Gene
KWMTBOMO12443 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA002365
Annotation
Endoprotease_FURIN_[Spodoptera_frugiperda]
Full name
Furin-like protease 2
Location in the cell
Extracellular Reliability : 3.398
Sequence
CDS
ATGAACGTGGCGCCAGCATGGCAAAAGGGATATACAGGCAAGGGCGTTGTAGTGTCAATATTAGATGACGGAATACAAACAAACCATCCAGATTTAGCACAGAATTACGATCCTCTAGCATCGACGGACATAAACGGCAACGATGATGACCCTATGCCACAGGACAATGGCGACAATAAACACGGCACAAGGTGTGCAGGCGAAGTCGCAGCTGTCGCTTACAATCGCTACTGTGGTGTGGGTATTGCCTACAACGCCAGCATAGGAGGAGTTAGGATGTTAGATGGAATTGTCAATGATGCAGTAGAAGCAAGAGCGTTGGGATTGAATCCAGATCATATTGACATCTATAGCGCTTCCTGGGGACCGGAAGACGATGGCAAAACAGTTGACGGTCCTGGACCTTTGGCCAGAAGAGCATTTATATATGGTGTTACGAGCGGTCGTCGTGGCAAAGGCTCTATCTTTGTCTGGGCTTCCGGTAACGGCGGAAGGCACACCGATTCATGTAACTGTGACGGCTACACAAACAGCATATTTACTCTATCGATATCTAGTGCAACACAAGGAGGCTACAAACCTTGGTATCTCGAAGAATGTTCTTCCACTTTAGCTACCACTTACAGTTCGGGTACTCCGGGTCACGACAAAAGTGTGGCGACTGTTGATATGGACGGTAGACTGAGGGCTGACCACATTTGTACAGTAGAACACACGGGAACTTCAGCTTCTGCACCATTAGCAGCTGGAATATGTGCTTTAGCTCTTGAAGCAAATCCTGAATTAACATGGAGAGATATGCAATATCTGGTTGTAATGACGTCTAGGCCTGAACCGTTAGAAAGAGAAAAGGGTTGGATCATCAATGGAGTCAAGAGGAAAGTTAGTCACAAATTCGGATACGGTCTTATGGATGCCACGGAAATGGTAAATTTGGCCGAGCAGTGGGTAACGGTTCCACCTCAACATATCTGTAAATCGCAGGAGTTGAACGAAGACAAGTCGATGGACTCTACTTTGGGCAATTCATTAAGGACATATATGGATGTAACTGGATGCAGTGGTACGGTGAATGAAGTGCGTTACCTAGAACATGTCCAATGTAAAATATCGTTGAGGTTCTTCCCCCGTGGTAACCTGCGTATATTATTAACTTCACCAATGGGTACTGTATCAACATTATTGTTCGAACGACCTCGTGACGTGGCTAGTTCTACCTTCGACGATTGGCCATTTTTGAGCGTTCATTTTTGGGGAGAACAAGCAGAAGGTAGATGGACATTAGAAATAATAAACGCCGGCAAAAGGCACGTGAACCAGGCGGGCGTTCTCAAGAAATGGCAGCTGATATTCTACGGAACTGTCATAGATCCCGTTCGGTTACGATCAAAGAGACCGTCGCCATTTGCACCGTCATACGCGTTTCCAACGGCAGCGGATGGCTACGACACAATCAGTGATTCTTTTTACAATACTGACGCGTTTGCTAATTACCAGAACTTTCCAAATTTATTCGCCGCTGGGTCCAACCCGGAAAAGGCGGTAGCCCGTCTCGACGGACATAACGTCCCTTCACCGCATGGGGAAAATGTCCTCGCCGATAGTAATGACAAGCGCGTCATGCACGACTGTGATCCCGAATGCGATTCTCAAGGATGCTACGGAAAAGGACCGTCCCAATGTGTGGCTTGTAAACATTACCGTTTAGATGACACGTGCGTTTCTCGTTGCCCACCGAGAAGCTATGCCAATCAAGGGGGAGTTTGTTGGCAATGTCACGAATCGTGTGAAACTTGCATGGGACCGGGGCAAGATTCATGTTTCACATGTGCTCCGGCTCATTTTTTAGTGGCAGATTTGGCTGTTTGCCTACAACAATGTCCTGATGGATATTGGGAGGACACAGATTCTGCAACTTGTCGGCCTTGTGCAGCTCATTGTGCGACTTGTTCGGAAAGAGCGAATGCGTGCACGTCGTGTGAACATCACCTACTCCTGCACGAGGGCAGCTGTCTGGTCACATGTCCGGCAGCGCACTACGAAACCGACGATTACACTTGCGCCAAATGCCACAATTCGTGTGACACATGCGTTGGACCAAAGGAAAACCAATGCATAACTTGTCACACTACAAATTACGTGTTAGATGGTAGCTGTGTGGCGACATGTCCAGGCGGATATTACGTGGACAAAAAAAGGAAGGAATGCGTGAGGTGTCCAGTTGGATGTGCGACTTGCACAAGCGCTTTCTGTCTCACATGCGAAACAAATTGGGAAATAAATAAGAAGGGGAGATGTCTCCCAGCTGGATCTGATCGATGTTCGACCGGACAATTCATAGATAAACTCGGCTGTAGCAGATGTGACGACGCTTGTGAATCGTGCTATGGAGAGGGCGAAGGACACTGTCTAACTTGTCCATCGCCTAATTTGCTAGAAGATTACAGATGCGTTCCGGAATGTAGCAGCGGATATTACGCGGAGGCCGGTCGGTGCATTCGCTGCGTGCACGGTTGTACTGAATGCGTTTCAAGACTCAACTGTACCAGCTGTGCTGGGTCCCTGAGACTGCAGTCTGGAGCTTGCAGGACTTCATGCGCTGATGGGTACTACGCTGATCGCGGCGCCTGTTCCAAGTGCTATTTATCTTGCCGGACATGTATTGGGCCCCGTCGCGATCAGTGCGCGTCCTGCCCTCAGGGTTGGAGGCTAGCCGCTGGAGAATGTCACCCCGAATGTCCTCAAGGATTCTACAAAAGCGACGACGGCTGCCGTCATTGTCACCACTACTGCCGGGAGTGCTCGGACTCCGGTCCGCTACACTGCACGTCGTGTCCGCCTCGTTTCGTGTTAGACGGCGGGCTTTGTATGGAGTGCCTCGGCTCTCAGTACTACGAGGCGGGCAATGGAACGTGCAGCACGTGCGATCCGTCGTGTCGCACCTGCTACGGTTCGGGACAGTTCAGTTGTACAGGATGCTCCAGGCCGCTGAGGCTCGATAGATTAAACAATCAGTGCGTTCCGTGTTGTACGGACAGGGGGACGACGACCTCGCCCGGGACCGAATGCTGCCATTGTCATCCCGAAACTGGTGAATGCATTAACTCATCGGTGGCGGGAAAACGTCGTATAGCCGAATGGGGCGCTCTGCACACAGTGGAGAACGCTCGCAGTACGTCAGCTCCCAGCGCCGCCGCGATAACCGTCATGGTGTGTCTGTCCGCCGGTGCACTGTTCTTTGTCGTGCTGATGATACTACAGTGGCGTTCACCGCGCGACAAGCGAACACGAACCCGCGGGCTCGTCTACTACCCCCTCGCCTACAACGACGACGGCAAGGTTGTCGTCCTTGCTCCGCATACGGCGTTCCCCTCAAGCGAGAGCAGGACGGAGGAACTACACGAGCACGAAGACGAACCGTTGCTCGAACATTCCACATAG
Protein
MNVAPAWQKGYTGKGVVVSILDDGIQTNHPDLAQNYDPLASTDINGNDDDPMPQDNGDNKHGTRCAGEVAAVAYNRYCGVGIAYNASIGGVRMLDGIVNDAVEARALGLNPDHIDIYSASWGPEDDGKTVDGPGPLARRAFIYGVTSGRRGKGSIFVWASGNGGRHTDSCNCDGYTNSIFTLSISSATQGGYKPWYLEECSSTLATTYSSGTPGHDKSVATVDMDGRLRADHICTVEHTGTSASAPLAAGICALALEANPELTWRDMQYLVVMTSRPEPLEREKGWIINGVKRKVSHKFGYGLMDATEMVNLAEQWVTVPPQHICKSQELNEDKSMDSTLGNSLRTYMDVTGCSGTVNEVRYLEHVQCKISLRFFPRGNLRILLTSPMGTVSTLLFERPRDVASSTFDDWPFLSVHFWGEQAEGRWTLEIINAGKRHVNQAGVLKKWQLIFYGTVIDPVRLRSKRPSPFAPSYAFPTAADGYDTISDSFYNTDAFANYQNFPNLFAAGSNPEKAVARLDGHNVPSPHGENVLADSNDKRVMHDCDPECDSQGCYGKGPSQCVACKHYRLDDTCVSRCPPRSYANQGGVCWQCHESCETCMGPGQDSCFTCAPAHFLVADLAVCLQQCPDGYWEDTDSATCRPCAAHCATCSERANACTSCEHHLLLHEGSCLVTCPAAHYETDDYTCAKCHNSCDTCVGPKENQCITCHTTNYVLDGSCVATCPGGYYVDKKRKECVRCPVGCATCTSAFCLTCETNWEINKKGRCLPAGSDRCSTGQFIDKLGCSRCDDACESCYGEGEGHCLTCPSPNLLEDYRCVPECSSGYYAEAGRCIRCVHGCTECVSRLNCTSCAGSLRLQSGACRTSCADGYYADRGACSKCYLSCRTCIGPRRDQCASCPQGWRLAAGECHPECPQGFYKSDDGCRHCHHYCRECSDSGPLHCTSCPPRFVLDGGLCMECLGSQYYEAGNGTCSTCDPSCRTCYGSGQFSCTGCSRPLRLDRLNNQCVPCCTDRGTTTSPGTECCHCHPETGECINSSVAGKRRIAEWGALHTVENARSTSAPSAAAITVMVCLSAGALFFVVLMILQWRSPRDKRTRTRGLVYYPLAYNDDGKVVVLAPHTAFPSSESRTEELHEHEDEPLLEHST
Summary
Description
Furin is likely to represent the ubiquitous endoprotease activity within constitutive secretory pathways and capable of cleavage at the RX(K/R)R consensus motif.
Catalytic Activity
Release of mature proteins from their proproteins by cleavage of -Arg-Xaa-Yaa-Arg-|-Zaa- bonds, where Xaa can be any amino acid and Yaa is Arg or Lys. Releases albumin, complement component C3 and von Willebrand factor from their respective precursors.
Cofactor
Ca(2+)
Similarity
Belongs to the peptidase S8 family.
Belongs to the peptidase S8 family. Furin subfamily.
Belongs to the peptidase S8 family. Furin subfamily.
Keywords
Alternative splicing
Cleavage on pair of basic residues
Complete proteome
Disulfide bond
Glycoprotein
Hydrolase
Membrane
Protease
Reference proteome
Repeat
Serine protease
Signal
Transmembrane
Transmembrane helix
Zymogen
Feature
chain Furin-like protease 2
splice variant In isoform A.
splice variant In isoform A.
Uniprot
H9IYN1
Q26489
A0A2A4K7Y7
A0A437BM12
A0A194PYD1
A0A212F4A7
+ More
A0A194RFY5 A0A0A1XGS4 A0A0A1X1M3 A0A0K8U8C6 W8APW3 A0A0L0BQD1 T1P9R1 A0A1I8Q1I4 A0A1Q3FU12 A0A1Q3FTV5 A0A1Q3FUD1 A0A1Q3FTU0 A0A1B6GHW2 W4VRF9 A0A2M4DS50 A0A2M3Z0N1 A0A2M4DS69 A0A2M4A589 A0A2M4B9D5 A0A2M4B9F8 A0A2M4B9B2 A0A2M4A5Q5 A0A2M4A4R7 A0A1B6CCM9 A0A2H1W496 A0A0L7LUE0 A0A2C9GWR4 A0A1J1IF38 E0V9N9 A0A3B0K7Y0 E1JJL4 A0A0R3P116 Q29GI8 B4NCL2 A0A0R3P6H0 P30432 P30432-2 A0A0R3P5B9 B4M1S6 A0A0Q9WMJ9 A0A158NEY8 B4PXS5 A0A1I8Q1H9 B3NX47 A0A0R1EGT9 A0A0R1EEA9 A0A0R1EG80 A0A195FL77 A0A0Q5T532 A0A0Q5T754 A0A437B8F0 B4JKM2 B4R5T4 A0A151IEG1 A0A336LIK8 A0A2A4JGI8 A0A336LJ86 A0A2H1WK85 A0A212FLJ6 A0A2M4CJY4 A0A195BWS1 A0A026X0H0 B4L8R4 A0A0P8ZKG0 A0A0Q9XGD4 A0A0Q9XJA6 B3N0S2 A0A0Q9XRF1 A0A0M4ELX6 E2BLN8 A0A0C9QQN8 A0A194PZ75 A0A1W4VVH0 A0A1W4VTY2 A0A1W4VUQ9 A0A1W4VHD4 E2AVN1 A0A2M4CHY2 A0A2M4CI92 A0A336LJV0 F4WVD8 A0A0C9QJY7 A0A151XIV9 W5JF16 E9ID68 A0A195EN83 A0A2A3EGB8 A0A2J7QX97 A0A182QSE6 A0A182XHH2 A0A067RUA5 A0A0N0U668 A0A2H8TUQ2
A0A194RFY5 A0A0A1XGS4 A0A0A1X1M3 A0A0K8U8C6 W8APW3 A0A0L0BQD1 T1P9R1 A0A1I8Q1I4 A0A1Q3FU12 A0A1Q3FTV5 A0A1Q3FUD1 A0A1Q3FTU0 A0A1B6GHW2 W4VRF9 A0A2M4DS50 A0A2M3Z0N1 A0A2M4DS69 A0A2M4A589 A0A2M4B9D5 A0A2M4B9F8 A0A2M4B9B2 A0A2M4A5Q5 A0A2M4A4R7 A0A1B6CCM9 A0A2H1W496 A0A0L7LUE0 A0A2C9GWR4 A0A1J1IF38 E0V9N9 A0A3B0K7Y0 E1JJL4 A0A0R3P116 Q29GI8 B4NCL2 A0A0R3P6H0 P30432 P30432-2 A0A0R3P5B9 B4M1S6 A0A0Q9WMJ9 A0A158NEY8 B4PXS5 A0A1I8Q1H9 B3NX47 A0A0R1EGT9 A0A0R1EEA9 A0A0R1EG80 A0A195FL77 A0A0Q5T532 A0A0Q5T754 A0A437B8F0 B4JKM2 B4R5T4 A0A151IEG1 A0A336LIK8 A0A2A4JGI8 A0A336LJ86 A0A2H1WK85 A0A212FLJ6 A0A2M4CJY4 A0A195BWS1 A0A026X0H0 B4L8R4 A0A0P8ZKG0 A0A0Q9XGD4 A0A0Q9XJA6 B3N0S2 A0A0Q9XRF1 A0A0M4ELX6 E2BLN8 A0A0C9QQN8 A0A194PZ75 A0A1W4VVH0 A0A1W4VTY2 A0A1W4VUQ9 A0A1W4VHD4 E2AVN1 A0A2M4CHY2 A0A2M4CI92 A0A336LJV0 F4WVD8 A0A0C9QJY7 A0A151XIV9 W5JF16 E9ID68 A0A195EN83 A0A2A3EGB8 A0A2J7QX97 A0A182QSE6 A0A182XHH2 A0A067RUA5 A0A0N0U668 A0A2H8TUQ2
EC Number
3.4.21.75
Pubmed
EMBL
BABH01037900
Z68888
CAA93116.1
NWSH01000051
PCG80179.1
RSAL01000033
+ More
RVE51494.1 KQ459585 KPI98346.1 AGBW02010411 OWR48567.1 KQ460211 KPJ16512.1 GBXI01004172 JAD10120.1 GBXI01009632 JAD04660.1 GDHF01029508 JAI22806.1 GAMC01018483 JAB88072.1 JRES01001522 KNC22262.1 KA645434 AFP60063.1 GFDL01004017 JAV31028.1 GFDL01004015 JAV31030.1 GFDL01004012 JAV31033.1 GFDL01004014 JAV31031.1 GECZ01007763 JAS62006.1 GANO01003419 JAB56452.1 GGFL01016196 MBW80374.1 GGFM01001305 MBW22056.1 GGFL01016237 MBW80415.1 GGFK01002655 MBW35976.1 GGFJ01000508 MBW49649.1 GGFJ01000501 MBW49642.1 GGFJ01000503 MBW49644.1 GGFK01002802 MBW36123.1 GGFK01002391 MBW35712.1 GEDC01026106 JAS11192.1 ODYU01006244 SOQ47911.1 JTDY01000105 KOB78841.1 CVRI01000047 CRK97614.1 DS234995 EEB10108.1 OUUW01000015 SPP88772.1 AE014298 ACZ95309.1 CH379064 KRT06942.1 EAL32121.2 CH964239 EDW82571.2 KRT06944.1 KRT06945.1 M94375 L33831 BT021414 AY070553 KRT06943.1 CH940651 EDW65630.1 KRF82320.1 KRF82319.1 ADTU01013412 ADTU01013413 CM000162 EDX01911.1 CH954180 EDV46876.2 KRK06434.1 KRK06436.1 KRK06435.1 KRK06433.1 KQ981490 KYN41168.1 KQS30223.1 KQS30224.1 KQS30225.1 RSAL01000126 RVE46569.1 CH916370 EDW00125.1 CM000366 EDX18084.1 KQ977871 KYM99195.1 UFQT01000021 SSX17944.1 NWSH01001572 PCG70828.1 SSX17946.1 ODYU01009227 SOQ53495.1 AGBW02007756 OWR54611.1 GGFL01001478 MBW65656.1 KQ976401 KYM92418.1 KK107064 EZA60894.1 CH933815 EDW08039.1 CH902644 KPU75290.1 KRG07562.1 KRG07564.1 KRG07565.1 KRG07567.1 KRG07561.1 EDV34801.1 KRG07566.1 CP012528 ALC48584.1 GL449036 EFN83372.1 GBYB01002977 JAG72744.1 KPI98343.1 GL443146 EFN62505.1 GGFL01000782 MBW64960.1 GGFL01000777 MBW64955.1 SSX17945.1 GL888387 EGI61759.1 GBYB01003844 JAG73611.1 KQ982080 KYQ60346.1 ADMH02001671 ETN61480.1 GL762454 EFZ21411.1 KQ978625 KYN29621.1 KZ288254 PBC30823.1 NEVH01009384 PNF33201.1 AXCN02000011 KK852463 KDR23424.1 KQ435735 KOX77124.1 GFXV01006180 MBW17985.1
RVE51494.1 KQ459585 KPI98346.1 AGBW02010411 OWR48567.1 KQ460211 KPJ16512.1 GBXI01004172 JAD10120.1 GBXI01009632 JAD04660.1 GDHF01029508 JAI22806.1 GAMC01018483 JAB88072.1 JRES01001522 KNC22262.1 KA645434 AFP60063.1 GFDL01004017 JAV31028.1 GFDL01004015 JAV31030.1 GFDL01004012 JAV31033.1 GFDL01004014 JAV31031.1 GECZ01007763 JAS62006.1 GANO01003419 JAB56452.1 GGFL01016196 MBW80374.1 GGFM01001305 MBW22056.1 GGFL01016237 MBW80415.1 GGFK01002655 MBW35976.1 GGFJ01000508 MBW49649.1 GGFJ01000501 MBW49642.1 GGFJ01000503 MBW49644.1 GGFK01002802 MBW36123.1 GGFK01002391 MBW35712.1 GEDC01026106 JAS11192.1 ODYU01006244 SOQ47911.1 JTDY01000105 KOB78841.1 CVRI01000047 CRK97614.1 DS234995 EEB10108.1 OUUW01000015 SPP88772.1 AE014298 ACZ95309.1 CH379064 KRT06942.1 EAL32121.2 CH964239 EDW82571.2 KRT06944.1 KRT06945.1 M94375 L33831 BT021414 AY070553 KRT06943.1 CH940651 EDW65630.1 KRF82320.1 KRF82319.1 ADTU01013412 ADTU01013413 CM000162 EDX01911.1 CH954180 EDV46876.2 KRK06434.1 KRK06436.1 KRK06435.1 KRK06433.1 KQ981490 KYN41168.1 KQS30223.1 KQS30224.1 KQS30225.1 RSAL01000126 RVE46569.1 CH916370 EDW00125.1 CM000366 EDX18084.1 KQ977871 KYM99195.1 UFQT01000021 SSX17944.1 NWSH01001572 PCG70828.1 SSX17946.1 ODYU01009227 SOQ53495.1 AGBW02007756 OWR54611.1 GGFL01001478 MBW65656.1 KQ976401 KYM92418.1 KK107064 EZA60894.1 CH933815 EDW08039.1 CH902644 KPU75290.1 KRG07562.1 KRG07564.1 KRG07565.1 KRG07567.1 KRG07561.1 EDV34801.1 KRG07566.1 CP012528 ALC48584.1 GL449036 EFN83372.1 GBYB01002977 JAG72744.1 KPI98343.1 GL443146 EFN62505.1 GGFL01000782 MBW64960.1 GGFL01000777 MBW64955.1 SSX17945.1 GL888387 EGI61759.1 GBYB01003844 JAG73611.1 KQ982080 KYQ60346.1 ADMH02001671 ETN61480.1 GL762454 EFZ21411.1 KQ978625 KYN29621.1 KZ288254 PBC30823.1 NEVH01009384 PNF33201.1 AXCN02000011 KK852463 KDR23424.1 KQ435735 KOX77124.1 GFXV01006180 MBW17985.1
Proteomes
UP000005204
UP000218220
UP000283053
UP000053268
UP000007151
UP000053240
+ More
UP000037069 UP000095300 UP000037510 UP000075900 UP000183832 UP000009046 UP000268350 UP000000803 UP000001819 UP000007798 UP000008792 UP000005205 UP000002282 UP000008711 UP000078541 UP000001070 UP000000304 UP000078542 UP000078540 UP000053097 UP000009192 UP000007801 UP000092553 UP000008237 UP000192221 UP000000311 UP000007755 UP000075809 UP000000673 UP000078492 UP000242457 UP000235965 UP000075886 UP000076407 UP000027135 UP000053105
UP000037069 UP000095300 UP000037510 UP000075900 UP000183832 UP000009046 UP000268350 UP000000803 UP000001819 UP000007798 UP000008792 UP000005205 UP000002282 UP000008711 UP000078541 UP000001070 UP000000304 UP000078542 UP000078540 UP000053097 UP000009192 UP000007801 UP000092553 UP000008237 UP000192221 UP000000311 UP000007755 UP000075809 UP000000673 UP000078492 UP000242457 UP000235965 UP000075886 UP000076407 UP000027135 UP000053105
Pfam
Interpro
IPR000209
Peptidase_S8/S53_dom
+ More
IPR032778 GF_recep_IV
IPR015500 Peptidase_S8_subtilisin-rel
IPR006212 Furin_repeat
IPR036852 Peptidase_S8/S53_dom_sf
IPR008979 Galactose-bd-like_sf
IPR022398 Peptidase_S8_His-AS
IPR009030 Growth_fac_rcpt_cys_sf
IPR002884 P_dom
IPR023827 Peptidase_S8_Asp-AS
IPR023828 Peptidase_S8_Ser-AS
IPR034182 Kexin/furin
IPR032815 S8_pro-domain
IPR038466 S8_pro-domain_sf
IPR000742 EGF-like_dom
IPR011031 Multihaem_cyt
IPR032778 GF_recep_IV
IPR015500 Peptidase_S8_subtilisin-rel
IPR006212 Furin_repeat
IPR036852 Peptidase_S8/S53_dom_sf
IPR008979 Galactose-bd-like_sf
IPR022398 Peptidase_S8_His-AS
IPR009030 Growth_fac_rcpt_cys_sf
IPR002884 P_dom
IPR023827 Peptidase_S8_Asp-AS
IPR023828 Peptidase_S8_Ser-AS
IPR034182 Kexin/furin
IPR032815 S8_pro-domain
IPR038466 S8_pro-domain_sf
IPR000742 EGF-like_dom
IPR011031 Multihaem_cyt
Gene 3D
ProteinModelPortal
H9IYN1
Q26489
A0A2A4K7Y7
A0A437BM12
A0A194PYD1
A0A212F4A7
+ More
A0A194RFY5 A0A0A1XGS4 A0A0A1X1M3 A0A0K8U8C6 W8APW3 A0A0L0BQD1 T1P9R1 A0A1I8Q1I4 A0A1Q3FU12 A0A1Q3FTV5 A0A1Q3FUD1 A0A1Q3FTU0 A0A1B6GHW2 W4VRF9 A0A2M4DS50 A0A2M3Z0N1 A0A2M4DS69 A0A2M4A589 A0A2M4B9D5 A0A2M4B9F8 A0A2M4B9B2 A0A2M4A5Q5 A0A2M4A4R7 A0A1B6CCM9 A0A2H1W496 A0A0L7LUE0 A0A2C9GWR4 A0A1J1IF38 E0V9N9 A0A3B0K7Y0 E1JJL4 A0A0R3P116 Q29GI8 B4NCL2 A0A0R3P6H0 P30432 P30432-2 A0A0R3P5B9 B4M1S6 A0A0Q9WMJ9 A0A158NEY8 B4PXS5 A0A1I8Q1H9 B3NX47 A0A0R1EGT9 A0A0R1EEA9 A0A0R1EG80 A0A195FL77 A0A0Q5T532 A0A0Q5T754 A0A437B8F0 B4JKM2 B4R5T4 A0A151IEG1 A0A336LIK8 A0A2A4JGI8 A0A336LJ86 A0A2H1WK85 A0A212FLJ6 A0A2M4CJY4 A0A195BWS1 A0A026X0H0 B4L8R4 A0A0P8ZKG0 A0A0Q9XGD4 A0A0Q9XJA6 B3N0S2 A0A0Q9XRF1 A0A0M4ELX6 E2BLN8 A0A0C9QQN8 A0A194PZ75 A0A1W4VVH0 A0A1W4VTY2 A0A1W4VUQ9 A0A1W4VHD4 E2AVN1 A0A2M4CHY2 A0A2M4CI92 A0A336LJV0 F4WVD8 A0A0C9QJY7 A0A151XIV9 W5JF16 E9ID68 A0A195EN83 A0A2A3EGB8 A0A2J7QX97 A0A182QSE6 A0A182XHH2 A0A067RUA5 A0A0N0U668 A0A2H8TUQ2
A0A194RFY5 A0A0A1XGS4 A0A0A1X1M3 A0A0K8U8C6 W8APW3 A0A0L0BQD1 T1P9R1 A0A1I8Q1I4 A0A1Q3FU12 A0A1Q3FTV5 A0A1Q3FUD1 A0A1Q3FTU0 A0A1B6GHW2 W4VRF9 A0A2M4DS50 A0A2M3Z0N1 A0A2M4DS69 A0A2M4A589 A0A2M4B9D5 A0A2M4B9F8 A0A2M4B9B2 A0A2M4A5Q5 A0A2M4A4R7 A0A1B6CCM9 A0A2H1W496 A0A0L7LUE0 A0A2C9GWR4 A0A1J1IF38 E0V9N9 A0A3B0K7Y0 E1JJL4 A0A0R3P116 Q29GI8 B4NCL2 A0A0R3P6H0 P30432 P30432-2 A0A0R3P5B9 B4M1S6 A0A0Q9WMJ9 A0A158NEY8 B4PXS5 A0A1I8Q1H9 B3NX47 A0A0R1EGT9 A0A0R1EEA9 A0A0R1EG80 A0A195FL77 A0A0Q5T532 A0A0Q5T754 A0A437B8F0 B4JKM2 B4R5T4 A0A151IEG1 A0A336LIK8 A0A2A4JGI8 A0A336LJ86 A0A2H1WK85 A0A212FLJ6 A0A2M4CJY4 A0A195BWS1 A0A026X0H0 B4L8R4 A0A0P8ZKG0 A0A0Q9XGD4 A0A0Q9XJA6 B3N0S2 A0A0Q9XRF1 A0A0M4ELX6 E2BLN8 A0A0C9QQN8 A0A194PZ75 A0A1W4VVH0 A0A1W4VTY2 A0A1W4VUQ9 A0A1W4VHD4 E2AVN1 A0A2M4CHY2 A0A2M4CI92 A0A336LJV0 F4WVD8 A0A0C9QJY7 A0A151XIV9 W5JF16 E9ID68 A0A195EN83 A0A2A3EGB8 A0A2J7QX97 A0A182QSE6 A0A182XHH2 A0A067RUA5 A0A0N0U668 A0A2H8TUQ2
PDB
1P8J
E-value=1.57853e-151,
Score=1379
Ontologies
GO
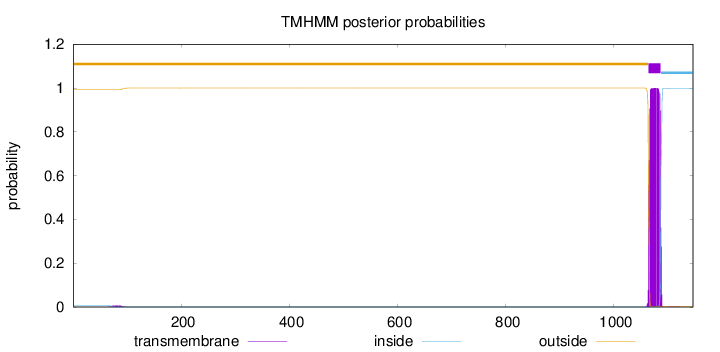
Topology
Subcellular location
Membrane
Length:
1146
Number of predicted TMHs:
1
Exp number of AAs in TMHs:
22.95659
Exp number, first 60 AAs:
9e-05
Total prob of N-in:
0.00672
outside
1 - 1063
TMhelix
1064 - 1086
inside
1087 - 1146
Population Genetic Test Statistics
Pi
213.461858
Theta
178.545172
Tajima's D
0.564621
CLR
0.414994
CSRT
0.527323633818309
Interpretation
Uncertain
