Gene
KWMTBOMO12855
Pre Gene Modal
BGIBMGA012668
Annotation
PREDICTED:_netrin_receptor_UNC5C-like_isoform_X1_[Bombyx_mori]
Location in the cell
Extracellular Reliability : 1.986 Nuclear Reliability : 1.583
Sequence
CDS
ATGGCGGGAGGGGTTGCGCTCCTCGTGCTATGGGTGGTCGCAGTCACAGCTGATGGTAAACACGACGTGTCAAACTCAACTGAAGCCGCAACGACAGCGTTTCCCACAGCACCCATACCACTGCGACCAGAATATGCTCATCCGGCTGGAGTAGAAGCATTAGACCCTTACAACTCAGAGAAGAACAACCCGCCAATTCATCACAAAGACTATTTAGAAGAAGACTACGACCATGAATCTCACAAAGAAGACGACCACGACAACAGAGACGATACAGAAGAATTACCTTCGCTATCAGGAGACCATTTGCTCCAAAGCGCGACGGATAATCTGCCAATGTTTCTATTAGAGCCACAACACGCGTTTGTTGTGAGAAATAAACCGGCCACGTTAAGATGTCGGGCGGCTAATGCACTCCAAGTGTATTTTAAGTGCAACGAAGTTCGATCGGTTGCGAGCACACAGTTCGAGTTCGTGGATCCCCAAAGCGGAGTTAGGATCGTTGAAGCCGAATGTAACGTGACGCGGGATCAGTTAGATGAATATTTCGGTGAGGACAGATTTACGTGCACTTGCTACGCTTGGAGCAGCAGAGGAGACATCCGAAGCCAGCCGGCTGTCGTGGAACTTGCCTATCTAAAGAAGCCATTCGCGGTATCGCCGTCAGCGATTTCAGTAGAGGCCGGTTCAGCCGCTTCTCTGCGCTGCTCTCCACCAATGGCTGCCCCACCTCCTCACATATCCTGGTTGAAGCACGGGGCCCCTCTGCAGCAGGATCACAACGTGCTGGTTTCTGCAGAAGGAAATCTGCTGATATCTCGTTCCACTTTGCAAGACATGGCAAACTACAGCTGCGTTGCTGAGAACATAGCCGGGAAGAGAATATCTGAGCCCGCTCTCCTAACTGTTTACGTGAACGGAGGGTGGAGTGCCTGGGGTCCCTGGACTCAGTGTCGTTGCAACGGCCGAATCTCGAGCGGGCAGCGTCGCACGAGGACCTGCACCGAACCTCATCCTCTCAACGGAGGAGCTCCGTGTCAAGGTCAGGCCGTGCAGAGGACCGCGGACTGCCTTCCGTGTCAAAGTGCACTCTCTGGACGATGGGGGGCTTGGTCGGAGTGGTCAGTGTGCGGTGCAGACTGCCGGCAGATCCGACGAAGATCTTGTACCGGCGATACGCCTTGTTCCGGGGCACACGCGCAACACGCCGATTGCGTTGGAGACTACTGTGGAGTTCATGGAGTTGCCGCGCACAGTGACATCCCTCTGTACATCGGAATCGCCGTCGCCGTGGTCGTATTCCTGGCTGGTGCTGCCGTGGTCTACAAACTGCTGCAGAGGAAGACCCGAGACCATTCTTTGTACACTATGACTAGGACAGATTTCCAACCAGAAATATACCCGACAGTGGAGAAGCAGTCCTTATCGCTAGCTCCTGATCTAATGCAGCACAGACCCCCAAGATACGAACATCCCCAACCCGAACATCACTATGATGTGCCACATCTAACGAACAGTTACGCTTCACCTGTGGACCACCAAGTGACTCCATCTGCCTCTACCAAGGGACAGAGCGACGATTATGAAAGCAAACCGTACAGCGAATCAGAACATTCAGCCTCTAGTTGTTTCACGAGCTCCGGTTCGATGTACGACGCGGCCAATGAATCCGTGACACTCCAACTGGCAGAGTTCGTCTCCAATTCGCCCACGTCACAATCCATGTCGGTCTCCAGTGCCGGAGCCAGACTGGCCTTGCCCGCAGCGGGGGTGGCCCTTTCAGTTCCTGAAGGAGCCCTGGCCAGGGGTAGACGAGAGCAGATCTATGTGGCTGTCGTCAAGGATGACAGATACAGGCCGCGACTCGGAAAAGGTATCACGCAGCTGAGTCCGGTGGTTAAATGCGGCCCGCCGCGTCAGACATTAAACAAGTCCGTCGTCTTACAAATACCACATTGCGCCAGCCTCAAGCATGGCTTCTGGAACCTGTCTCTGTACGCCATAGACCACAACAGTAAGCCAGAGGGCGGCCAGTCTCAGTGGAAGAAGATCGTGGGCCTCGGTCAAGAAACCATCAACACCCCAATCTTCACACAGCTCGACAACGACAAGATATTCCTTGTGACGGACATGTTGTCCACGTTCGTATTGGTCGGTGAAAGTTTCAATGGTAAGGCCGTAAAGGCACTACAATTAGCGATCTACGCCCCGACCGTGCTCAACGAAACCAGTTCCGAATATAGCATTAGATTGTACGTGTTCGAAGACACGCCTTGTGCGGCTTACTATTGTCAGGAACAAGAGAAAAAACTCGGCGGAGTGTTGCTTGAACGACCGAAAACTTTATTGTTCCAAGACGGAGGCTCCCATCTTTGTTTGAATCTGGAGCACGTCAGTCCCGGCTGGAAAGCGAAGCCGGGCATCGGTTATCAGGAGATCCCATTCAATCACGTGTGGAGCTCGAACTACAACGCTCTGCATTGCAGCTTCACTCTCGATCGGACGCACGACTGCGACAATATTGATTTTACGATCTCAGCCTGTCAGAGAACGAATCCATCTTATAAGCAAACGTTCAGGATATGCCTCGAAGACATGCAGAGTCGGTCGCCACGGGGCAGATCGCCGCGCGAATCCCAGCGCAGTTTTAATATTACCGAGAATCTCTTGGATGGTCGTATGGGTCGAAACGACGAGGGCAGTGAGGGTAGCGTGAAAAGAAACATCACTGTTAATGAGACGGGCGTGAACGAATGCTCCGGAGAGGGCCCTTTCAAATTGAGCGGGAAGATAAAAAGGCAGCTTTGTGCCATTTTAGACCCTCCTAACGCCCGTGGAAATGATTGGCGTGCTCTGGCATCGCGGTTGGGCGTAGATCGCTATACAACTTGGTTCGCCACAAAGAGCTCACCCACTGAAGCTATATTAGAGCTGTGGGAATGCAGGGAGCGTCATCCTGGGGCCATTCCCGCTCTGGCGACCGCATTGCGACAGATGGACAGACACGACGCCGCTGATCTACTTCATATACGGCCATCTTGGTTATGA
Protein
MAGGVALLVLWVVAVTADGKHDVSNSTEAATTAFPTAPIPLRPEYAHPAGVEALDPYNSEKNNPPIHHKDYLEEDYDHESHKEDDHDNRDDTEELPSLSGDHLLQSATDNLPMFLLEPQHAFVVRNKPATLRCRAANALQVYFKCNEVRSVASTQFEFVDPQSGVRIVEAECNVTRDQLDEYFGEDRFTCTCYAWSSRGDIRSQPAVVELAYLKKPFAVSPSAISVEAGSAASLRCSPPMAAPPPHISWLKHGAPLQQDHNVLVSAEGNLLISRSTLQDMANYSCVAENIAGKRISEPALLTVYVNGGWSAWGPWTQCRCNGRISSGQRRTRTCTEPHPLNGGAPCQGQAVQRTADCLPCQSALSGRWGAWSEWSVCGADCRQIRRRSCTGDTPCSGAHAQHADCVGDYCGVHGVAAHSDIPLYIGIAVAVVVFLAGAAVVYKLLQRKTRDHSLYTMTRTDFQPEIYPTVEKQSLSLAPDLMQHRPPRYEHPQPEHHYDVPHLTNSYASPVDHQVTPSASTKGQSDDYESKPYSESEHSASSCFTSSGSMYDAANESVTLQLAEFVSNSPTSQSMSVSSAGARLALPAAGVALSVPEGALARGRREQIYVAVVKDDRYRPRLGKGITQLSPVVKCGPPRQTLNKSVVLQIPHCASLKHGFWNLSLYAIDHNSKPEGGQSQWKKIVGLGQETINTPIFTQLDNDKIFLVTDMLSTFVLVGESFNGKAVKALQLAIYAPTVLNETSSEYSIRLYVFEDTPCAAYYCQEQEKKLGGVLLERPKTLLFQDGGSHLCLNLEHVSPGWKAKPGIGYQEIPFNHVWSSNYNALHCSFTLDRTHDCDNIDFTISACQRTNPSYKQTFRICLEDMQSRSPRGRSPRESQRSFNITENLLDGRMGRNDEGSEGSVKRNITVNETGVNECSGEGPFKLSGKIKRQLCAILDPPNARGNDWRALASRLGVDRYTTWFATKSSPTEAILELWECRERHPGAIPALATALRQMDRHDAADLLHIRPSWL
Summary
Similarity
Belongs to the serpin family.
Uniprot
A0A2A4JLB3
A0A2W1BXA1
A0A2A4JM37
A0A437BJV8
A0A212EVY5
A0A194Q8C5
+ More
A0A1Y1KGW7 A0A1B6BY64 A0A067RP65 A0A146LCK0 A0A0K8TDJ2 A0A0A9VZT2 A0A1B6LH58 A0A195CJV7 A0A1B6KVK9 A0A158NTE6 A0A195DVW8 A0A026VX76 A0A088A971 A0A195B423 A0A3L8DS02 A0A154PEH5 A0A2A4IZN1 A0A0L7RHT7 A0A151X1U2 A0A2W1BMT9 F4W406 A0A212EZX5 A0A1B6CT56 A0A0N1I685 A0A0M9A636 A0A194PXF9 A0A3S2NWX0 J9KU90 E9G837 A0A0P6BFD9 A0A0N8BMV1 A0A0P5B8K0 A0A2H8TRB2 T1IHT4 A0A0P5LXZ4 A0A0P5MDR1 A0A139WHJ8 A0A026VUN0 A0A151X1E9 A0A195ETG5 E2APQ4 V3ZRF4 A0A195CJV1 F4W410 A0A195EUI3 A0A3L8DRW3 A0A0N8BDX1 A0A195DVL0 A0A158NK16 A0A195B393 A0A210QTQ7 A0A1S3JPZ4 A0A088AWA8 A0A1S3JPB8 A0A1S3JQG5 A0A154PEA8 A0A2S2PVW1 A0A0P5UBF6 A0A0L7QTL0 W8B2P2 W8AYM5 A0A0A1X6V4 A0A3B3B8P1 A0A3B3I060 A0A3P9IY59 A0A0M9A8F8 A0A3P9MHT7 A0A2A3EFS9 A0A3Q0SUT7 A0A2I4AUK5 A0A3Q4GB99 A0A3B3B8Q1 I3K934 H2LUM0 A0A3B4FMA2 A0A3B5KZF6 A0A3P9HB44 A0A3S2MSF4 A0A3P8PY04 A0A3P9CRB2 A0A3P9L932 A0A3P8SXW2 A0A1A8EAU7 A0A1A8P3X2 A0A1A8HFL4 A0A3Q3S554 A0A3Q2QYD0 A0A1A8Q6R1 A0A3Q3CCL9 A0A1A8JPX1
A0A1Y1KGW7 A0A1B6BY64 A0A067RP65 A0A146LCK0 A0A0K8TDJ2 A0A0A9VZT2 A0A1B6LH58 A0A195CJV7 A0A1B6KVK9 A0A158NTE6 A0A195DVW8 A0A026VX76 A0A088A971 A0A195B423 A0A3L8DS02 A0A154PEH5 A0A2A4IZN1 A0A0L7RHT7 A0A151X1U2 A0A2W1BMT9 F4W406 A0A212EZX5 A0A1B6CT56 A0A0N1I685 A0A0M9A636 A0A194PXF9 A0A3S2NWX0 J9KU90 E9G837 A0A0P6BFD9 A0A0N8BMV1 A0A0P5B8K0 A0A2H8TRB2 T1IHT4 A0A0P5LXZ4 A0A0P5MDR1 A0A139WHJ8 A0A026VUN0 A0A151X1E9 A0A195ETG5 E2APQ4 V3ZRF4 A0A195CJV1 F4W410 A0A195EUI3 A0A3L8DRW3 A0A0N8BDX1 A0A195DVL0 A0A158NK16 A0A195B393 A0A210QTQ7 A0A1S3JPZ4 A0A088AWA8 A0A1S3JPB8 A0A1S3JQG5 A0A154PEA8 A0A2S2PVW1 A0A0P5UBF6 A0A0L7QTL0 W8B2P2 W8AYM5 A0A0A1X6V4 A0A3B3B8P1 A0A3B3I060 A0A3P9IY59 A0A0M9A8F8 A0A3P9MHT7 A0A2A3EFS9 A0A3Q0SUT7 A0A2I4AUK5 A0A3Q4GB99 A0A3B3B8Q1 I3K934 H2LUM0 A0A3B4FMA2 A0A3B5KZF6 A0A3P9HB44 A0A3S2MSF4 A0A3P8PY04 A0A3P9CRB2 A0A3P9L932 A0A3P8SXW2 A0A1A8EAU7 A0A1A8P3X2 A0A1A8HFL4 A0A3Q3S554 A0A3Q2QYD0 A0A1A8Q6R1 A0A3Q3CCL9 A0A1A8JPX1
Pubmed
EMBL
NWSH01001062
PCG72847.1
KZ149907
PZC78254.1
PCG72846.1
RSAL01000046
+ More
RVE50538.1 AGBW02012107 OWR45646.1 KQ459324 KPJ01644.1 GEZM01086422 JAV59460.1 GEDC01031076 JAS06222.1 KK852508 KDR22415.1 GDHC01018809 GDHC01013807 JAP99819.1 JAQ04822.1 GBRD01002257 JAG63564.1 GBHO01043896 JAF99707.1 GEBQ01016935 JAT23042.1 KQ977642 KYN01011.1 GEBQ01024488 JAT15489.1 ADTU01025884 ADTU01025885 KQ980295 KYN16887.1 KK107847 EZA47459.1 KQ976625 KYM78959.1 QOIP01000005 RLU23184.1 KQ434870 KZC09610.1 NWSH01004342 PCG65211.1 KQ414585 KOC70552.1 KQ982585 KYQ54236.1 KZ150134 PZC73003.1 GL887491 EGI71002.1 AGBW02011206 OWR47055.1 GEDC01020737 JAS16561.1 KQ460874 KPJ11666.1 KQ435727 KOX77995.1 KQ459586 KPI98031.1 RSAL01000124 RVE46623.1 ABLF02041018 GL732534 EFX84751.1 GDIP01015914 JAM87801.1 GDIQ01162251 JAK89474.1 GDIP01188765 JAJ34637.1 GFXV01004337 MBW16142.1 JH430009 GDIQ01163557 JAK88168.1 GDIQ01168858 JAK82867.1 KQ971343 KYB27433.1 EZA47457.1 KYQ54241.1 KQ981979 KYN31545.1 GL441572 EFN64605.1 KB203683 ESO83456.1 KYN01006.1 EGI71006.1 KYN31539.1 RLU23185.1 GDIQ01187291 JAK64434.1 KYN16881.1 ADTU01018638 ADTU01018639 KYM78963.1 NEDP02001973 OWF52102.1 KZC09608.1 GGMS01000443 MBY69646.1 GDIP01115424 JAL88290.1 KQ414755 KOC61811.1 GAMC01015247 JAB91308.1 GAMC01015248 JAB91307.1 GBXI01007470 JAD06822.1 KOX77999.1 KZ288263 PBC30344.1 AERX01032132 CM012448 RVE65808.1 HAEA01013744 SBQ42224.1 HAEI01001394 SBR75986.1 HAEB01008131 HAEC01013243 SBQ81460.1 HAEH01010337 SBR89445.1 HAEE01002135 SBR22155.1
RVE50538.1 AGBW02012107 OWR45646.1 KQ459324 KPJ01644.1 GEZM01086422 JAV59460.1 GEDC01031076 JAS06222.1 KK852508 KDR22415.1 GDHC01018809 GDHC01013807 JAP99819.1 JAQ04822.1 GBRD01002257 JAG63564.1 GBHO01043896 JAF99707.1 GEBQ01016935 JAT23042.1 KQ977642 KYN01011.1 GEBQ01024488 JAT15489.1 ADTU01025884 ADTU01025885 KQ980295 KYN16887.1 KK107847 EZA47459.1 KQ976625 KYM78959.1 QOIP01000005 RLU23184.1 KQ434870 KZC09610.1 NWSH01004342 PCG65211.1 KQ414585 KOC70552.1 KQ982585 KYQ54236.1 KZ150134 PZC73003.1 GL887491 EGI71002.1 AGBW02011206 OWR47055.1 GEDC01020737 JAS16561.1 KQ460874 KPJ11666.1 KQ435727 KOX77995.1 KQ459586 KPI98031.1 RSAL01000124 RVE46623.1 ABLF02041018 GL732534 EFX84751.1 GDIP01015914 JAM87801.1 GDIQ01162251 JAK89474.1 GDIP01188765 JAJ34637.1 GFXV01004337 MBW16142.1 JH430009 GDIQ01163557 JAK88168.1 GDIQ01168858 JAK82867.1 KQ971343 KYB27433.1 EZA47457.1 KYQ54241.1 KQ981979 KYN31545.1 GL441572 EFN64605.1 KB203683 ESO83456.1 KYN01006.1 EGI71006.1 KYN31539.1 RLU23185.1 GDIQ01187291 JAK64434.1 KYN16881.1 ADTU01018638 ADTU01018639 KYM78963.1 NEDP02001973 OWF52102.1 KZC09608.1 GGMS01000443 MBY69646.1 GDIP01115424 JAL88290.1 KQ414755 KOC61811.1 GAMC01015247 JAB91308.1 GAMC01015248 JAB91307.1 GBXI01007470 JAD06822.1 KOX77999.1 KZ288263 PBC30344.1 AERX01032132 CM012448 RVE65808.1 HAEA01013744 SBQ42224.1 HAEI01001394 SBR75986.1 HAEB01008131 HAEC01013243 SBQ81460.1 HAEH01010337 SBR89445.1 HAEE01002135 SBR22155.1
Proteomes
UP000218220
UP000283053
UP000007151
UP000053268
UP000027135
UP000078542
+ More
UP000005205 UP000078492 UP000053097 UP000005203 UP000078540 UP000279307 UP000076502 UP000053825 UP000075809 UP000007755 UP000053240 UP000053105 UP000007819 UP000000305 UP000007266 UP000078541 UP000000311 UP000030746 UP000242188 UP000085678 UP000261560 UP000001038 UP000265200 UP000265180 UP000242457 UP000261340 UP000192220 UP000261580 UP000005207 UP000261460 UP000261380 UP000265100 UP000265160 UP000265080 UP000261640 UP000265000 UP000264840
UP000005205 UP000078492 UP000053097 UP000005203 UP000078540 UP000279307 UP000076502 UP000053825 UP000075809 UP000007755 UP000053240 UP000053105 UP000007819 UP000000305 UP000007266 UP000078541 UP000000311 UP000030746 UP000242188 UP000085678 UP000261560 UP000001038 UP000265200 UP000265180 UP000242457 UP000261340 UP000192220 UP000261580 UP000005207 UP000261460 UP000261380 UP000265100 UP000265160 UP000265080 UP000261640 UP000265000 UP000264840
Interpro
IPR033772
UPA
+ More
IPR013098 Ig_I-set
IPR007110 Ig-like_dom
IPR003599 Ig_sub
IPR000488 Death_domain
IPR013783 Ig-like_fold
IPR011029 DEATH-like_dom_sf
IPR000906 ZU5_dom
IPR003598 Ig_sub2
IPR036179 Ig-like_dom_sf
IPR037936 UNC5
IPR000884 TSP1_rpt
IPR036383 TSP1_rpt_sf
IPR042178 Serpin_sf_1
IPR042185 Serpin_sf_2
IPR023796 Serpin_dom
IPR036186 Serpin_sf
IPR042154 Death_UNC5C
IPR013098 Ig_I-set
IPR007110 Ig-like_dom
IPR003599 Ig_sub
IPR000488 Death_domain
IPR013783 Ig-like_fold
IPR011029 DEATH-like_dom_sf
IPR000906 ZU5_dom
IPR003598 Ig_sub2
IPR036179 Ig-like_dom_sf
IPR037936 UNC5
IPR000884 TSP1_rpt
IPR036383 TSP1_rpt_sf
IPR042178 Serpin_sf_1
IPR042185 Serpin_sf_2
IPR023796 Serpin_dom
IPR036186 Serpin_sf
IPR042154 Death_UNC5C
Gene 3D
ProteinModelPortal
A0A2A4JLB3
A0A2W1BXA1
A0A2A4JM37
A0A437BJV8
A0A212EVY5
A0A194Q8C5
+ More
A0A1Y1KGW7 A0A1B6BY64 A0A067RP65 A0A146LCK0 A0A0K8TDJ2 A0A0A9VZT2 A0A1B6LH58 A0A195CJV7 A0A1B6KVK9 A0A158NTE6 A0A195DVW8 A0A026VX76 A0A088A971 A0A195B423 A0A3L8DS02 A0A154PEH5 A0A2A4IZN1 A0A0L7RHT7 A0A151X1U2 A0A2W1BMT9 F4W406 A0A212EZX5 A0A1B6CT56 A0A0N1I685 A0A0M9A636 A0A194PXF9 A0A3S2NWX0 J9KU90 E9G837 A0A0P6BFD9 A0A0N8BMV1 A0A0P5B8K0 A0A2H8TRB2 T1IHT4 A0A0P5LXZ4 A0A0P5MDR1 A0A139WHJ8 A0A026VUN0 A0A151X1E9 A0A195ETG5 E2APQ4 V3ZRF4 A0A195CJV1 F4W410 A0A195EUI3 A0A3L8DRW3 A0A0N8BDX1 A0A195DVL0 A0A158NK16 A0A195B393 A0A210QTQ7 A0A1S3JPZ4 A0A088AWA8 A0A1S3JPB8 A0A1S3JQG5 A0A154PEA8 A0A2S2PVW1 A0A0P5UBF6 A0A0L7QTL0 W8B2P2 W8AYM5 A0A0A1X6V4 A0A3B3B8P1 A0A3B3I060 A0A3P9IY59 A0A0M9A8F8 A0A3P9MHT7 A0A2A3EFS9 A0A3Q0SUT7 A0A2I4AUK5 A0A3Q4GB99 A0A3B3B8Q1 I3K934 H2LUM0 A0A3B4FMA2 A0A3B5KZF6 A0A3P9HB44 A0A3S2MSF4 A0A3P8PY04 A0A3P9CRB2 A0A3P9L932 A0A3P8SXW2 A0A1A8EAU7 A0A1A8P3X2 A0A1A8HFL4 A0A3Q3S554 A0A3Q2QYD0 A0A1A8Q6R1 A0A3Q3CCL9 A0A1A8JPX1
A0A1Y1KGW7 A0A1B6BY64 A0A067RP65 A0A146LCK0 A0A0K8TDJ2 A0A0A9VZT2 A0A1B6LH58 A0A195CJV7 A0A1B6KVK9 A0A158NTE6 A0A195DVW8 A0A026VX76 A0A088A971 A0A195B423 A0A3L8DS02 A0A154PEH5 A0A2A4IZN1 A0A0L7RHT7 A0A151X1U2 A0A2W1BMT9 F4W406 A0A212EZX5 A0A1B6CT56 A0A0N1I685 A0A0M9A636 A0A194PXF9 A0A3S2NWX0 J9KU90 E9G837 A0A0P6BFD9 A0A0N8BMV1 A0A0P5B8K0 A0A2H8TRB2 T1IHT4 A0A0P5LXZ4 A0A0P5MDR1 A0A139WHJ8 A0A026VUN0 A0A151X1E9 A0A195ETG5 E2APQ4 V3ZRF4 A0A195CJV1 F4W410 A0A195EUI3 A0A3L8DRW3 A0A0N8BDX1 A0A195DVL0 A0A158NK16 A0A195B393 A0A210QTQ7 A0A1S3JPZ4 A0A088AWA8 A0A1S3JPB8 A0A1S3JQG5 A0A154PEA8 A0A2S2PVW1 A0A0P5UBF6 A0A0L7QTL0 W8B2P2 W8AYM5 A0A0A1X6V4 A0A3B3B8P1 A0A3B3I060 A0A3P9IY59 A0A0M9A8F8 A0A3P9MHT7 A0A2A3EFS9 A0A3Q0SUT7 A0A2I4AUK5 A0A3Q4GB99 A0A3B3B8Q1 I3K934 H2LUM0 A0A3B4FMA2 A0A3B5KZF6 A0A3P9HB44 A0A3S2MSF4 A0A3P8PY04 A0A3P9CRB2 A0A3P9L932 A0A3P8SXW2 A0A1A8EAU7 A0A1A8P3X2 A0A1A8HFL4 A0A3Q3S554 A0A3Q2QYD0 A0A1A8Q6R1 A0A3Q3CCL9 A0A1A8JPX1
PDB
3G5B
E-value=8.26336e-73,
Score=700
Ontologies
GO
PANTHER
Topology
SignalP
Position: 1 - 19,
Likelihood: 0.989115
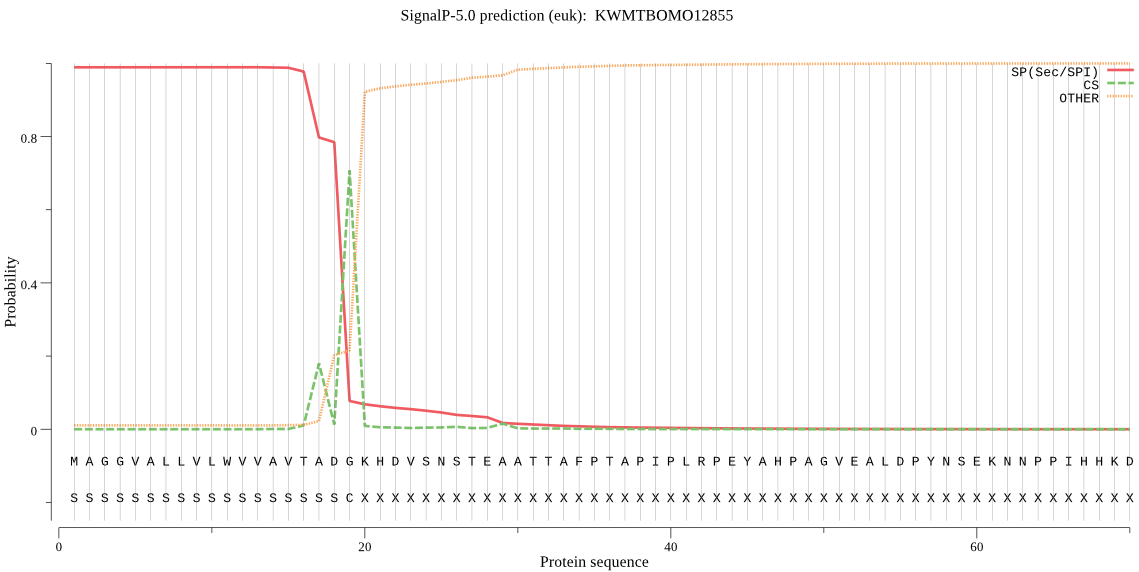
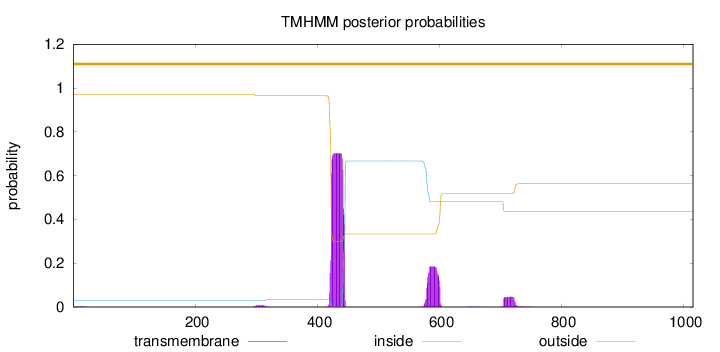
Length:
1015
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
20.40881
Exp number, first 60 AAs:
0.02491
Total prob of N-in:
0.02915
outside
1 - 1015
Population Genetic Test Statistics
Pi
288.574485
Theta
187.358997
Tajima's D
1.655391
CLR
0.104521
CSRT
0.819059047047648
Interpretation
Uncertain
