3 A0A0J7KIY5 A0A0K8VFR3 A0A0J7KIY3 A0A0J7K3R6 A0A0J7KIW1 J9KLZ3 J9KPP8 X1X6F1 A0A0J7KFG9 A0A2H8TIT6 A0A142LX37 A0A0K8TSE5 A0A0J7K840 J9LJ80 A0A2S2NS54 A0A0J7K6Z4 A0A0A9Z5Y1 J9KM84 A0A2S2PDK9 A0A0J7KJH4 J9KRD0 V5I8F3 A0A0J7NAB3 A0A0J7JZ13 A0A0J7KMU1 A0A0J7JZF7 X1WZ64 A0A0J7KJG6 A0A2S2PYG9 A0A0J7KYD2 J9K6F0 A0A0J7K3F1 A0A3Q0JIB3 A0A0Q9WRU0 J9LDW8 A0A023EPW0 K7JP42 T1DI88 A0A0J7NMN5 A0A023EQT4 J9KJG8 D7EIB0 A0A2H8TR36 J9KZ24 A0A1W7R6C5 X1WJY4 J9LAC1 A0A023EQZ1 A0A139WNC9
Q8MY35 Q5KTM7 A0A0J7KCI1 A0A2W1B333 A0A2W1BSQ5 A0A2W1C3J5 Q868Q4 A0A0J7N1D9 A0A2H1WNZ5 A0A3S2NGT9 A0A0J7MZM4 A0A3L8DDV9 A0A3L8DDS1 A0A0J7N7H9 A0A0J7K2M2 A0A0J7KJB9 A0A3S2LNH8 A0A3L8D2G3 A0A0J7KCQ3 A0A0J7MY07 A0A194R5D3 A0A0J7KS67 A0A0J7KLT9 J9KV04 A0A0J7NEG9 A0A2S2PFB9 A0A0J7KPZ3 A0A0J7KIY5 A0A0K8VFR3 A0A0J7KIY3 A0A0J7K3R6 A0A0J7KIW1 J9KLZ3 J9KPP8 X1X6F1 A0A0J7KFG9 A0A2H8TIT6 A0A142LX37 A0A0K8TSE5 A0A0J7K840 J9LJ80 A0A2S2NS54 A0A0J7K6Z4 A0A0A9Z5Y1 J9KM84 A0A2S2PDK9 A0A0J7KJH4 J9KRD0 V5I8F3 A0A0J7NAB3 A0A0J7JZ13 A0A0J7KMU1 A0A0J7JZF7 X1WZ64 A0A0J7KJG6 A0A2S2PYG9 A0A0J7KYD2 J9K6F0 A0A0J7K3F1 A0A3Q0JIB3 A0A0Q9WRU0 J9LDW8 A0A023EPW0 K7JP42 T1DI88 A0A0J7NMN5 A0A023EQT4 J9KJG8 D7EIB0 A0A2H8TR36 J9KZ24 A0A1W7R6C5 X1WJY4 J9LAC1 A0A023EQZ1 A0A139WNC9
Ontologies
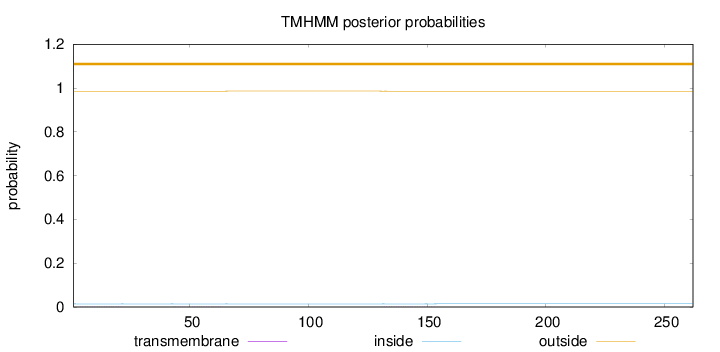
Topology
Length:
262
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.01843
Exp number, first 60 AAs:
0.00703
Total prob of N-in:
0.01411
outside
1 - 262
Population Genetic Test Statistics
Pi
59.783246
Theta
48.723775
Tajima's D
0.557075
CLR
0
CSRT
0.530623468826559
Interpretation
Uncertain
