Gene
KWMTBOMO12870
Pre Gene Modal
BGIBMGA012706
Annotation
PREDICTED:_X-linked_retinitis_pigmentosa_GTPase_regulator-like_isoform_X1_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 1.484 Nuclear Reliability : 1.855
Sequence
CDS
ATGCTTTCTTGGTACACAGAGAATGGTCGTGTGTTTGTGTTCGGAGCTAACACGTGGGGTCAGCTGGGCCTTGGTCATAAAGATGAGGTGACGCGGCCTAGTTGCGTGAAATGGCTCAAACCGCAGCGAACAATGTTCGTTGCGTGCGGACGAGCGCACACCGTTTTCGTTACAGAAACAAACGCTATATACACCGTGGGCTGCAACGATGAGGGGCAGCTCGGCACCGGCGATATGGAACACAGCACAGTTCCGCAAAGTGTCGAACTTGAGGAGCCCATAGTCGTGAAACAGATATCCGCTGGAAGCAACCACACGGCTCTACTGACTGAGGATGGTCGCGTGTTTATGTGCGGCTCCAACTCCGAAGGGCAGCTCGGTTTAGGCGAAGACTCACGATCCGCTGTACGGTTCACTGAACTCAAGTTCATGGAGAAGATAGCCTTCATCGAATGCGGGTATTATCACACCGTCTTTATAACTACCAAAGGAGCAGTCTTTGTAACTGGCGACAACGAAAATCAGAAACTAGGTATCCAGAAAATAGCCACAAGCATCTACGTTCCGGAAGTTTTGCCCATAAGTACTCCAATAAAGAGCGCCTGCTGTGGGGCTGCACACACGTTCCTTCTGACAATGGATGAATCTAAAATATTGGGCTTCGGTTCCAATGAAAAGGGTCAGCTGGGAATGCCTCGAGAGGTCGAGTTTGTTACTGAGCCTAGCGAAATTGACATGGACATTATGTTCGACGGATACCAACTAAAGTTGGTCGCCTACAACGGACTACTGTACACGTGCGGCGAGTCGAGGCATAATAAACTGTGTTTGGAAGATATCGACGACACCAACTCAAACGAGGTGGTCCCCAATCAATACACCCCTAAACGGGTCACCGCCATGAATGGGCACATCGTAGACAATGTGGCGTGTGGGGGCTGCCACACTCTACTTACGGCCACCAAAGGCTCGAACGCGGACTTCACGAACAACGATTTCAACATTATTAACGAAGCCACCAGACAAATAACTCTCACCGAATTACCGCCCTTACAGAAAATACCAGCGATGAACAAAACCGAAGAGAATCCGTTCTTAATTGAAGACGTGACAAACTCTAACGTGGACGACAAACAGAAAGCGGATAACTTGAATGGAAACACGAACGAGGAATCCGGATCAATAGACTCGTTAGAGGCCAACGTGAGCGGATCACACATCGTAAACATAAACGACCTTGAAATTGAAAATGGAAGTACGCATGACCTGGCTGACCTAAACAAAGACAACATTGATCGTGTTTCCGACACAGGTGACTCGGCTATGGACGTGGTCGAAATTATGGACACGGTGCTTGACGATATGCAGACAAATGTTCATGATTCTGTAAAAGAGGTCACCGATACAGTCCATGAGAAAAAGGAAGTGATTGAAGAAGTAATTCACAAAACGGAAGCAAATGCAAAAGGGCTGCGCGATGATGTCACAAAGATTTTGACCGTTCCTGATAAAGTAGTCGAAATTGCAAATTCCCCGCTGCCTAAAACACCCATATTGCAAGAACTACTCCACGTTCCCGATATATCACAACTCAGTCCTGCTCACAGTAAGCACAGTGACGAGAAAAGTGAAACAGGCCAGTCCGAAAACGAAACAATAACATGCGCATCGGATAAATCTAGTCCTCAAGCTGGGAAGACAGCTAAAACACAAGCCGTTATGCCGATTGTCAGGAGCGAGGACCATATAGACTCTCATGTTGATGAAACCAATGAAGAATCAGATGTTGTTGTTAATAAATTGGAAGAGAAGGGTCGATTTGCGAGGATGTTTCAGTCAATACGAGACAAGGAAAACTCTTGTATGGGGAAGGGCAAAGTCATTGAAGAGAAAGTTGTTGAGACGGTTCACAAAGGCATCGACAACGCCGAAGAGTCGATTCACAAAGTTGAAGAAACCGTTGAAACAGAGATACGGAACCAGACAAACTCTAGAACCTGCACTATATTGTAA
Protein
MLSWYTENGRVFVFGANTWGQLGLGHKDEVTRPSCVKWLKPQRTMFVACGRAHTVFVTETNAIYTVGCNDEGQLGTGDMEHSTVPQSVELEEPIVVKQISAGSNHTALLTEDGRVFMCGSNSEGQLGLGEDSRSAVRFTELKFMEKIAFIECGYYHTVFITTKGAVFVTGDNENQKLGIQKIATSIYVPEVLPISTPIKSACCGAAHTFLLTMDESKILGFGSNEKGQLGMPREVEFVTEPSEIDMDIMFDGYQLKLVAYNGLLYTCGESRHNKLCLEDIDDTNSNEVVPNQYTPKRVTAMNGHIVDNVACGGCHTLLTATKGSNADFTNNDFNIINEATRQITLTELPPLQKIPAMNKTEENPFLIEDVTNSNVDDKQKADNLNGNTNEESGSIDSLEANVSGSHIVNINDLEIENGSTHDLADLNKDNIDRVSDTGDSAMDVVEIMDTVLDDMQTNVHDSVKEVTDTVHEKKEVIEEVIHKTEANAKGLRDDVTKILTVPDKVVEIANSPLPKTPILQELLHVPDISQLSPAHSKHSDEKSETGQSENETITCASDKSSPQAGKTAKTQAVMPIVRSEDHIDSHVDETNEESDVVVNKLEEKGRFARMFQSIRDKENSCMGKGKVIEEKVVETVHKGIDNAEESIHKVEETVETEIRNQTNSRTCTIL
Summary
Uniprot
EMBL
Proteomes
PRIDE
Pfam
PF00415 RCC1
SUPFAM
SSF50985
SSF50985
Gene 3D
ProteinModelPortal
PDB
4JHN
E-value=5.04055e-55,
Score=545
Ontologies
GO
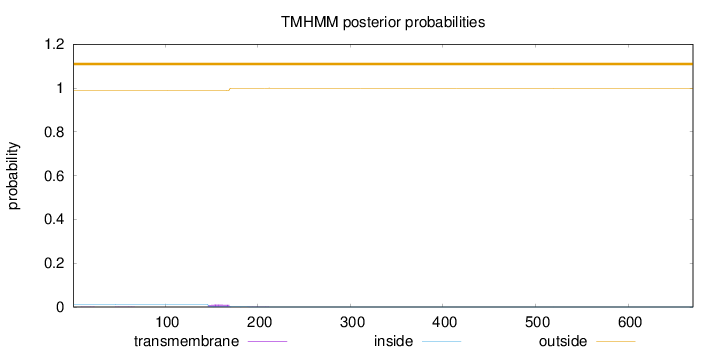
Topology
Length:
670
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.2516
Exp number, first 60 AAs:
0.00802
Total prob of N-in:
0.01147
outside
1 - 670
Population Genetic Test Statistics
Pi
148.59268
Theta
144.833189
Tajima's D
0.098727
CLR
22.719337
CSRT
0.397630118494075
Interpretation
Uncertain
