Gene
KWMTBOMO13256 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA004726
Annotation
PREDICTED:_antichymotrypsin-2-like_[Bombyx_mori]
Full name
Antitrypsin
+ More
Antichymotrypsin-2
Alaserpin
Antichymotrypsin-2
Alaserpin
Alternative Name
Antichymotrypsin II
Serpin-1
Serpin-1
Location in the cell
Extracellular Reliability : 1.794
Sequence
CDS
ATGTATCCCGAAGACAGAGTGCCAATTGATCCTAAATACTGTAAAGGAGGCTGTCTCTGTGAAGACGGCTATGTAAAAGCTAAGAATGGTACTTGCATACCAGAGAAAGAATGTCCATCATGTGGCGGTGATCCGAATGCCGAAGCTGGTTGTGGGATAAATTGTAACAGGCGTTGTTCAGATGTTGGCAAGGACCCTGGAGTGTGCGAGGACGATATATGTGCTGTAAACGGTTGTGATTGTAAACAAGGATATTACTATGATTACGTCTCATCGAAATGTGTTCTACCAGATCGATGTCCTACTACCAATTCATCGGGAAATGTCGACCAATCCTTGGAAAAACTCCGCAGAGGAAACATGGCATTAGTTGGCCAGTTTCTTTGGGAATTATACAAAATGAATCCAGGACAAAGTTTCGTTACATCACCCGTATCTGTACTCATTCCTTACGGTCAACTAGCGCTATACGCCGTTGGAGAAACTTTAGATCAACTATTAAAGTTTATAAACCTCGACAATAAGGATCAGATCAAGAAAGCTTTCCCCGCGTTGCTTAAGAATTATCGGTCTCAAACCTCAGTTCTTTTGACGGTGGCAGTTAAATGCTACGGCAACATAAATTATCCTTTTACGGACCAGTTTAAGAACGACGCTGTAACTTATTTTGACTCAGAAGGAGAAAATATTGATTTTAAATATGCGAAACAAGCTGCAGACACAATAAACGGATGGGTCGCAAACAAAACCAATAACTTAATTCAAAATCTTGTCAGCGAAGATTTGTTCAGCGATGACACTAGATTGGTTCTAGTTGACGCCATTTATTTGCATGGAATTTGGCAAAATCAGTTCAATCCTAATAACACTCACGACAGAGACTTCTACATCACACCAAACAAAACTAAGTCCATAAAGACTATGTATCAAGAAAACTTCTTTAATTATGGAGAAAACGAGATTCTTGATGCACAGATCTTAGAATTGTTCTACAAAGGCGGCAATTCTAGTTTAGTAATCTTGTTACCAAAGAAAAAAGATGGCCTACCCCAGATGTCACATAAATTACGAAATTCAGATGAATTTTTTAATAGCATTAAATCACTTAGGCGAGAGAAAGTCAAATGTTTCCTACCTAAGATGGATATAAAAACGAACATCGATATGGCAGACCTTTCCCAGAAGATGAACGTAACCAAAATGTTCGATCCACAATCAAATGATTTCGAAGGAATTTTATTAAATTCGGACCCCGTATATGTTTCTGCTGCAATACAGAAAGCTCAAATCATAGTTGATGAAACTGGAACCGAGGCAGCGGTCGCAACTGCCGTCGTACTCACGAAAAGAAGTAGAACAGATATGTATTCTTTTGAAGCAGATCATCCGTTTGTATACGCAATCAAAGTTCAAGATGACATTATATTTAGTGGGGTTTTCGTCGGATAA
Protein
MYPEDRVPIDPKYCKGGCLCEDGYVKAKNGTCIPEKECPSCGGDPNAEAGCGINCNRRCSDVGKDPGVCEDDICAVNGCDCKQGYYYDYVSSKCVLPDRCPTTNSSGNVDQSLEKLRRGNMALVGQFLWELYKMNPGQSFVTSPVSVLIPYGQLALYAVGETLDQLLKFINLDNKDQIKKAFPALLKNYRSQTSVLLTVAVKCYGNINYPFTDQFKNDAVTYFDSEGENIDFKYAKQAADTINGWVANKTNNLIQNLVSEDLFSDDTRLVLVDAIYLHGIWQNQFNPNNTHDRDFYITPNKTKSIKTMYQENFFNYGENEILDAQILELFYKGGNSSLVILLPKKKDGLPQMSHKLRNSDEFFNSIKSLRREKVKCFLPKMDIKTNIDMADLSQKMNVTKMFDPQSNDFEGILLNSDPVYVSAAIQKAQIIVDETGTEAAVATAVVLTKRSRTDMYSFEADHPFVYAIKVQDDIIFSGVFVG
Summary
Miscellaneous
The N-terminal section (1-336) is identical to the N-terminal section of the silk moth antitrypsin I (17-352).
Similarity
Belongs to the serpin family.
Keywords
Complete proteome
Direct protein sequencing
Protease inhibitor
Reference proteome
Secreted
Serine protease inhibitor
Signal
3D-structure
Alternative splicing
Glycoprotein
Feature
chain Antitrypsin
Uniprot
C0J8H7
A0A3G1T1R3
A0A3S2M6W8
G9F9J3
A0A3G1T1Q1
A0A194R0R3
+ More
A0A2A4J7B9 A0A194PXE0 G9F9J2 A0A212EGT4 A0A1E1WCC9 A0A194PXB8 A0A1E1WM18 A0A1E1W9X2 K4P392 Q25498 B4XH99 F5B4G8 B4XHA0 Q25495 B4XHA1 Q25492 A0A385XRR7 A0A3S2NJW5 Q25500 C7ASM2 P22922 Q25499 Q25526 Q25497 A3KEZ7 B4XH98 Q25491 A0A194PZ13 C7ASM3 C7ASM9 P80034 I4DJC8 Q25494 P14754 C7ASM7 A0A0M4K7J2 Q60FS0 C7ASM4 A0A3G1T1E7 A3KEZ8 C7ASM6 A0A194R1A3 A0A212FD15 B0FZ68 B0FZ61 C5I794 B0FZ67 J7HI16 B0FZ65 Q8IS84 Q8IS86 Q9NGS0 A0A194PIN8 B0FZ66 Q8IS85 A0A3S2NC81 E2GHX4 B0FZ60 A0A0M4JBR8 A0A212EXP2 B0FZ63 C5I795 B0FZ62 A0A2H1VE36 A0A2A4JPH1 B2X122 C0J8H0 A0A3S2TR38 Q5MGH1 A0A2A4JQ63 A0A0L7LMC8 A0A2H1VAC5 Q8IS50 Q5MGH2
A0A2A4J7B9 A0A194PXE0 G9F9J2 A0A212EGT4 A0A1E1WCC9 A0A194PXB8 A0A1E1WM18 A0A1E1W9X2 K4P392 Q25498 B4XH99 F5B4G8 B4XHA0 Q25495 B4XHA1 Q25492 A0A385XRR7 A0A3S2NJW5 Q25500 C7ASM2 P22922 Q25499 Q25526 Q25497 A3KEZ7 B4XH98 Q25491 A0A194PZ13 C7ASM3 C7ASM9 P80034 I4DJC8 Q25494 P14754 C7ASM7 A0A0M4K7J2 Q60FS0 C7ASM4 A0A3G1T1E7 A3KEZ8 C7ASM6 A0A194R1A3 A0A212FD15 B0FZ68 B0FZ61 C5I794 B0FZ67 J7HI16 B0FZ65 Q8IS84 Q8IS86 Q9NGS0 A0A194PIN8 B0FZ66 Q8IS85 A0A3S2NC81 E2GHX4 B0FZ60 A0A0M4JBR8 A0A212EXP2 B0FZ63 C5I795 B0FZ62 A0A2H1VE36 A0A2A4JPH1 B2X122 C0J8H0 A0A3S2TR38 Q5MGH1 A0A2A4JQ63 A0A0L7LMC8 A0A2H1VAC5 Q8IS50 Q5MGH2
Pubmed
EMBL
EU935629
ACG61191.1
MG770321
AXY94923.1
RSAL01000018
RVE52848.1
+ More
JN033738 AEW46891.2 MG770320 AXY94922.1 KQ460883 KPJ11292.1 NWSH01002542 PCG68017.1 KQ459586 KPI97992.1 JN033737 AEW46890.2 AGBW02015025 OWR40695.1 GDQN01006457 JAT84597.1 KPI97991.1 GDQN01003022 JAT88032.1 GDQN01007288 JAT83766.1 JX524148 AFV46312.1 U58361 AAC47342.1 EU040208 ABW17156.1 HQ615869 ADT63775.1 EU040209 ABW17157.1 AAC47341.1 EU040210 ABW17158.1 AAC47338.1 MG753799 AYC12556.1 RVE52846.1 AAC47335.1 FJ613793 ACT36272.1 D00738 AAC47340.1 L20793 AAA29334.1 AAC47333.1 AB295498 BAF48334.1 EU040207 ABW17155.1 AAC47334.1 KPI97989.1 FJ613794 FJ613796 ACT36273.1 ACT36275.1 FJ613797 ACT36276.1 AK401396 BAM18018.1 AAC47332.1 M23438 L20792 ACT36277.1 ACT36279.1 KR003729 ALD62507.1 AB189029 BAD52261.1 FJ613795 ACT36274.1 MG992370 AXY94808.1 AB295499 BAF48335.1 ACT36278.1 KPJ11294.1 AGBW02009121 OWR51607.1 EU368832 ABY68555.1 ABY68562.1 FJ763763 ACR56864.1 ABY68556.1 JN400442 AFQ01141.1 ABY68558.1 AY148485 AAN71634.1 AY148483 AAN71632.1 AF242200 AAF61252.1 KQ459602 KPI93286.1 ABY68557.1 AY148484 AAN71633.1 RSAL01000236 RVE43711.1 HM357255 ADN52338.1 ABY68563.1 KR003725 ALD62503.1 AGBW02011729 OWR46227.1 ABY68560.1 FJ763764 ACR56865.1 ABY68561.1 ODYU01002062 SOQ39103.1 NWSH01000840 PCG73901.1 EF584499 ABU62829.1 EU935622 ACG61184.1 RVE52836.1 AY829815 AAV91429.1 PCG73906.1 JTDY01000556 KOB76688.1 ODYU01001314 SOQ37342.1 AY150533 AAN73322.1 AY829814 AAV91428.1
JN033738 AEW46891.2 MG770320 AXY94922.1 KQ460883 KPJ11292.1 NWSH01002542 PCG68017.1 KQ459586 KPI97992.1 JN033737 AEW46890.2 AGBW02015025 OWR40695.1 GDQN01006457 JAT84597.1 KPI97991.1 GDQN01003022 JAT88032.1 GDQN01007288 JAT83766.1 JX524148 AFV46312.1 U58361 AAC47342.1 EU040208 ABW17156.1 HQ615869 ADT63775.1 EU040209 ABW17157.1 AAC47341.1 EU040210 ABW17158.1 AAC47338.1 MG753799 AYC12556.1 RVE52846.1 AAC47335.1 FJ613793 ACT36272.1 D00738 AAC47340.1 L20793 AAA29334.1 AAC47333.1 AB295498 BAF48334.1 EU040207 ABW17155.1 AAC47334.1 KPI97989.1 FJ613794 FJ613796 ACT36273.1 ACT36275.1 FJ613797 ACT36276.1 AK401396 BAM18018.1 AAC47332.1 M23438 L20792 ACT36277.1 ACT36279.1 KR003729 ALD62507.1 AB189029 BAD52261.1 FJ613795 ACT36274.1 MG992370 AXY94808.1 AB295499 BAF48335.1 ACT36278.1 KPJ11294.1 AGBW02009121 OWR51607.1 EU368832 ABY68555.1 ABY68562.1 FJ763763 ACR56864.1 ABY68556.1 JN400442 AFQ01141.1 ABY68558.1 AY148485 AAN71634.1 AY148483 AAN71632.1 AF242200 AAF61252.1 KQ459602 KPI93286.1 ABY68557.1 AY148484 AAN71633.1 RSAL01000236 RVE43711.1 HM357255 ADN52338.1 ABY68563.1 KR003725 ALD62503.1 AGBW02011729 OWR46227.1 ABY68560.1 FJ763764 ACR56865.1 ABY68561.1 ODYU01002062 SOQ39103.1 NWSH01000840 PCG73901.1 EF584499 ABU62829.1 EU935622 ACG61184.1 RVE52836.1 AY829815 AAV91429.1 PCG73906.1 JTDY01000556 KOB76688.1 ODYU01001314 SOQ37342.1 AY150533 AAN73322.1 AY829814 AAV91428.1
Proteomes
Interpro
Gene 3D
ProteinModelPortal
C0J8H7
A0A3G1T1R3
A0A3S2M6W8
G9F9J3
A0A3G1T1Q1
A0A194R0R3
+ More
A0A2A4J7B9 A0A194PXE0 G9F9J2 A0A212EGT4 A0A1E1WCC9 A0A194PXB8 A0A1E1WM18 A0A1E1W9X2 K4P392 Q25498 B4XH99 F5B4G8 B4XHA0 Q25495 B4XHA1 Q25492 A0A385XRR7 A0A3S2NJW5 Q25500 C7ASM2 P22922 Q25499 Q25526 Q25497 A3KEZ7 B4XH98 Q25491 A0A194PZ13 C7ASM3 C7ASM9 P80034 I4DJC8 Q25494 P14754 C7ASM7 A0A0M4K7J2 Q60FS0 C7ASM4 A0A3G1T1E7 A3KEZ8 C7ASM6 A0A194R1A3 A0A212FD15 B0FZ68 B0FZ61 C5I794 B0FZ67 J7HI16 B0FZ65 Q8IS84 Q8IS86 Q9NGS0 A0A194PIN8 B0FZ66 Q8IS85 A0A3S2NC81 E2GHX4 B0FZ60 A0A0M4JBR8 A0A212EXP2 B0FZ63 C5I795 B0FZ62 A0A2H1VE36 A0A2A4JPH1 B2X122 C0J8H0 A0A3S2TR38 Q5MGH1 A0A2A4JQ63 A0A0L7LMC8 A0A2H1VAC5 Q8IS50 Q5MGH2
A0A2A4J7B9 A0A194PXE0 G9F9J2 A0A212EGT4 A0A1E1WCC9 A0A194PXB8 A0A1E1WM18 A0A1E1W9X2 K4P392 Q25498 B4XH99 F5B4G8 B4XHA0 Q25495 B4XHA1 Q25492 A0A385XRR7 A0A3S2NJW5 Q25500 C7ASM2 P22922 Q25499 Q25526 Q25497 A3KEZ7 B4XH98 Q25491 A0A194PZ13 C7ASM3 C7ASM9 P80034 I4DJC8 Q25494 P14754 C7ASM7 A0A0M4K7J2 Q60FS0 C7ASM4 A0A3G1T1E7 A3KEZ8 C7ASM6 A0A194R1A3 A0A212FD15 B0FZ68 B0FZ61 C5I794 B0FZ67 J7HI16 B0FZ65 Q8IS84 Q8IS86 Q9NGS0 A0A194PIN8 B0FZ66 Q8IS85 A0A3S2NC81 E2GHX4 B0FZ60 A0A0M4JBR8 A0A212EXP2 B0FZ63 C5I795 B0FZ62 A0A2H1VE36 A0A2A4JPH1 B2X122 C0J8H0 A0A3S2TR38 Q5MGH1 A0A2A4JQ63 A0A0L7LMC8 A0A2H1VAC5 Q8IS50 Q5MGH2
PDB
1SEK
E-value=3.79165e-59,
Score=579
Ontologies
PANTHER
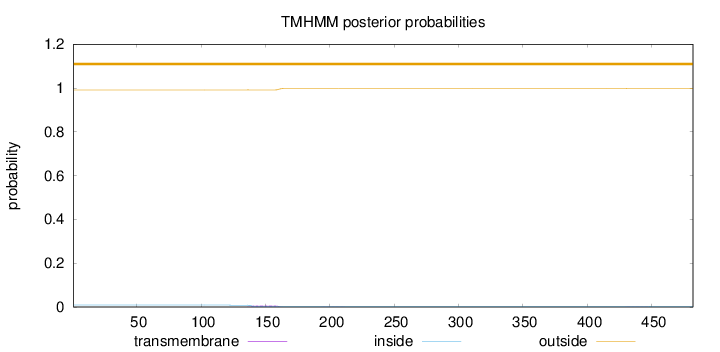
Topology
Subcellular location
Secreted
Length:
482
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.15592
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00857
outside
1 - 482
Population Genetic Test Statistics
Pi
179.7112
Theta
165.043365
Tajima's D
-1.069082
CLR
3.583155
CSRT
0.127243637818109
Interpretation
Uncertain
