Gene
KWMTBOMO12424
Annotation
PREDICTED:_uncharacterized_protein_LOC107046166_[Diachasma_alloeum]
Location in the cell
Mitochondrial Reliability : 1.324 Nuclear Reliability : 1.919
Sequence
CDS
ATGAATGGTAGCTCTGTAGTTCTGCTGGCTACTGCTAACGTTGAAGTGCGCGACTCATCTGGTCACTTTCATACTCTTCGCGCACTCATCGACCCTGGTAGTCAATCACATTTCATTTCTAATCAAGCTATGGAACGATTGGGCTTAAAGCCTAAGCAAACTTTGCGAAATGTTTGTGGCTTGGGGCAATCGTCGACGTCAGTGACTGGAATGGTGGAGCTTCAACTCGGAGTTCATCAAAAGAATTTATTTAATTTAACGGCTCTCATCTTACCATCTATTTGTGCACAAATGCCCTCTGCTCGTCTTGATCTTTCTTCATGCGAACACCTACGTAATTTGCGACTCGCTGATCCGAATTATAATAAGCCTGGACCAATAGATCTGCTTATCGGTGCTGAAATGTTTTCAACGTTGTTACTGCCCGGAAATATTTCAGGTAGTCCTCATGAACCCTCAGCCTTGAACACCATCTTCGGATGGATCCTGATGGGAAGTGTCAAATGCAGGGATGAGACAAAACACTCAAATTCTTTTTTTCTAAGAGAACACGATACTCCTCTCAATGAGGAATTGAGGAGACTGTGGGAGTTGGAGGTCATACCTAAATCATCAGTTCTAACGCCGGACCAGCAGCAATGTGAGAAACTTTTCACTGAAACCCACGCTCGAGGTACATCTGGAAGGTACATCGTTTCTTTGCCGTTTAGGCAGAGGAATGGGGAGTTACAGTTTCCGGGTTCGCGTGAAAATACATTGCGAAGATTCCATTCATTGGAACGAAGACTTCTTTCAAGTCCAAACCTTCATCAGCAGTATTCGGACTTCATGCTGGACTACTTACATAATGGTCATAGGAGTATCCTTTTATATGCTAAGTCTCGCGTGGCCCCAATTAAGAGAGTTTCATTGCCGAGACTAGAGCTCTGCGCTGCGGTTCTGCTCGCTGACCTGCTCAAATTCGTAAAAGAAGTGTATGTACCGTTACTTCCGCTTTCTGATACATTTGCTTGGTCAGATTCAACGGTAACTCTGTCAGGGATTCGTGGCCATTCCTCGCGTTGGAAAACATTTGTTGCTAATAGGGTTAGTCATATCCAAGAAACTGTCCCCGAAACTCACTGGAATCACGTGCGCTCTAATGAAAATCCTGCGGACATAGCTTCCCGCGGTATGTGTCCGAGCGACCTACTCCAGAGTGAGTTATGGTGGGCGGGACCGAACTGGCTTCGGCAACCTTCATCTTCATGGCCTTCAAATTCCGGTCTCGAGGTGAACGAGACTGATATGCTTGCCGAGGAACGCTGTTTTTCGTTGGTCGTGAATGATGAACCTAACTTTATCGATTCTTTGCTCGAAAGGTTTTCATCAATTCGTAAGGTTCAACGTATTATTGGTTATTGTCTTCGTTTTATCAGAAATGTTAACAAAAAACAGACACGTGTTCCGGGTACCTTTTTAGGTCGTGCAGAACTTCATGAGGCCCTGATGACGATTACCAAGCGAATTCAATTTCTCAACTTTCAGGATGATATTCGTAAGCTCGAGAATGGTATACCATTACCCAAACCGTGGAGAAAACTCAATCCATTCCTGGACGACCAAGGAGTTGTCAGGGTGGGCGGCCGTTTGTCTCGTTCTGGTCTTGAGTTTAGTCATAAGCATCCTGCTCTGTTGCCGCGTCATTCTAGGTTCACATACATGATCATCGAGCAATTCACAAAGAGAATTGTCATCCTGGGTTGGGAACGACACAATATCTTTTGA
Protein
MNGSSVVLLATANVEVRDSSGHFHTLRALIDPGSQSHFISNQAMERLGLKPKQTLRNVCGLGQSSTSVTGMVELQLGVHQKNLFNLTALILPSICAQMPSARLDLSSCEHLRNLRLADPNYNKPGPIDLLIGAEMFSTLLLPGNISGSPHEPSALNTIFGWILMGSVKCRDETKHSNSFFLREHDTPLNEELRRLWELEVIPKSSVLTPDQQQCEKLFTETHARGTSGRYIVSLPFRQRNGELQFPGSRENTLRRFHSLERRLLSSPNLHQQYSDFMLDYLHNGHRSILLYAKSRVAPIKRVSLPRLELCAAVLLADLLKFVKEVYVPLLPLSDTFAWSDSTVTLSGIRGHSSRWKTFVANRVSHIQETVPETHWNHVRSNENPADIASRGMCPSDLLQSELWWAGPNWLRQPSSSWPSNSGLEVNETDMLAEERCFSLVVNDEPNFIDSLLERFSSIRKVQRIIGYCLRFIRNVNKKQTRVPGTFLGRAELHEALMTITKRIQFLNFQDDIRKLENGIPLPKPWRKLNPFLDDQGVVRVGGRLSRSGLEFSHKHPALLPRHSRFTYMIIEQFTKRIVILGWERHNIF
Summary
Uniprot
EMBL
Proteomes
Interpro
SUPFAM
SSF53098
SSF53098
Gene 3D
ProteinModelPortal
Ontologies
KEGG
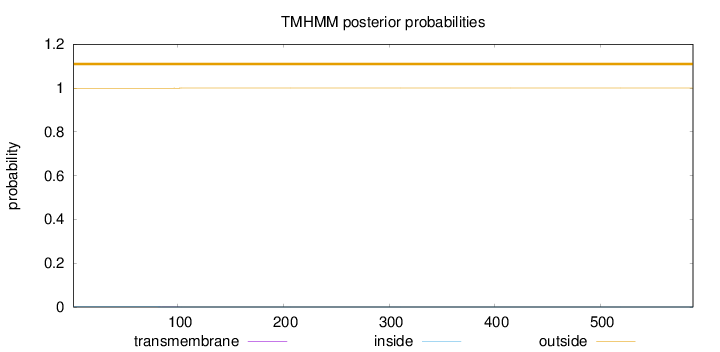
Topology
Length:
588
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.01102
Exp number, first 60 AAs:
0.00024
Total prob of N-in:
0.00067
outside
1 - 588
Population Genetic Test Statistics
Pi
331.619149
Theta
188.303069
Tajima's D
2.313109
CLR
0
CSRT
0.927253637318134
Interpretation
Uncertain
