Gene
KWMTBOMO13491
Annotation
PREDICTED:_uncharacterized_protein_LOC107046166_[Diachasma_alloeum]
Location in the cell
Mitochondrial Reliability : 1.54 Nuclear Reliability : 1.936
Sequence
CDS
ATGAATGGTAGCTCCGTAGTTCTGCTGGCTACTGCTAACGTTGAAGTGCGCGACTCATCTGGTCACTTTCATACTCTTCGCGCACTCATCGACCCTGGTAGTCAATCACATTTCATTTCTAATCAAGCTATGGAACGATTGGGCTTAAAGCCTAAGCAAACTTCGCGAAATGTTTGTGGCTTGGGGCAATCATCGACGTCAGTGACTGGAATGGTGGAGCTTCAACTCGGAGTTCATCAAAAGAATTTATTTAATTTAACGGCTCTCATCTTACCATCTATTTGTGCACAAATGCCCACTGCTCATCTTGATCTTTCTTCATGCGAACACCTACGTAATTTGCGACTCGCTGATCCGAATTATAATAAGCCTGGACCAATAGATCTGCTTATCGGTGCTGAAATGTTTTCAACGCTGTTACTGCCCGGAAATATTTCAGGTAGTCCTCATGAACCCTCAGCCTTGAACACCATCTTCGGATGGATCCTGATGGGAAGTGTCAAATGCAGGGATGAGACAAAACACTCAAATTCTTTTTTTCTAAGAGAACACGATACTCCTCTCCATGAGGAATTGAAGAGACTGTGGGAGTTGGAGGTCATACCTAAATCATCAGTTCTAACGCCGGACGAGCAGCAATGTGAGAAACTTTTCACTGAAACCCACGCTCGAGGTACATCTGGAAGGTACATCGTTTCTTTGCCGTTTAGGCTGAGGGATGGGGAGTTACAGTTTCCGGGTTCGCGTGAAAATACATTGCGAAGATTCCATTCATTGGAACGAAGACTTCTTTCAAGTCCAAACCTTCATCAGCAGTATTCGGACTTCATGCTGGACTACTTACATAATGGTCATAGGAGTATCCTTTTATATGCTAAGTCTCGCGTGGCCCCAATTAAGAGAGTTTCATTGCCGAGACTAGAGCTCTGCGCTGCGGTTGTGCTCGCTGACCTGCTCAAATTCGTAAAAGAAGTGTATGTACCGTTACTTCCGCTTTCTGATACCTTTGCTTGGTCAGATTCAACGGTAACTCTGTCATGGATTCGTGGCCATTCCTCGCGTTGGAAAACATTTGTTGCTAATAGGGTTAGTCATATCCAAGAAACTGTCCCCGAAACTCACTGGAATCACGTGCGCTCTAATGAAAATCCTGCGGACATAGCTTCCCGCGGTATGTGTCCGAGCGACCTACTCCAGAGTGAGTTATGGTGGGCGGGACCGAGCTGGCTTCAGCAACCTTCATCTTCATGGCCTTCAAATTCCGGTCTCGAGGTGAACGAGACTGATATGCTTGCCGAGGAACGCTGTTTTTCGTTGATCCTGAATGATGAACCTAACTTTATCGATTCTTTGCTCGAAAGGTTTTCATCAATTCGTAAGGTTCAACGTATTATTGGTTATTGTCTTCGTTTTATCAGAAATGCTAACAAAAAACAGACACGTGTTCCGGGTACCCTTTTAGGTCGTGCAGAACTTCATGGGGCCCTGATGACGATTACCAAGCGAATTCAAATTCTCAACTTTCAGGATGAAATTCGTAAGCTCGAGAATGGTATACCATTACCCAAACCGTGGAGAAAACTCAATCCATTCCTGGACGACCAAGGAGTTGTCAGGGTGGGCGGCCGTTTGTCTCGTTCTGGTCTTGAGTTTAGTCATAAGCATCCTGCTCTGTTGCCGCGTCATTCTAGGTTCACATACATGATAATCGAAGCAATTCACAAAGAGAATTGTCATCCTGGGTTGGGAACGACACAATATCTTTTGATGCAGCTGTTTTGGAGAGCAAAGTGGAACAAAAGGGGCGAAACTTTGAATGATGGTTCTGTTGTACTGATTAAGAGTGAAAGTACAATGCCGTTGCACTGGCCTTTAGCTCGTGTTTCCAAGCTTCACGCCAGCCCTGATGGCGTGGTTCGTATTGCCACTGTCCGTACAGCCAATGGCAAATTCTACACTAGGCCATTAACGAAGTTAGCTCCCCTACCTATTTATTCTGATTAA
Protein
MNGSSVVLLATANVEVRDSSGHFHTLRALIDPGSQSHFISNQAMERLGLKPKQTSRNVCGLGQSSTSVTGMVELQLGVHQKNLFNLTALILPSICAQMPTAHLDLSSCEHLRNLRLADPNYNKPGPIDLLIGAEMFSTLLLPGNISGSPHEPSALNTIFGWILMGSVKCRDETKHSNSFFLREHDTPLHEELKRLWELEVIPKSSVLTPDEQQCEKLFTETHARGTSGRYIVSLPFRLRDGELQFPGSRENTLRRFHSLERRLLSSPNLHQQYSDFMLDYLHNGHRSILLYAKSRVAPIKRVSLPRLELCAAVVLADLLKFVKEVYVPLLPLSDTFAWSDSTVTLSWIRGHSSRWKTFVANRVSHIQETVPETHWNHVRSNENPADIASRGMCPSDLLQSELWWAGPSWLQQPSSSWPSNSGLEVNETDMLAEERCFSLILNDEPNFIDSLLERFSSIRKVQRIIGYCLRFIRNANKKQTRVPGTLLGRAELHGALMTITKRIQILNFQDEIRKLENGIPLPKPWRKLNPFLDDQGVVRVGGRLSRSGLEFSHKHPALLPRHSRFTYMIIEAIHKENCHPGLGTTQYLLMQLFWRAKWNKRGETLNDGSVVLIKSESTMPLHWPLARVSKLHASPDGVVRIATVRTANGKFYTRPLTKLAPLPIYSD
Summary
Uniprot
Proteomes
Pfam
Interpro
SUPFAM
SSF53098
SSF53098
Gene 3D
ProteinModelPortal
Ontologies
KEGG
Topology
Length:
667
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.01538
Exp number, first 60 AAs:
0.00013
Total prob of N-in:
0.00079
outside
1 - 667
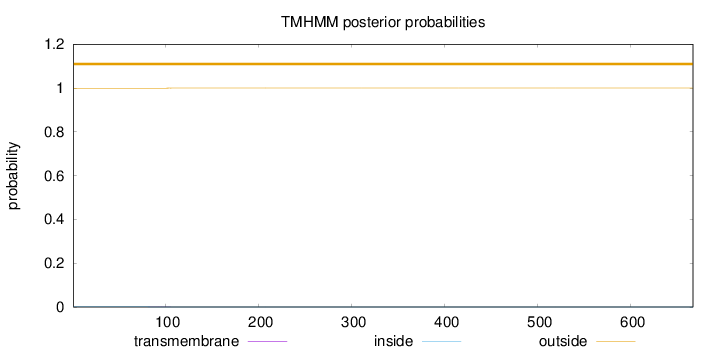
Population Genetic Test Statistics
Pi
3.440445
Theta
17.941971
Tajima's D
-2.090808
CLR
2.145311
CSRT
0.00849957502124894
Interpretation
Uncertain
