Gene
KWMTBOMO12840
Pre Gene Modal
BGIBMGA012722
Annotation
PREDICTED:_esterase_FE4-like_[Bombyx_mori]
Full name
Carboxylic ester hydrolase
Location in the cell
Cytoplasmic Reliability : 1.052 PlasmaMembrane Reliability : 1.425
Sequence
CDS
ATGCTACGACCAGTGACTCTAGCGGTGGTCGTGGTCGCCTTGTTGTCGGCGGCGGGGGCGAGGCTGAGAGTAGACCCGCTCGTTGAGACCCGGAAGGGACTCATCAAGGGTCTTAGGTCCGAAAATGGAGAGTTCTCCAAGTTCCTGGGTGTGCCCTACGCGTTAGTAGACGAGGATAATCCTTTTGGGTCCTCAGTGCCGCACCCCGGCTTCGAGGAGACATTCGAGGCGTTCGACGATTCCGTGGTTTGTCCTCAACTCACCGACAATGTCGCGGTCGGAAGCCTCCAGTGCTTGAACCTGAACATCTACGTGCCCAACACGGCGACGTCGCGGAATAAGCGCCCGGTGATGATTTGGATCCACGGAGGGGGCTTCATGTCTGGCAGCGGAACTAGCAGAGATTACTCCTTCGAAGATCTGGTCCGACATGACGTTATCATTGTGTCGGTTAACTACCGGCTTGGAGCGTTTGGATTCTTTTGTTTGGACAGCGCCGATGTTCCTGGGAATCAGGGGTTGAAGGACCAAGTTCTTGCTTTAAAATGGATCAAGGAAAATATCGAGGCTTTCGGGGGAGATACCTCTAAGATAACTCTGTTCGGGGAAAGTGCGGGAGGCGTCGCCGTCGAGCTACACCTACTAACCGAGCAAGAGAAGTTCTTCAACCAGGTCATCGTTCAGAGCGCATCCGCCTTTATTCCTGGAGCCATCATGAAACCTGACAACAGTATCCCGATACGAATAGCCAGCCGATTGGGATTCGAAACTGACAACTTCGTAGAAGCTATAGCGTTTTTAGCCGATCAAGACCCGCTCCTGGTGGTCGGAGCGACTAGATCTGATGGCCCGGTCACTTTGACGGACGCCACCTTCGGCTTGTTCAGAGCTTGTATCGAAAATGAATATAATGGAGTTTATAGATTTCTTGCTAATACTCCAGAAAATTTAGACCCTAAAAAGCTTCGCAATATTCCGGTTATCTACGGAATGAACGATCGAGAAATGCTAACTCTGCACGCCTTGATCAGCCCGGAAGAATATGAGCAGCCGGCCATTAGAAACTACATGAATATAGCTTTCGACTTGAAGGAGAACGAGATAGAAGAAGACGTAACTCATTTCTACATCGCAGACGAACCCTTCAATCCGGAGGTCTTTGATAAATTCATTAATTTTGCTTCAAACTACTACTTTCATTATCCCGTCGAGCGGTCCCTTAAGAAGATATTGAACGCGGGCAACGAAGACATATATTACTATGTATTCTCGTACGATGGCGGTCGAAACGCTATGAAAAACCATCTTGGAATAACAGCGCCGGGCGCGGCGCACGCGGACGAGCTCGGCTACCTGCTGGCCGTGGACTTGCAGCCGGGTGAGCTTGTTGCGGAGGAGGATCAACTAGTCATTGACAGAATCACTACGCTTTGGACCAATTTTGCTAAGTTCGGCAATCCGACCCCTGAAACATCTGACCTGCTGCCAGTCGTGTGGTCGCCAGTGACGGCGAGCAGGCGGCCCTTCCTGGAGATCGGCGCCGACTTCGGACTCAAGAGCAGGCCTTTCCACGACCGCATGGCCTTCTGGGATCTCTTCTACAAGCTGTATGGAAAACTGGAACGCATTCACTAA
Protein
MLRPVTLAVVVVALLSAAGARLRVDPLVETRKGLIKGLRSENGEFSKFLGVPYALVDEDNPFGSSVPHPGFEETFEAFDDSVVCPQLTDNVAVGSLQCLNLNIYVPNTATSRNKRPVMIWIHGGGFMSGSGTSRDYSFEDLVRHDVIIVSVNYRLGAFGFFCLDSADVPGNQGLKDQVLALKWIKENIEAFGGDTSKITLFGESAGGVAVELHLLTEQEKFFNQVIVQSASAFIPGAIMKPDNSIPIRIASRLGFETDNFVEAIAFLADQDPLLVVGATRSDGPVTLTDATFGLFRACIENEYNGVYRFLANTPENLDPKKLRNIPVIYGMNDREMLTLHALISPEEYEQPAIRNYMNIAFDLKENEIEEDVTHFYIADEPFNPEVFDKFINFASNYYFHYPVERSLKKILNAGNEDIYYYVFSYDGGRNAMKNHLGITAPGAAHADELGYLLAVDLQPGELVAEEDQLVIDRITTLWTNFAKFGNPTPETSDLLPVVWSPVTASRRPFLEIGADFGLKSRPFHDRMAFWDLFYKLYGKLERIH
Summary
Similarity
Belongs to the type-B carboxylesterase/lipase family.
Feature
chain Carboxylic ester hydrolase
Uniprot
H9J066
H9IZW1
H9J067
H9J064
A0A2H1VCT7
A0A2A4JU91
+ More
A0A2W1BZX4 A0A2H1VCP0 A0A2H1VCN6 H9IWJ7 A0A0N1IPF0 A0A2H1VCN0 A0A194Q254 A0A2H1VCQ9 A0A2A4K075 A0A1U9X1V6 A0A2A4K7E9 A0A194PEF7 A0A2A4JZ33 D5G3E3 A0A194Q887 A0A2H1VEA4 A0A2H1WRP2 A0A212FJ60 B1Q137 A0A2W1BQY3 A0A194Q2Q8 A0A2H1VAJ0 D5G3E0 H9IUA7 H9J1Y0 A0A0N1PJ90 A0A0L7LED6 B1Q141 M1RGF2 A0A437AWV6 A0A0L7L0K6 A0A1U9X1V1 A0A2H1VCR3 A0A2I6RJM2 A0A2A4JR93 A0A194PDI7 H9IWJ8 A0A194Q249 A0A2W1BTR6 A0A194PJ15 A0A0N1IGW1 D5G3E1 D5G3E2 A0A3G1T174 A0A1U9X1V3 A0A2W1BJM9 A0A2H1VCT9 A0A2A4JSH8 A0A194RIH0 A0A2H1WHB4 A0A1U9X1U7 A0A212FJ67 A0A2A4JHL2 B3GEF6 A0A2H1VCP9 A0A2W1BTK5 A0A2W1BXP7 A0A0B4RXS5 A0A2H1VAI9 A0A1U9X1V0 A0A3S2LSF5 A0A2W1BVP4 A0A2A4JU71 A0A212FJ53 A0A0L7L3V0 A0A2H1W648 A0A3S2NSH9 A0A0L7L1R0 A0A1J0M1Z7 D3GDM3 A0A1U9X1Z0 H9IUE5 A0A0C5C1P7 A0A194QYN7 H9JDS3 B1Q139 A4UA26 A0A194PD51 B8YPE7 D5G3E4 A0A0L7KY57 A0A2A4JU07 A0A0L7KQ39 A0A2W1BNW5 A0A0F7QHL9 H9JF44 A0A2H1VH39 A0A1U9X1Z4 A0A2A4JAV9 A0A0C5BXH5 D9ILE0 A0A2H1VJA9 A0A0L7LMD6
A0A2W1BZX4 A0A2H1VCP0 A0A2H1VCN6 H9IWJ7 A0A0N1IPF0 A0A2H1VCN0 A0A194Q254 A0A2H1VCQ9 A0A2A4K075 A0A1U9X1V6 A0A2A4K7E9 A0A194PEF7 A0A2A4JZ33 D5G3E3 A0A194Q887 A0A2H1VEA4 A0A2H1WRP2 A0A212FJ60 B1Q137 A0A2W1BQY3 A0A194Q2Q8 A0A2H1VAJ0 D5G3E0 H9IUA7 H9J1Y0 A0A0N1PJ90 A0A0L7LED6 B1Q141 M1RGF2 A0A437AWV6 A0A0L7L0K6 A0A1U9X1V1 A0A2H1VCR3 A0A2I6RJM2 A0A2A4JR93 A0A194PDI7 H9IWJ8 A0A194Q249 A0A2W1BTR6 A0A194PJ15 A0A0N1IGW1 D5G3E1 D5G3E2 A0A3G1T174 A0A1U9X1V3 A0A2W1BJM9 A0A2H1VCT9 A0A2A4JSH8 A0A194RIH0 A0A2H1WHB4 A0A1U9X1U7 A0A212FJ67 A0A2A4JHL2 B3GEF6 A0A2H1VCP9 A0A2W1BTK5 A0A2W1BXP7 A0A0B4RXS5 A0A2H1VAI9 A0A1U9X1V0 A0A3S2LSF5 A0A2W1BVP4 A0A2A4JU71 A0A212FJ53 A0A0L7L3V0 A0A2H1W648 A0A3S2NSH9 A0A0L7L1R0 A0A1J0M1Z7 D3GDM3 A0A1U9X1Z0 H9IUE5 A0A0C5C1P7 A0A194QYN7 H9JDS3 B1Q139 A4UA26 A0A194PD51 B8YPE7 D5G3E4 A0A0L7KY57 A0A2A4JU07 A0A0L7KQ39 A0A2W1BNW5 A0A0F7QHL9 H9JF44 A0A2H1VH39 A0A1U9X1Z4 A0A2A4JAV9 A0A0C5BXH5 D9ILE0 A0A2H1VJA9 A0A0L7LMD6
EC Number
3.1.1.-
EMBL
BABH01018991
BABH01018992
BABH01019018
BABH01018995
BABH01018984
BABH01018985
+ More
ODYU01001845 SOQ38611.1 SOQ38612.1 NWSH01000660 PCG74962.1 KZ149903 PZC78410.1 SOQ38610.1 SOQ38609.1 BABH01021273 KQ460400 KPJ14974.1 SOQ38608.1 KQ459564 KPI99621.1 SOQ38607.1 NWSH01000356 PCG77070.1 KY021858 AQY62747.1 NWSH01000067 PCG79999.1 KQ459606 KPI91074.1 PCG77069.1 FJ997307 ADF43472.1 KPI99620.1 SOQ38604.1 ODYU01010533 SOQ55708.1 AGBW02008309 OWR53771.1 EU523532 ACB12411.1 KZ150007 PZC75160.1 KPI99618.1 ODYU01001525 SOQ37859.1 FJ997304 ADF43469.1 BABH01001500 BABH01007753 KQ459837 KPJ19757.1 JTDY01001549 KOB73566.1 EU523536 ACB12415.1 KC200270 AGG20205.1 RSAL01000314 RVE42639.1 JTDY01003780 KOB69017.1 KY021842 AQY62731.1 SOQ38605.1 KY304479 AUN86410.1 NWSH01000834 PCG73942.1 KPI91073.1 KPI99616.1 PZC78409.1 KPI91075.1 KPJ19760.1 FJ997305 ADF43470.1 FJ997306 ADF43471.1 MG846884 AXY94736.1 KY021841 AQY62730.1 KZ150132 PZC73036.1 SOQ38606.1 PCG74961.1 KQ460205 KPJ17125.1 ODYU01008680 SOQ52463.1 KY021848 AQY62737.1 OWR53772.1 NWSH01001418 PCG71299.1 EU688968 ACD80422.1 SOQ38613.1 PZC78412.1 PZC78414.1 KJ023212 AIY69040.1 SOQ37858.1 KY021850 AQY62739.1 RSAL01000258 RVE43396.1 PZC78411.1 PCG74959.1 PCG74960.1 OWR53773.1 JTDY01003106 KOB70142.1 ODYU01006598 SOQ48575.1 RVE43394.1 JTDY01003509 KOB69443.1 KU375432 APD13874.1 FJ652455 ACV60239.1 KY021887 AQY62776.1 BABH01001182 KC200271 AGG20206.1 KP001149 AJN91194.1 KQ460947 KPJ10567.1 BABH01003066 EU523534 ACB12413.2 DQ680829 ABH01082.1 KPI91162.1 FJ556877 ACL83460.1 FJ997308 ADF43473.1 JTDY01004572 KOB67996.1 PCG74963.1 JTDY01007420 KOB65210.1 KZ149946 PZC76749.1 LC017770 BAR64784.1 BABH01026894 ODYU01002500 SOQ40071.1 KY021884 AQY62773.1 NWSH01002285 PCG68674.1 KP001158 AJN91203.2 HM150791 ADI47117.1 ODYU01002878 SOQ40908.1 JTDY01000638 KOB76381.1
ODYU01001845 SOQ38611.1 SOQ38612.1 NWSH01000660 PCG74962.1 KZ149903 PZC78410.1 SOQ38610.1 SOQ38609.1 BABH01021273 KQ460400 KPJ14974.1 SOQ38608.1 KQ459564 KPI99621.1 SOQ38607.1 NWSH01000356 PCG77070.1 KY021858 AQY62747.1 NWSH01000067 PCG79999.1 KQ459606 KPI91074.1 PCG77069.1 FJ997307 ADF43472.1 KPI99620.1 SOQ38604.1 ODYU01010533 SOQ55708.1 AGBW02008309 OWR53771.1 EU523532 ACB12411.1 KZ150007 PZC75160.1 KPI99618.1 ODYU01001525 SOQ37859.1 FJ997304 ADF43469.1 BABH01001500 BABH01007753 KQ459837 KPJ19757.1 JTDY01001549 KOB73566.1 EU523536 ACB12415.1 KC200270 AGG20205.1 RSAL01000314 RVE42639.1 JTDY01003780 KOB69017.1 KY021842 AQY62731.1 SOQ38605.1 KY304479 AUN86410.1 NWSH01000834 PCG73942.1 KPI91073.1 KPI99616.1 PZC78409.1 KPI91075.1 KPJ19760.1 FJ997305 ADF43470.1 FJ997306 ADF43471.1 MG846884 AXY94736.1 KY021841 AQY62730.1 KZ150132 PZC73036.1 SOQ38606.1 PCG74961.1 KQ460205 KPJ17125.1 ODYU01008680 SOQ52463.1 KY021848 AQY62737.1 OWR53772.1 NWSH01001418 PCG71299.1 EU688968 ACD80422.1 SOQ38613.1 PZC78412.1 PZC78414.1 KJ023212 AIY69040.1 SOQ37858.1 KY021850 AQY62739.1 RSAL01000258 RVE43396.1 PZC78411.1 PCG74959.1 PCG74960.1 OWR53773.1 JTDY01003106 KOB70142.1 ODYU01006598 SOQ48575.1 RVE43394.1 JTDY01003509 KOB69443.1 KU375432 APD13874.1 FJ652455 ACV60239.1 KY021887 AQY62776.1 BABH01001182 KC200271 AGG20206.1 KP001149 AJN91194.1 KQ460947 KPJ10567.1 BABH01003066 EU523534 ACB12413.2 DQ680829 ABH01082.1 KPI91162.1 FJ556877 ACL83460.1 FJ997308 ADF43473.1 JTDY01004572 KOB67996.1 PCG74963.1 JTDY01007420 KOB65210.1 KZ149946 PZC76749.1 LC017770 BAR64784.1 BABH01026894 ODYU01002500 SOQ40071.1 KY021884 AQY62773.1 NWSH01002285 PCG68674.1 KP001158 AJN91203.2 HM150791 ADI47117.1 ODYU01002878 SOQ40908.1 JTDY01000638 KOB76381.1
Proteomes
PRIDE
Pfam
PF00135 COesterase
Interpro
SUPFAM
SSF53474
SSF53474
Gene 3D
ProteinModelPortal
H9J066
H9IZW1
H9J067
H9J064
A0A2H1VCT7
A0A2A4JU91
+ More
A0A2W1BZX4 A0A2H1VCP0 A0A2H1VCN6 H9IWJ7 A0A0N1IPF0 A0A2H1VCN0 A0A194Q254 A0A2H1VCQ9 A0A2A4K075 A0A1U9X1V6 A0A2A4K7E9 A0A194PEF7 A0A2A4JZ33 D5G3E3 A0A194Q887 A0A2H1VEA4 A0A2H1WRP2 A0A212FJ60 B1Q137 A0A2W1BQY3 A0A194Q2Q8 A0A2H1VAJ0 D5G3E0 H9IUA7 H9J1Y0 A0A0N1PJ90 A0A0L7LED6 B1Q141 M1RGF2 A0A437AWV6 A0A0L7L0K6 A0A1U9X1V1 A0A2H1VCR3 A0A2I6RJM2 A0A2A4JR93 A0A194PDI7 H9IWJ8 A0A194Q249 A0A2W1BTR6 A0A194PJ15 A0A0N1IGW1 D5G3E1 D5G3E2 A0A3G1T174 A0A1U9X1V3 A0A2W1BJM9 A0A2H1VCT9 A0A2A4JSH8 A0A194RIH0 A0A2H1WHB4 A0A1U9X1U7 A0A212FJ67 A0A2A4JHL2 B3GEF6 A0A2H1VCP9 A0A2W1BTK5 A0A2W1BXP7 A0A0B4RXS5 A0A2H1VAI9 A0A1U9X1V0 A0A3S2LSF5 A0A2W1BVP4 A0A2A4JU71 A0A212FJ53 A0A0L7L3V0 A0A2H1W648 A0A3S2NSH9 A0A0L7L1R0 A0A1J0M1Z7 D3GDM3 A0A1U9X1Z0 H9IUE5 A0A0C5C1P7 A0A194QYN7 H9JDS3 B1Q139 A4UA26 A0A194PD51 B8YPE7 D5G3E4 A0A0L7KY57 A0A2A4JU07 A0A0L7KQ39 A0A2W1BNW5 A0A0F7QHL9 H9JF44 A0A2H1VH39 A0A1U9X1Z4 A0A2A4JAV9 A0A0C5BXH5 D9ILE0 A0A2H1VJA9 A0A0L7LMD6
A0A2W1BZX4 A0A2H1VCP0 A0A2H1VCN6 H9IWJ7 A0A0N1IPF0 A0A2H1VCN0 A0A194Q254 A0A2H1VCQ9 A0A2A4K075 A0A1U9X1V6 A0A2A4K7E9 A0A194PEF7 A0A2A4JZ33 D5G3E3 A0A194Q887 A0A2H1VEA4 A0A2H1WRP2 A0A212FJ60 B1Q137 A0A2W1BQY3 A0A194Q2Q8 A0A2H1VAJ0 D5G3E0 H9IUA7 H9J1Y0 A0A0N1PJ90 A0A0L7LED6 B1Q141 M1RGF2 A0A437AWV6 A0A0L7L0K6 A0A1U9X1V1 A0A2H1VCR3 A0A2I6RJM2 A0A2A4JR93 A0A194PDI7 H9IWJ8 A0A194Q249 A0A2W1BTR6 A0A194PJ15 A0A0N1IGW1 D5G3E1 D5G3E2 A0A3G1T174 A0A1U9X1V3 A0A2W1BJM9 A0A2H1VCT9 A0A2A4JSH8 A0A194RIH0 A0A2H1WHB4 A0A1U9X1U7 A0A212FJ67 A0A2A4JHL2 B3GEF6 A0A2H1VCP9 A0A2W1BTK5 A0A2W1BXP7 A0A0B4RXS5 A0A2H1VAI9 A0A1U9X1V0 A0A3S2LSF5 A0A2W1BVP4 A0A2A4JU71 A0A212FJ53 A0A0L7L3V0 A0A2H1W648 A0A3S2NSH9 A0A0L7L1R0 A0A1J0M1Z7 D3GDM3 A0A1U9X1Z0 H9IUE5 A0A0C5C1P7 A0A194QYN7 H9JDS3 B1Q139 A4UA26 A0A194PD51 B8YPE7 D5G3E4 A0A0L7KY57 A0A2A4JU07 A0A0L7KQ39 A0A2W1BNW5 A0A0F7QHL9 H9JF44 A0A2H1VH39 A0A1U9X1Z4 A0A2A4JAV9 A0A0C5BXH5 D9ILE0 A0A2H1VJA9 A0A0L7LMD6
PDB
5YDJ
E-value=8.50898e-46,
Score=464
Ontologies
Topology
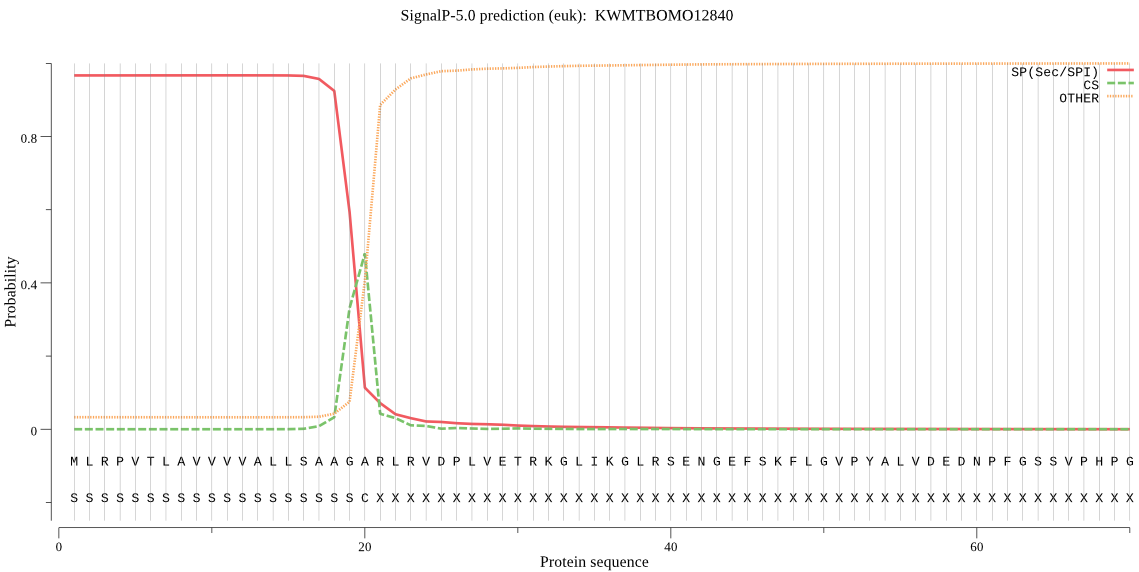
SignalP
Position: 1 - 20,
Likelihood: 0.966649
Length:
544
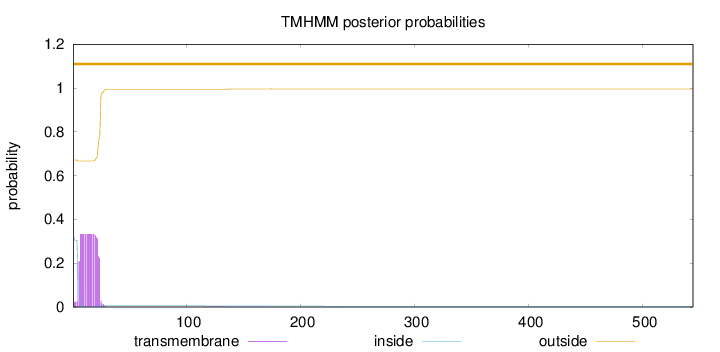
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
6.33693999999999
Exp number, first 60 AAs:
6.29157
Total prob of N-in:
0.32811
outside
1 - 544
Population Genetic Test Statistics
Pi
208.11696
Theta
176.665756
Tajima's D
0.669794
CLR
3.281597
CSRT
0.565671716414179
Interpretation
Uncertain
