Gene
KWMTBOMO13684
Annotation
reverse_transcriptase_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 1.258 Mitochondrial Reliability : 1.369
Sequence
CDS
ATGCAAGTTGCGTTCAGTGGTCGAGCTTGGGAGCTGGAGGCGGAGTCGCTCGCTGCCGACTATCGGTGGCGCAGCGAGCTTCGTGCTCGGGGCGTGGCGCGTGTCCCCGAGAGTGAGCTGCGGGCGCGGAAGGCCCATTCTCGGCGGTCCGTGCTCGAGTCGTGGTCGAGGCGATTGGCCAACCCCACGTGGGGGCTACGGACCGTCGAGGCGGTTTACCCGGTCTTTGATGACTGGGTGAATCGTGGCGAGGGACGTCTCACCTTTCGTCTGGTGCAGGTGCTGACCGGGCACGGATGCTTCGGGAAGTACCTGCGCCGGATAGGGGCTGAGCCGACGACGAGGTGTCACCATTGTGGACACGACCTGGACACGGCGGAGCATACGCTCGCTGTCTGCCTCGCTTGGGAGGTGCAGCGCCGTGTCCTGGTCGCAAAGATAGGACCTGACTTGTCGCTGCCTGGCGTCGTGGCGTCGATGCTTGGCAGCGATGAGTCATGGAAGGCTATGCTCGACTTCTGCGAGTGCACCATCTCGCAGAAGGAGGCGGCGGGGCGAGTGAGGGAAAGCTCTCCTCACTACGCAGAAACCCGCCGCCGCCGAGCAGGGGGTCGGGACCGGGGTCGTATCCGTGACCTGGCCCCCTAA
Protein
MQVAFSGRAWELEAESLAADYRWRSELRARGVARVPESELRARKAHSRRSVLESWSRRLANPTWGLRTVEAVYPVFDDWVNRGEGRLTFRLVQVLTGHGCFGKYLRRIGAEPTTRCHHCGHDLDTAEHTLAVCLAWEVQRRVLVAKIGPDLSLPGVVASMLGSDESWKAMLDFCECTISQKEAAGRVRESSPHYAETRRRRAGGRDRGRIRDLAP
Summary
Cofactor
FAD
Similarity
Belongs to the GMC oxidoreductase family.
Uniprot
Q868Q4
Q5KTM7
A0A0J7KCI1
A0A2W1B333
A0A0J7K2M2
A0A2W1BSQ5
+ More
A0A2W1BDU5 A0A0J7N1D9 A0A2W1C3J5 Q8MY27 A0A0J7KJB9 A0A3L8DDS1 Q8MY31 Q8MY33 A0A0J7MZM4 Q8MY35 A0A0J7KCQ3 A0A3S2NGT9 A0A0J7KIY3 A0A3S2LNH8 A0A2H1WNZ5 A0A0J7KS67 A0A2H1VYR8 A0A3S2N998 E2BVR6 A0A0J7N7H9 A0A0L7QYH7 O01419 A0A0J7KIQ5 A0A194R5D3 A0A3L8DDV9 A0A3L8D2G3 A0A3L8DXT9 A0A0J7KQU9
A0A2W1BDU5 A0A0J7N1D9 A0A2W1C3J5 Q8MY27 A0A0J7KJB9 A0A3L8DDS1 Q8MY31 Q8MY33 A0A0J7MZM4 Q8MY35 A0A0J7KCQ3 A0A3S2NGT9 A0A0J7KIY3 A0A3S2LNH8 A0A2H1WNZ5 A0A0J7KS67 A0A2H1VYR8 A0A3S2N998 E2BVR6 A0A0J7N7H9 A0A0L7QYH7 O01419 A0A0J7KIQ5 A0A194R5D3 A0A3L8DDV9 A0A3L8D2G3 A0A3L8DXT9 A0A0J7KQU9
EMBL
AB090825
BAC57926.1
AB126050
BAD86650.1
LBMM01009501
KMQ88083.1
+ More
KZ150386 PZC71062.1 LBMM01016102 KMQ84557.1 KZ149950 PZC76665.1 KZ150308 PZC71477.1 LBMM01012027 KMQ86470.1 KZ149896 PZC78733.1 AB078935 BAC06462.1 LBMM01006538 KMQ90523.1 QOIP01000009 RLU18544.1 AB078931 BAC06456.1 AB078930 BAC06454.1 LBMM01012957 LBMM01012956 KMQ85900.1 KMQ85901.1 AB078929 BAC06452.1 LBMM01009494 KMQ88097.1 RSAL01000109 RVE47213.1 LBMM01006909 KMQ90209.1 RSAL01003504 RVE40273.1 ODYU01009945 SOQ54696.1 LBMM01003679 KMQ93242.1 ODYU01005282 SOQ45985.1 RSAL01000481 RVE41577.1 GL450953 EFN80213.1 LBMM01008861 KMQ88580.1 KQ414692 KOC63606.1 D85594 BAA19776.1 LBMM01006996 KMQ90129.1 KQ460685 KPJ12887.1 QOIP01000010 RLU18098.1 QOIP01000031 RLU14697.1 QOIP01000003 RLU25053.1 LBMM01004152 KMQ92762.1
KZ150386 PZC71062.1 LBMM01016102 KMQ84557.1 KZ149950 PZC76665.1 KZ150308 PZC71477.1 LBMM01012027 KMQ86470.1 KZ149896 PZC78733.1 AB078935 BAC06462.1 LBMM01006538 KMQ90523.1 QOIP01000009 RLU18544.1 AB078931 BAC06456.1 AB078930 BAC06454.1 LBMM01012957 LBMM01012956 KMQ85900.1 KMQ85901.1 AB078929 BAC06452.1 LBMM01009494 KMQ88097.1 RSAL01000109 RVE47213.1 LBMM01006909 KMQ90209.1 RSAL01003504 RVE40273.1 ODYU01009945 SOQ54696.1 LBMM01003679 KMQ93242.1 ODYU01005282 SOQ45985.1 RSAL01000481 RVE41577.1 GL450953 EFN80213.1 LBMM01008861 KMQ88580.1 KQ414692 KOC63606.1 D85594 BAA19776.1 LBMM01006996 KMQ90129.1 KQ460685 KPJ12887.1 QOIP01000010 RLU18098.1 QOIP01000031 RLU14697.1 QOIP01000003 RLU25053.1 LBMM01004152 KMQ92762.1
Proteomes
Pfam
Interpro
IPR000477
RT_dom
+ More
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR002123 Plipid/glycerol_acylTrfase
IPR024079 MetalloPept_cat_dom_sf
IPR001590 Peptidase_M12B
IPR036188 FAD/NAD-bd_sf
IPR012132 GMC_OxRdtase
IPR000172 GMC_OxRdtase_N
IPR012934 Znf_AD
IPR013087 Znf_C2H2_type
IPR036236 Znf_C2H2_sf
IPR011990 TPR-like_helical_dom_sf
IPR036691 Endo/exonu/phosph_ase_sf
IPR005135 Endo/exonuclease/phosphatase
IPR002123 Plipid/glycerol_acylTrfase
IPR024079 MetalloPept_cat_dom_sf
IPR001590 Peptidase_M12B
IPR036188 FAD/NAD-bd_sf
IPR012132 GMC_OxRdtase
IPR000172 GMC_OxRdtase_N
IPR012934 Znf_AD
IPR013087 Znf_C2H2_type
IPR036236 Znf_C2H2_sf
IPR011990 TPR-like_helical_dom_sf
Gene 3D
ProteinModelPortal
Q868Q4
Q5KTM7
A0A0J7KCI1
A0A2W1B333
A0A0J7K2M2
A0A2W1BSQ5
+ More
A0A2W1BDU5 A0A0J7N1D9 A0A2W1C3J5 Q8MY27 A0A0J7KJB9 A0A3L8DDS1 Q8MY31 Q8MY33 A0A0J7MZM4 Q8MY35 A0A0J7KCQ3 A0A3S2NGT9 A0A0J7KIY3 A0A3S2LNH8 A0A2H1WNZ5 A0A0J7KS67 A0A2H1VYR8 A0A3S2N998 E2BVR6 A0A0J7N7H9 A0A0L7QYH7 O01419 A0A0J7KIQ5 A0A194R5D3 A0A3L8DDV9 A0A3L8D2G3 A0A3L8DXT9 A0A0J7KQU9
A0A2W1BDU5 A0A0J7N1D9 A0A2W1C3J5 Q8MY27 A0A0J7KJB9 A0A3L8DDS1 Q8MY31 Q8MY33 A0A0J7MZM4 Q8MY35 A0A0J7KCQ3 A0A3S2NGT9 A0A0J7KIY3 A0A3S2LNH8 A0A2H1WNZ5 A0A0J7KS67 A0A2H1VYR8 A0A3S2N998 E2BVR6 A0A0J7N7H9 A0A0L7QYH7 O01419 A0A0J7KIQ5 A0A194R5D3 A0A3L8DDV9 A0A3L8D2G3 A0A3L8DXT9 A0A0J7KQU9
Ontologies
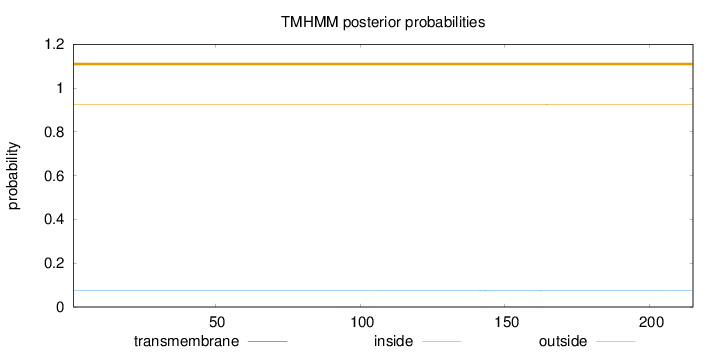
Topology
Length:
215
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.01126
Exp number, first 60 AAs:
0
Total prob of N-in:
0.07626
outside
1 - 215
Population Genetic Test Statistics
Pi
155.35833
Theta
77.865903
Tajima's D
2.079318
CLR
64.411469
CSRT
0.893755312234388
Interpretation
Uncertain
