Gene
KWMTBOMO13449 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA010889
Annotation
reverse_transcriptase_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 3.467
Sequence
CDS
ATGTCGTTCAGTTGGATCGTAGTGTTAGCGTTCGTCAACATCATCGTCCTCTGTACAGCTACTTGTCCAGAAAACGAAGAACGCACCTGCCTTCAAGGTTTATGCAGACCTCAGAAGTGCATCGAAAAAAATGATATCATCTTCTGCCAATTAGTTGATGAGGAGAAGTGCGAATATGGATGTGCCTGCAAAATAGGATATCTGAGAGACGAAAATGGAACTTGTATACCGCAAGACAAGTGTCCAACTGTGCCTTGTCCAGTGAATGAATACTTCACGAACTGCGCTAAGGGCATGTGTCGTCAGGAGAACTGCACAGAGCTGGGCAAGTTGTCTGAATGTAAAACGCAATCGACAGAGCTGTGCGAGCCGGGCTGCGTTTGTGAAGGGGGCTTCTTGAGATCAAAAAACGGAACGTGCGTATCTATCGATGAATGCCATAGGGAACTATGTCCCGTGAACGAAGTGTACTCGAGCTGTCGACAGCCCAATTGTAATTCTGATAAATGCGAGTACAAATACAGGTCACAGTCGTGTCCCTCGGACGAACCCTGCGAAGTCGGTTGTGTTTGTAAAAGAGGATTTCGTAGGGCTGACAATGGCACCTGTGTCGATGAAAGAGATTGTGAATCCCAGCTTTGTTCAGTAAATGAACAGTATTTAAGTTGCATCCAAGCTGTTTGTCGGGTCGAGAAGTGTTCAGACCTGGGGGGATCCCTCAGTTGCAAGGGGGTGTCGGAAAGGGAATGCGTCGGTGGCTGCGTCTGTAAGGACAACTACTTCCGAGCTAAAAACGACACTTGTATTAAACTCAGTGATTGTGACGCTGACCTTTGCTCTGAAAATGAAATACACGTGAACTGTGTCCTAGCACAGTGTGGCCCAATGACGTGCTCAGAGAAGGACGCGGTAAATGTGTCGCCCGAGAAAATTGCCCAAATTAGGAATGTTCAGGCGAAAATGAAGAGTTTACCAACTGCACGAACCCATGTCCGCCGCGTACTTGCAATTCACTCGTCGCTCGTTTTGATTGTAACCCCCGCTCGCGATTCTGCCTGCTGTAGCACTATACAGCCCGAGCGGGTTCCTTTCCTTTTCCCCGCCGATTGCCGTTTTCACGGCATAGGCATTCCCCCCCCTGCTCCTTGCTGTCCTGCAGCACAGGGGGGTTACAAATCCTATCCGGGGCCTCGCCTTTCGGCGAGTTCAACAAAAGTCCCGGGGCCGCCCCCCCGGGCAAGAAGACCGAAGATGAGCGAAGATTTTGTGTGTGACAGTGAAAGTGCTAGGGACATTGTCGGGTTCCTCCGGGATAGGTACCCGTCAATTAGGGCCGAGTATCTCAGCTATAGGGCCGCTCGTGGCAATCCCTCCCCCCCCGCGTCCGTCGCCGCGTTTCTCAACCGTGCCTCGGAGGCACGCCGCGCGCCCCCCGCCGCCGCGTCAAGTTCGAGCGCACGTTCCCACGATGCGGCTCACATAGCGCCTAACGCTACCCCCACCCCCGCGCCCGCGTCGTTTACGTCATCGCGCGCTTCGTCCGTCGCGACCTCGATCGTTTCGAGTTCAGGCTCCGATACCGAATCGGAGATGGATTTCGAATCCGCGTCAAGCCCTCAGCCTGGAACTTCGGATGGGTTCCAAACCGTAACGCGCGGCAAAAAGCGTACTCGCGCCGTGGAGTCCCGGAGCTCCACGACGAAACAGACTAAATCCGCGACCGCCTCTCGACCAAAAGTAGTAGTTACACCGGAGTCGGACTCCGCGCGCCGCGTAACCCCACCGCCGCGTCCCAAGCCCGCGCCCGCTCCCCAAGTAGCTGCCCCGCCCCCGCTGATACTTCAAGAGAAGACAGCGTGGAATCGCGTGTCCCAGGCCCTTCAGGCAAATAAAATTAACTACACCCATGCGCGTAACGTCGCGCATGGGATTCAGATCAAGGTCGCAACGCCGGGCGACCATAGGGCCCTCTCAGCATACCTCCGAAAGGAGAACATAGGTTTCCACACCTATGCTCTTCAGGAGGACCGCGAGCTCCGCGTAGTAATACGCGGAGTCCCTAAGGAGCTAGAAATAGACTACATTAAGGAGGATCTGACCGCTCAGGCCCTCCCGATAGTTAGTGTGCACCGGATGCACAGCGGGCGGGGCAAACAGCCCTATAATATGATACTCGTGGCGTTAGAACCCACCCCGGAGGCCAAAAAGAAGATCGCATGTCTCACGACAATTTGCGGCCTATCCGGGATCTCGATCGAAGCCCCCCATAAGCGTGGCACTCCCGGACAGTGCTACAGGTGCCAACTCTATGGCCACTCAGCGCGTAATTGCCACGCGCGCCCCCGCTGCGTGAAATGCCTCGGCGATCACGCTACGTTAGATTGCGTTAGAGATAGGGAAACCGCGACCGAACCTCCCAGCTGTGTCCTATGCCTTAAGCAAGGGCACCCCGCGAACTACCGTGGATGTCCCAGGGCTCCGCGGAAACGTCCAACCCACCCAGCTCCGCGAACCGTCGGCCCAAAGACTTCGGCACCTTCAGTGCCGAAACCCGCCTTCGTGCCGGCTGCAGTTCCCACGGTCTCGGCGTGGAAAAAGCCGCTGCCGTATACGAAGGCGGGAACGGAAACCGCACCCCCGCCCCCGCTCGCGACACGCCCCGCGCCTCAGCCCCTGCTCGCACCTCGCCCCGCGCCCGCGTTCCGTCCCCCGCAACCCTCTGCCGCCAACGACTTCGCGCTCGTGCGCGACTTCGTCACCGCGGTGAACTTCGACCGTCTGCGATCGTTCGCTGACGCTATCCGTAGGTCGACAACCCCCGAACAGCGGCTCGCGGCCGCGTTCGATCACATGGACGTTTACGAGTCCGTGTCGCCATTCTATAATGCTAACGGACTCGCGCGGCAACGCGATCAAATTTTCGAATTCCTCCGCGATAATCTTGTAGATATTCTATTAGTGCAGGAAACCTGTCTGAAGCCCTCGCGTCGCGACCCGAAAGTAGCGAATTACGTCATGGTTAGGAATGACAGACTCACCGCCTCCAAAGGCGGGACTGCCATTTACTATAGGCGGGCCCTGCACGTTGTCCCTCTCGATACTCCCTCGCTCTCACATATCGAGGCGTCAGTGTGCCGTATCTCGCTGACGGGACACCAGCCGATCGTCATCGCATCCGTTTATCTCCCCCCGGACAAGCCCCTTCTGAGCAGTGACATCGAGTCACTGTTCGGCATGGGAGACTCCGTCATCCTGGCAGGCGATTTAAATTGCCACCACACTAGGTGGAACTGCCATCGTACTAACGTTAACGGTAGGCGTCTCGACGCGTTTATAGACGACCTCACCTTTGAAATAGTCGGTCCCCCAACTCCAACATGTTATCCGTATAACATCGCGCTCCGTCCGAGCACTATAGACCTGGCATTGCTTAGGAACGTAACTCTGCGCTTGCGTTCCATCGAAGCAATGTCAGAGCTCGACTCAGACCACCGACCTGTCGTTATGCAGCTCGGTCGCCCCCACAACCCAGTCACTGTTACGAGGACCATGGTGGATTGGAATAAGCTGGGCACATCTTCTAAAGAAGTCGATGTGGAGGACAGCTTCCACCGCATCAGACTGTCCCCCGATCTTAGGAATCTCTTAAGAGTTAGGAACGCGGCAATCCGGGCCTACGATCGTCTTCCCACGCATCCAAACCGGATTCGGATGCGTCGTCTACAACGCGAAGTCCACTCCCGCTTAAGCGACGCGCGTAACGATAATTGGCATAGTTATTTAGAACAACTCGCGCCCTCCCACCAAGCATACTGGCGACTAGCTAGGACTCTCAAATCCGAAACTACCGCTACTATGCCTCCCCTCGTACGCCCTTCAGGCCAACCACCGGCATTCGATGACGATGACAAGGCTGAGCTGCTGGCCGATGCACTGCAAGAGCAGTGCACCACCAGCACTCAACACGCGGACCCCGAACACACCGAGTTAGTCGACAGGGAGGTCGAGCGCAGAGCTTCCCTGCCGCCCTCGGACGCGTTACCCCCCATTACCACTGACGAAGTTAGAGACGCGATCCACAACCTACAACCTAGGAAGGCACCCGGCTCCGACGGCATCCACAACCGTGCGCTTAAACTCTTGCCAGTCCAACTGATAGCAATGTTGGCTACAATTTTAAATGCCGCTATGACGCACTGCATCTTTCCCGCGGTGTGGAAAGAAGCGGACGTTATCGGTATACATAAGCCGGGCAAACCGACAAACGAAACTTCTAGTTACCGTCCGATTAGTCTCCTCTCGACGATAGGAAAAATTTACGAACGGCTCCTTAGGAAACGCCTCTGGGATTTTGTTACCGCGAACAAAATTCTCATAGACGAACAGTTCGGATTCCGCTCAAAACACTCGTGCGTACAACAAGTGCACCGCCTCACGGAGCACATTCTGATAGGACTAAATAGGCGTAAACAAATCCCGACCGGCGCCCTCTTCTTCGACATCGCGAAGGCGTTCGACAAAGTCTGGCACAACGGTTTAATTTATAAACTATACAACATGGGAGTGCCAGACAGACTCGTGCTCATCATACGAGACTTCTTGTCGAACCGTTCGTTTCGATATCGAGTAGAGGGAACTCGTTCTCGTCCCCGCCAACTGACCGCCGGAGTCCCGCAAGGCTCCGCGCTCTCCCCGTTATTATTTAGTTTGTATATCAATGATATACCCCGGTCTCCGGAGACCCATCTAGCGCTCTTCGCCGATGACACGGCTATCTACTACTCGTGTAGGAAGATGTCGCTGCTTCATCGGCGACTCCAGATCGCAGTAGCCACCCTGGGACAGTGGTTCCGGAAGTGGCGAATAGACATCAACCCCACGAAAAGCGCAGCGGTGCTCTTCAAAAGGGGTCGCCCTCCGAACATCACTTCGAGCATCCCACTCCGTAGTAGGCGCGCAAACACCTCCGCCGTTAGTCCCATCACTCTCTTTGGCCAGCCCATTCCGTGGGTCTCGAAGGTCAAATATCTAGGCGTCACCCTCGACAGAGGGATGACATTCCGTCCCCATATAAAAACGGTACGCGACCGCGCCGCGTTTATTCTAGGACGACTCTATCCAATGCTTTGTAGTCGAAGCAAACTGTCCCTCCGCAATAAGGTAACTCTCTACAAAACTTGTATACGCCCCGTCATGACGTATGCAAGCGTAGTGTTCGCTCACGCAGCCCGCACCAACTTGAAATCCCTTCAGGTTATTCAATCACGATTCTGCAGGATAGCCGTCGGAGCGCCTTGGTTCCTTAGGAATGTGGATCTCCACGACGACCTGGAGCTCGACTCTTTTAGTAAGTATCTACAGTCGGCATCGCTGCGCCATTTTGAGAAGGCGGCACGACATGAGAACCCTCTCATCGTAGCCGCTGGAAATTACATACCCGACCCAGTAGACCGAATGGTAAACCGTCGACGTCGCCCAAAGCACGTCATTACGGATCCTCCTGATCCATTAACGGTGCTTTTAGGCACCACAAGCACCGGTCACCGTCCTCGTCGAACCCGTCGCTTGCGACGAAGGGCTCGACGAGCGAACTAA
Protein
MSFSWIVVLAFVNIIVLCTATCPENEERTCLQGLCRPQKCIEKNDIIFCQLVDEEKCEYGCACKIGYLRDENGTCIPQDKCPTVPCPVNEYFTNCAKGMCRQENCTELGKLSECKTQSTELCEPGCVCEGGFLRSKNGTCVSIDECHRELCPVNEVYSSCRQPNCNSDKCEYKYRSQSCPSDEPCEVGCVCKRGFRRADNGTCVDERDCESQLCSVNEQYLSCIQAVCRVEKCSDLGGSLSCKGVSERECVGGCVCKDNYFRAKNDTCIKLSDCDADLCSENEIHVNCVLAQCGPMTCSEKDAVNVSPEKIAQIRNVQAKMKSLPTARTHVRRVLAIHSSLVLIVTPARDSACCSTIQPERVPFLFPADCRFHGIGIPPPAPCCPAAQGGYKSYPGPRLSASSTKVPGPPPRARRPKMSEDFVCDSESARDIVGFLRDRYPSIRAEYLSYRAARGNPSPPASVAAFLNRASEARRAPPAAASSSSARSHDAAHIAPNATPTPAPASFTSSRASSVATSIVSSSGSDTESEMDFESASSPQPGTSDGFQTVTRGKKRTRAVESRSSTTKQTKSATASRPKVVVTPESDSARRVTPPPRPKPAPAPQVAAPPPLILQEKTAWNRVSQALQANKINYTHARNVAHGIQIKVATPGDHRALSAYLRKENIGFHTYALQEDRELRVVIRGVPKELEIDYIKEDLTAQALPIVSVHRMHSGRGKQPYNMILVALEPTPEAKKKIACLTTICGLSGISIEAPHKRGTPGQCYRCQLYGHSARNCHARPRCVKCLGDHATLDCVRDRETATEPPSCVLCLKQGHPANYRGCPRAPRKRPTHPAPRTVGPKTSAPSVPKPAFVPAAVPTVSAWKKPLPYTKAGTETAPPPPLATRPAPQPLLAPRPAPAFRPPQPSAANDFALVRDFVTAVNFDRLRSFADAIRRSTTPEQRLAAAFDHMDVYESVSPFYNANGLARQRDQIFEFLRDNLVDILLVQETCLKPSRRDPKVANYVMVRNDRLTASKGGTAIYYRRALHVVPLDTPSLSHIEASVCRISLTGHQPIVIASVYLPPDKPLLSSDIESLFGMGDSVILAGDLNCHHTRWNCHRTNVNGRRLDAFIDDLTFEIVGPPTPTCYPYNIALRPSTIDLALLRNVTLRLRSIEAMSELDSDHRPVVMQLGRPHNPVTVTRTMVDWNKLGTSSKEVDVEDSFHRIRLSPDLRNLLRVRNAAIRAYDRLPTHPNRIRMRRLQREVHSRLSDARNDNWHSYLEQLAPSHQAYWRLARTLKSETTATMPPLVRPSGQPPAFDDDDKAELLADALQEQCTTSTQHADPEHTELVDREVERRASLPPSDALPPITTDEVRDAIHNLQPRKAPGSDGIHNRALKLLPVQLIAMLATILNAAMTHCIFPAVWKEADVIGIHKPGKPTNETSSYRPISLLSTIGKIYERLLRKRLWDFVTANKILIDEQFGFRSKHSCVQQVHRLTEHILIGLNRRKQIPTGALFFDIAKAFDKVWHNGLIYKLYNMGVPDRLVLIIRDFLSNRSFRYRVEGTRSRPRQLTAGVPQGSALSPLLFSLYINDIPRSPETHLALFADDTAIYYSCRKMSLLHRRLQIAVATLGQWFRKWRIDINPTKSAAVLFKRGRPPNITSSIPLRSRRANTSAVSPITLFGQPIPWVSKVKYLGVTLDRGMTFRPHIKTVRDRAAFILGRLYPMLCSRSKLSLRNKVTLYKTCIRPVMTYASVVFAHAARTNLKSLQVIQSRFCRIAVGAPWFLRNVDLHDDLELDSFSKYLQSASLRHFEKAARHENPLIVAAGNYIPDPVDRMVNRRRRPKHVITDPPDPLTVLLGTTSTGHRPRRTRRLRRRARRAN
Summary
Uniprot
ProteinModelPortal
PDB
6AR3
E-value=6.1325e-05,
Score=117
Ontologies
Topology
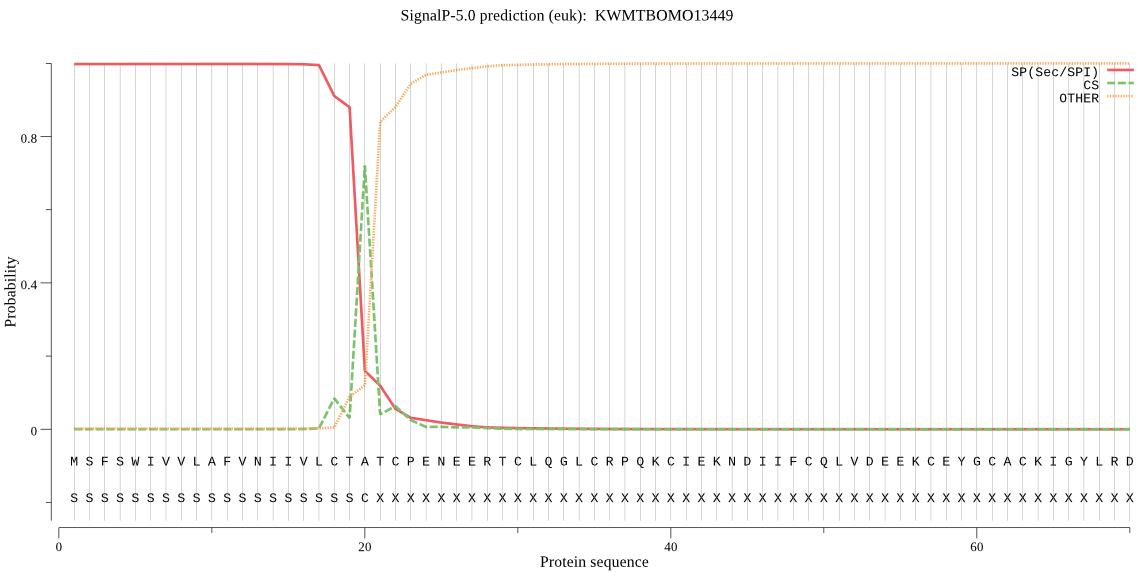
SignalP
Position: 1 - 20,
Likelihood: 0.998351
Length:
1864
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
5.37710999999997
Exp number, first 60 AAs:
4.93359
Total prob of N-in:
0.24742
outside
1 - 1864

Population Genetic Test Statistics
Pi
16.043634
Theta
14.67516
Tajima's D
0
CLR
1.703747
CSRT
0.380530973451327
Interpretation
Uncertain
