Gene
KWMTBOMO12769
Pre Gene Modal
BGIBMGA006410
Annotation
endonuclease-reverse_transcriptase_[Blattella_germanica]
Location in the cell
Nuclear Reliability : 2.243
Sequence
CDS
ATGGAAGCTCAAAGAAAAACCAGACAAGTTATAAGAACAGCCAAGCGAGAGCACATGAATGACATCATGAAGAATATGGAAAACCTGATAAGAACGAATGAGGCTCGAAAATTCTACCAGGAAATAAGATCGGTAAAGAAGGCATACCAACCAATCGCACAATTTCTTGAAGACGAAGACGGAAATCTTGTGACTAACATTCGTGAGATCAGTGTAATGTGGCGTCAGTACTTTCAAAACCTTCTGAATTGCCCTCCACCGACGGAAGATTTAGATGACAACCCCCAAGATCCTGCGGAGGAAGAAGAAGTGGAGCCACTGAACTTCGAAGAAATAAAAAACGCTATTATGAGGCTTAAGAACAACAAAGCGCCAGGTATTGACAACCTACCATCTGAACTGTGGAAATACGGCGGAGAAGCCTTGCAGTCCAGATTGTACAATCTTATTAAAATGATATGGGAATCTGAACAACAACCTGTCCAGTGGTATTCGGGGGTCATCTGCCCCATACACAAAAAAGGCAGCCGGAAGAAATGCAGTAACTACAGAGGAATTGCTTTACTACCTACAGCGTACAAAATACTGTCATACGTTTTACTAAGCAGGTTAGAGCCGTATTCAGACAAAATCATTGGGGATTATCAGTGCGGCTTCAGACGTAATCGTAGTACAGTCGACCAAATCTTTACTCTGAAACAGTTAATGGAGAAAAGATGGGAGTATGCACAAAGTATTCATTCGCTCTTTGTTGATTTTGCTAAGGCCTACGACAGCATAGATAGACCGACTCTATATAGAATCCTCACGCGCTTTAAAATCCCACAGAAACTTGTGAGAATGGTGGAGGTTGCTACTAGGGAAAGTAGAATGACGGTACGAGTTGGCGCTTCTCTCACTGATGAGTTTGACGTTGTGACTGGACTCAGGCAGGGGGACGCTCTGTCACCAATGCTGTTCAACCTGGCACTTGAACACGCCATACGGAAAGTGAACACACTTAGTGGCGGAGTGTGGCTCAACGGACAGCATAGGGTGATAGGATATGCCGATGACCTCGCTCTACTTGGAGAAAGAAAGCAGCACGTTATAGACGCACTCAACTGCCTCCAACAAGAAGCCTCTCGGATTGGCCTTCGCGTGAGCCACGAGAAGACAGAATATCTTCACATGCGTCGATATAAAAACGTAAGACGGAAGCGAGAAGACCTAGTGGTGGGAAATACGAGGTTCAAAGGAGTGGCTAAATTCCGGTACCTTGGCTGCACTGTAACAGACACCAATGACAGGGAAGATGAAATCGAAATTCGTATACAAAATACCCTACGATGCAGTGCAGCCCTACACCCGGTTCTCTCTTCGAAGCTACTAAGCAGGCGCACCAAGATTCGGATATACAAGACTCTAGTCAGACCGATCCTATTGTATGGATGCGAAGCATGGACGCTCACACAGAAAGAAGAGGCAAAACTTCTCGTGACGGAGCGAAAGGTATTGAGAAAAATACTAGGACCAGTTCGGAAAGAAGACGGTTCTTGGCACGTCCGGAAAAATAAGGATGTGGAGAGGCTTTTTGGTGAGCCAAATATTCTGGGGGAGATGAAGGCACGGAGGCTCGGATGGCTCGGACATGTAGTACGTATGGAGCAGGATCGGGCTGTGAAACGGGTGTACACGGGTGTACCGGAGGGACGCCGCCCCGTAGGGAGGCCCAAGTACCGCTGGTGTGACGCTGTGGAAGGAGATTTACGGGAGCTCCAAGTGTCGGACTGGCAAGGTGTTGCGCAGCGCCGGGATGAATGGCAAAGTTTATTGTCGGAGGCCAAGACCCACTTCGGGGTCATAGCGCCAGCTCAGTAG
Protein
MEAQRKTRQVIRTAKREHMNDIMKNMENLIRTNEARKFYQEIRSVKKAYQPIAQFLEDEDGNLVTNIREISVMWRQYFQNLLNCPPPTEDLDDNPQDPAEEEEVEPLNFEEIKNAIMRLKNNKAPGIDNLPSELWKYGGEALQSRLYNLIKMIWESEQQPVQWYSGVICPIHKKGSRKKCSNYRGIALLPTAYKILSYVLLSRLEPYSDKIIGDYQCGFRRNRSTVDQIFTLKQLMEKRWEYAQSIHSLFVDFAKAYDSIDRPTLYRILTRFKIPQKLVRMVEVATRESRMTVRVGASLTDEFDVVTGLRQGDALSPMLFNLALEHAIRKVNTLSGGVWLNGQHRVIGYADDLALLGERKQHVIDALNCLQQEASRIGLRVSHEKTEYLHMRRYKNVRRKREDLVVGNTRFKGVAKFRYLGCTVTDTNDREDEIEIRIQNTLRCSAALHPVLSSKLLSRRTKIRIYKTLVRPILLYGCEAWTLTQKEEAKLLVTERKVLRKILGPVRKEDGSWHVRKNKDVERLFGEPNILGEMKARRLGWLGHVVRMEQDRAVKRVYTGVPEGRRPVGRPKYRWCDAVEGDLRELQVSDWQGVAQRRDEWQSLLSEAKTHFGVIAPAQ
Summary
Uniprot
A0A2P8ZGH9
A0A2P8YPE1
A0A2P8XT91
A0A060Q6A2
A0A2P8ZIH1
A0A2P8ZLT6
+ More
A0A2P8Z488 A0A2P8YWA2 A0A2P8Z313 A0A2P8YLY7 A0A2P8ZFZ0 A0A2P8Y2S1 A0A2P8YLJ1 A0A2P8XR21 A0A1B0DRK3 A0A023F0H0 A0A0K8TRS6 A0A2P8ZCA4 A0A0J7KMY2 A0A2S2NQ16 A0A367YPS2 A0A0U4HM41 A0A2P8YGI5 A0A2M4A3Y0 A0A0K8TJE5 A0A2M4A3Z2 A0A023F0M0 A0A2S2PCU1 A0A2P8YFW0 A0A2P8YYC9 D7F170 A0A2P8ZAY7 A0A023F0F3 A0A2M4A3H0 A0A2M4ACI4 J9L287 A0A146KNZ7 A0A2M4A2X8 A0A2M4A809 A0A2P8Z0K7 A0A2P8XC08 A0A2M4CRR0 J9LFP1 A0A2M4CRT4 A0A3L8DEM2 A0A023EZ68 A0A069DWW0 A0A0P8Y4M6 A0A0V0G616 A0A2P8YTB7 A0A2M4A6U1 A0A0P8Y6K1 J9KNQ8 A0A0V0G8V1 A0A0K8SN56 A0A2S2PT92 A0A2J7Q3Q5 A0A0N8NZU6 A0A0P9A604 X1X8T7 A0A0P8Y5F3 A0A2L2YFD0 A0A2S2N6G4 A0A2S2PIT5 A0A2P8ZEA9 A0A0P8ZWR4 A0A2P8YHZ7 J9KHI9 A0A2S2P2F5 T1E1T8 X1X9T6 A0A0K8S3X2 A0A2P8ZET0 A0A2P8XRQ1 A0A023EZ18 A0A0A9WI21 J9L6X2 A0A2P8Z3H5 A0A023EX67 A0A0P8Y3A6 X1WLM1 J9M6M7 J9KT01 A0A0P8XJ08 A0A023EXX8 A0A2S2N711 A0A0P8Y0P3 X1WT72 A0A0P8XDM7 A0A0P9AMX6 J9KUA2 A0A0P8XIG1 J9KH77 A0A0P9BSL5 A0A0P9AAF4 A0A0P8Y7V9 A0A2S2P3U1 J9LBF3 A0A1Y1LRT6 A0A023EWL3
A0A2P8Z488 A0A2P8YWA2 A0A2P8Z313 A0A2P8YLY7 A0A2P8ZFZ0 A0A2P8Y2S1 A0A2P8YLJ1 A0A2P8XR21 A0A1B0DRK3 A0A023F0H0 A0A0K8TRS6 A0A2P8ZCA4 A0A0J7KMY2 A0A2S2NQ16 A0A367YPS2 A0A0U4HM41 A0A2P8YGI5 A0A2M4A3Y0 A0A0K8TJE5 A0A2M4A3Z2 A0A023F0M0 A0A2S2PCU1 A0A2P8YFW0 A0A2P8YYC9 D7F170 A0A2P8ZAY7 A0A023F0F3 A0A2M4A3H0 A0A2M4ACI4 J9L287 A0A146KNZ7 A0A2M4A2X8 A0A2M4A809 A0A2P8Z0K7 A0A2P8XC08 A0A2M4CRR0 J9LFP1 A0A2M4CRT4 A0A3L8DEM2 A0A023EZ68 A0A069DWW0 A0A0P8Y4M6 A0A0V0G616 A0A2P8YTB7 A0A2M4A6U1 A0A0P8Y6K1 J9KNQ8 A0A0V0G8V1 A0A0K8SN56 A0A2S2PT92 A0A2J7Q3Q5 A0A0N8NZU6 A0A0P9A604 X1X8T7 A0A0P8Y5F3 A0A2L2YFD0 A0A2S2N6G4 A0A2S2PIT5 A0A2P8ZEA9 A0A0P8ZWR4 A0A2P8YHZ7 J9KHI9 A0A2S2P2F5 T1E1T8 X1X9T6 A0A0K8S3X2 A0A2P8ZET0 A0A2P8XRQ1 A0A023EZ18 A0A0A9WI21 J9L6X2 A0A2P8Z3H5 A0A023EX67 A0A0P8Y3A6 X1WLM1 J9M6M7 J9KT01 A0A0P8XJ08 A0A023EXX8 A0A2S2N711 A0A0P8Y0P3 X1WT72 A0A0P8XDM7 A0A0P9AMX6 J9KUA2 A0A0P8XIG1 J9KH77 A0A0P9BSL5 A0A0P9AAF4 A0A0P8Y7V9 A0A2S2P3U1 J9LBF3 A0A1Y1LRT6 A0A023EWL3
Pubmed
EMBL
PYGN01000064
PSN55609.1
PYGN01000451
PSN46127.1
PYGN01001382
PSN35227.1
+ More
HE799691 CCH14900.1 PYGN01000047 PSN56294.1 PYGN01000020 PSN57453.1 PYGN01000203 PSN51319.1 PYGN01000324 PSN48495.1 PYGN01000219 PSN50892.1 PYGN01000502 PSN45243.1 PYGN01000070 PSN55417.1 PYGN01001004 PSN38555.1 PYGN01000510 PSN45119.1 PYGN01001494 PSN34463.1 AJVK01001578 GBBI01004263 JAC14449.1 GDAI01000742 JAI16861.1 PYGN01000107 PSN54137.1 LBMM01005351 KMQ91586.1 GGMR01006433 MBY19052.1 QOUI01000020 RCK67885.1 KU311054 ALX81665.1 PYGN01000613 PSN43351.1 GGFK01002192 MBW35513.1 GBRD01000202 JAG65619.1 GGFK01002196 MBW35517.1 GBBI01004198 JAC14514.1 GGMR01014652 MBY27271.1 PYGN01000629 PSN43121.1 PYGN01000287 PSN49259.1 FJ265555 ADI61823.1 PYGN01000119 PSN53667.1 GBBI01004261 JAC14451.1 GGFK01001949 MBW35270.1 GGFK01005184 MBW38505.1 ABLF02027703 ABLF02061609 GDHC01021627 JAP97001.1 GGFK01001798 MBW35119.1 GGFK01003613 MBW36934.1 PYGN01000254 PSN50032.1 PYGN01003402 PSN29545.1 GGFL01003846 MBW68024.1 ABLF02066124 GGFL01003845 MBW68023.1 QOIP01000009 RLU18885.1 GBBI01004199 JAC14513.1 GBGD01000331 JAC88558.1 CH902619 KPU76535.1 GECL01002632 JAP03492.1 PYGN01000372 PSN47498.1 GGFK01003183 MBW36504.1 CH902653 KPU74858.1 ABLF02039015 GECL01001585 JAP04539.1 GBRD01011073 JAG54751.1 GGMR01020071 MBY32690.1 NEVH01019065 PNF23216.1 CH902657 KPU75308.1 CH902621 KPU81727.1 ABLF02009557 ABLF02016497 ABLF02041313 ABLF02041567 ABLF02062088 CH904714 KPU81814.1 IAAA01017079 LAA06070.1 GGMR01000126 MBY12745.1 GGMR01016675 MBY29294.1 PYGN01000082 PSN54842.1 CH902626 KPU79149.1 PYGN01000580 PSN43876.1 ABLF02019047 ABLF02019049 GGMR01010991 MBY23610.1 GALA01001775 JAA93077.1 ABLF02036907 ABLF02036915 ABLF02036922 ABLF02041419 GBRD01017846 JAG47981.1 PYGN01000078 PSN55011.1 PYGN01001465 PSN34679.1 GBBI01004262 JAC14450.1 GBHO01037471 GBRD01009355 GBRD01009354 GDHC01014856 GDHC01007898 JAG06133.1 JAG56469.1 JAQ03773.1 JAQ10731.1 ABLF02018097 ABLF02043469 ABLF02056908 PYGN01000212 PSN51056.1 GAPW01000107 JAC13491.1 CH902714 KPU81861.1 ABLF02010115 ABLF02030763 ABLF02005950 ABLF02005951 ABLF02003651 ABLF02007294 ABLF02007295 ABLF02056183 CH902627 KPU74872.1 GAPW01000199 JAC13399.1 GGMR01000301 MBY12920.1 CH902628 KPU75346.1 ABLF02020712 ABLF02020717 ABLF02054820 CH902623 KPU72899.1 CH902618 KPU79125.1 ABLF02026145 ABLF02026146 ABLF02042729 CH902666 KPU74610.1 ABLF02018925 CH902633 KPU74667.1 CH902622 KPU75266.1 KPU77326.1 CH902630 KPU77522.1 GGMR01011466 MBY24085.1 ABLF02003877 ABLF02042991 GEZM01048836 JAV76363.1 GAPW01000120 JAC13478.1
HE799691 CCH14900.1 PYGN01000047 PSN56294.1 PYGN01000020 PSN57453.1 PYGN01000203 PSN51319.1 PYGN01000324 PSN48495.1 PYGN01000219 PSN50892.1 PYGN01000502 PSN45243.1 PYGN01000070 PSN55417.1 PYGN01001004 PSN38555.1 PYGN01000510 PSN45119.1 PYGN01001494 PSN34463.1 AJVK01001578 GBBI01004263 JAC14449.1 GDAI01000742 JAI16861.1 PYGN01000107 PSN54137.1 LBMM01005351 KMQ91586.1 GGMR01006433 MBY19052.1 QOUI01000020 RCK67885.1 KU311054 ALX81665.1 PYGN01000613 PSN43351.1 GGFK01002192 MBW35513.1 GBRD01000202 JAG65619.1 GGFK01002196 MBW35517.1 GBBI01004198 JAC14514.1 GGMR01014652 MBY27271.1 PYGN01000629 PSN43121.1 PYGN01000287 PSN49259.1 FJ265555 ADI61823.1 PYGN01000119 PSN53667.1 GBBI01004261 JAC14451.1 GGFK01001949 MBW35270.1 GGFK01005184 MBW38505.1 ABLF02027703 ABLF02061609 GDHC01021627 JAP97001.1 GGFK01001798 MBW35119.1 GGFK01003613 MBW36934.1 PYGN01000254 PSN50032.1 PYGN01003402 PSN29545.1 GGFL01003846 MBW68024.1 ABLF02066124 GGFL01003845 MBW68023.1 QOIP01000009 RLU18885.1 GBBI01004199 JAC14513.1 GBGD01000331 JAC88558.1 CH902619 KPU76535.1 GECL01002632 JAP03492.1 PYGN01000372 PSN47498.1 GGFK01003183 MBW36504.1 CH902653 KPU74858.1 ABLF02039015 GECL01001585 JAP04539.1 GBRD01011073 JAG54751.1 GGMR01020071 MBY32690.1 NEVH01019065 PNF23216.1 CH902657 KPU75308.1 CH902621 KPU81727.1 ABLF02009557 ABLF02016497 ABLF02041313 ABLF02041567 ABLF02062088 CH904714 KPU81814.1 IAAA01017079 LAA06070.1 GGMR01000126 MBY12745.1 GGMR01016675 MBY29294.1 PYGN01000082 PSN54842.1 CH902626 KPU79149.1 PYGN01000580 PSN43876.1 ABLF02019047 ABLF02019049 GGMR01010991 MBY23610.1 GALA01001775 JAA93077.1 ABLF02036907 ABLF02036915 ABLF02036922 ABLF02041419 GBRD01017846 JAG47981.1 PYGN01000078 PSN55011.1 PYGN01001465 PSN34679.1 GBBI01004262 JAC14450.1 GBHO01037471 GBRD01009355 GBRD01009354 GDHC01014856 GDHC01007898 JAG06133.1 JAG56469.1 JAQ03773.1 JAQ10731.1 ABLF02018097 ABLF02043469 ABLF02056908 PYGN01000212 PSN51056.1 GAPW01000107 JAC13491.1 CH902714 KPU81861.1 ABLF02010115 ABLF02030763 ABLF02005950 ABLF02005951 ABLF02003651 ABLF02007294 ABLF02007295 ABLF02056183 CH902627 KPU74872.1 GAPW01000199 JAC13399.1 GGMR01000301 MBY12920.1 CH902628 KPU75346.1 ABLF02020712 ABLF02020717 ABLF02054820 CH902623 KPU72899.1 CH902618 KPU79125.1 ABLF02026145 ABLF02026146 ABLF02042729 CH902666 KPU74610.1 ABLF02018925 CH902633 KPU74667.1 CH902622 KPU75266.1 KPU77326.1 CH902630 KPU77522.1 GGMR01011466 MBY24085.1 ABLF02003877 ABLF02042991 GEZM01048836 JAV76363.1 GAPW01000120 JAC13478.1
Proteomes
PRIDE
Pfam
Interpro
Gene 3D
ProteinModelPortal
A0A2P8ZGH9
A0A2P8YPE1
A0A2P8XT91
A0A060Q6A2
A0A2P8ZIH1
A0A2P8ZLT6
+ More
A0A2P8Z488 A0A2P8YWA2 A0A2P8Z313 A0A2P8YLY7 A0A2P8ZFZ0 A0A2P8Y2S1 A0A2P8YLJ1 A0A2P8XR21 A0A1B0DRK3 A0A023F0H0 A0A0K8TRS6 A0A2P8ZCA4 A0A0J7KMY2 A0A2S2NQ16 A0A367YPS2 A0A0U4HM41 A0A2P8YGI5 A0A2M4A3Y0 A0A0K8TJE5 A0A2M4A3Z2 A0A023F0M0 A0A2S2PCU1 A0A2P8YFW0 A0A2P8YYC9 D7F170 A0A2P8ZAY7 A0A023F0F3 A0A2M4A3H0 A0A2M4ACI4 J9L287 A0A146KNZ7 A0A2M4A2X8 A0A2M4A809 A0A2P8Z0K7 A0A2P8XC08 A0A2M4CRR0 J9LFP1 A0A2M4CRT4 A0A3L8DEM2 A0A023EZ68 A0A069DWW0 A0A0P8Y4M6 A0A0V0G616 A0A2P8YTB7 A0A2M4A6U1 A0A0P8Y6K1 J9KNQ8 A0A0V0G8V1 A0A0K8SN56 A0A2S2PT92 A0A2J7Q3Q5 A0A0N8NZU6 A0A0P9A604 X1X8T7 A0A0P8Y5F3 A0A2L2YFD0 A0A2S2N6G4 A0A2S2PIT5 A0A2P8ZEA9 A0A0P8ZWR4 A0A2P8YHZ7 J9KHI9 A0A2S2P2F5 T1E1T8 X1X9T6 A0A0K8S3X2 A0A2P8ZET0 A0A2P8XRQ1 A0A023EZ18 A0A0A9WI21 J9L6X2 A0A2P8Z3H5 A0A023EX67 A0A0P8Y3A6 X1WLM1 J9M6M7 J9KT01 A0A0P8XJ08 A0A023EXX8 A0A2S2N711 A0A0P8Y0P3 X1WT72 A0A0P8XDM7 A0A0P9AMX6 J9KUA2 A0A0P8XIG1 J9KH77 A0A0P9BSL5 A0A0P9AAF4 A0A0P8Y7V9 A0A2S2P3U1 J9LBF3 A0A1Y1LRT6 A0A023EWL3
A0A2P8Z488 A0A2P8YWA2 A0A2P8Z313 A0A2P8YLY7 A0A2P8ZFZ0 A0A2P8Y2S1 A0A2P8YLJ1 A0A2P8XR21 A0A1B0DRK3 A0A023F0H0 A0A0K8TRS6 A0A2P8ZCA4 A0A0J7KMY2 A0A2S2NQ16 A0A367YPS2 A0A0U4HM41 A0A2P8YGI5 A0A2M4A3Y0 A0A0K8TJE5 A0A2M4A3Z2 A0A023F0M0 A0A2S2PCU1 A0A2P8YFW0 A0A2P8YYC9 D7F170 A0A2P8ZAY7 A0A023F0F3 A0A2M4A3H0 A0A2M4ACI4 J9L287 A0A146KNZ7 A0A2M4A2X8 A0A2M4A809 A0A2P8Z0K7 A0A2P8XC08 A0A2M4CRR0 J9LFP1 A0A2M4CRT4 A0A3L8DEM2 A0A023EZ68 A0A069DWW0 A0A0P8Y4M6 A0A0V0G616 A0A2P8YTB7 A0A2M4A6U1 A0A0P8Y6K1 J9KNQ8 A0A0V0G8V1 A0A0K8SN56 A0A2S2PT92 A0A2J7Q3Q5 A0A0N8NZU6 A0A0P9A604 X1X8T7 A0A0P8Y5F3 A0A2L2YFD0 A0A2S2N6G4 A0A2S2PIT5 A0A2P8ZEA9 A0A0P8ZWR4 A0A2P8YHZ7 J9KHI9 A0A2S2P2F5 T1E1T8 X1X9T6 A0A0K8S3X2 A0A2P8ZET0 A0A2P8XRQ1 A0A023EZ18 A0A0A9WI21 J9L6X2 A0A2P8Z3H5 A0A023EX67 A0A0P8Y3A6 X1WLM1 J9M6M7 J9KT01 A0A0P8XJ08 A0A023EXX8 A0A2S2N711 A0A0P8Y0P3 X1WT72 A0A0P8XDM7 A0A0P9AMX6 J9KUA2 A0A0P8XIG1 J9KH77 A0A0P9BSL5 A0A0P9AAF4 A0A0P8Y7V9 A0A2S2P3U1 J9LBF3 A0A1Y1LRT6 A0A023EWL3
Ontologies
GO
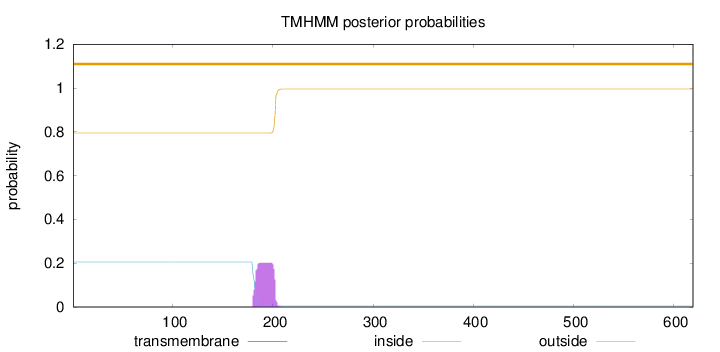
Topology
Length:
619
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
4.16481
Exp number, first 60 AAs:
0
Total prob of N-in:
0.20572
outside
1 - 619
Population Genetic Test Statistics
Pi
15.527612
Theta
18.874603
Tajima's D
-0.660436
CLR
3.955404
CSRT
0.2024398780061
Interpretation
Uncertain
