Gene
KWMTBOMO12297
Annotation
PREDICTED:_uncharacterized_protein_LOC101745010_[Bombyx_mori]
Location in the cell
PlasmaMembrane Reliability : 2.643
Sequence
CDS
ATGGAGTCGTTAGTAGTAGATAAAATTACTGAACGTTTACCGACTACCACAGTTGATGCCTCTCTTTTATCGAATTTCGAAGACATAGCATTGGCAGACGATACTTACTTTAACCCAGGCCCGATCCAACTGTTGATTGGAGCTTCGATTTTCCCGCATCTGCTCTTATCGAACAAGGTTCAAGGACGGCCTTCGCATGTCTCGCCGATCGCGTTAGAGACTATCCTCGGTTATGTTATTGTTGGTGACGCTCCCGTTCGTGTAGCTAACAACGAGAGCGTTTCATATTGCTGTACCGGAGACCCGTTAGACTCCACGATTCGGAAGTTTTGGGAATTAGAGGAGGTAGACGTGCCGTCTGTACTCAGCCCCGAGGACAAAATGTCTGAGGAGTTTTATACCAGTACTACCACACGAGATAGTGACGGACGATACGTGGTGGCTTTGCCCTTTAAAGGTGATGTTAACACTCTCGGTAATTCGCGTCAAGCAAGTGAAAGTCAATTTTACTGCTTAGAAAGAAAGATGCAAGCTTCTCCGCAGCTAAAATTAGCGTACGACAACATCATTCTCGAATATTTAGAGAAGAACTATCTCTCAGCTGTAGATTGTGACAGTTCAGATATTTCTTATTTTATTCCGCATCATGGTGTAATCAGACAAGATAAGGTCACAACCAAATTAAGAGTCGTACTCGATGGTAGTTGTAAGACTACTACTAATGTTTCGCTCAATGATCTGTTATATCGTGGACAAAATTTCCAAGGAAATTTATTTAATATTATTATTAATTTTCGTTTGTTTCCGGTTGCCCTTTCCGCTGACATTCGTCTGATGTTTCTGAGTATTGGAGTCAGAGAATCTGACAGGCGTTTTCAACGCATTCTATATCGTTTTTCTCCTCAACAACCATTACAAGTTTACGAATTTAATCGTGTAGTTTTCGGGCTTAAGTCTAGTCCTTTTCATGCGTTGCGTACTATTAAACAGCTGGCAGCTGATGAAGGTGTCGCTTACCCTCGGGCCAGAGAAATTATTGATACTAGCCTGTATATGGACGATTTTGTTTATTCAATTCCAACTGAAGACGTCGGCATATCAACCGCTTCTGAGATGATAGGTCTCATGAAGGCCGGTCAGTTCGAGCTGGTGAAGTGGACGAGTAACTCACAAGTAGTGTTAGATTCGATCCCGCCGTCGCATCGCCTGCCAGGTGTCAAGGAGTTCGATGAGTCGAACCAGCATAAAGTACTCGGCTTGTGTTGGTCTCCTACGTCGGACAATTTTTGTATAAAAATTGACACTCCGATTGAAATTTGTACTAAACGTACAATACTGTCATGTGTTGCTAAAATATGGGACGTCATGGGATTTGTCGCACCGGTCGTCCTGTTCGTTAAACTTATTATTAAGCAGTTGTGGCTGAGCAATTGTGACTGGGATGACACTCCATCCGAAGATATAGTAAAAATGTGGAGCCGATTCAAAGAAGAACTTCCTTTGCTTTCTCATATAAAGATCCCACGTCACGTAGGTATGAACGTCGACAGCATCGCGACTATATTAGGTTTCGCGGACGCCAGTGAGAAAGCATATGGGGGTGTGATATATTTTCACGTTCAAAATAACAATGATAGTACAATACGTTTGATAACAGCAAAATCAAAGGTTTCTCCTAAAAAGGTTGTGTCGCTCGCCAGACTTGAGCTTTGCGCGATTCTCATACTCGCTCAGCTCATAAAAAATGTAGTAGTCGCTTGTCAAAATCGCATTGTTTTGCGCAATATATTCGCGTTTTCAGACTCAACAGTCGCACTTTGTTGGACACATTCATCGCCTCATCGATGGAGCACCTTTGTGGCTAACAGAGTGACGAAGATACAAAGCTATTTATCTCCACAAAATCTATATCACGTGTCTGGTAAAGACAATCCCGCTGATTGTTTGTCGCGTGGCCTCACTCCTTCCCAAATAATTAATCACCCGCATTGGTTTAGTGGCCCCTCTTGGGCACATTTGGATGTCGCGCAGTGGCCTGCTGTTCCTTTTAATCCTTCCGTAGAGCCTGAGGCGCCGGAAGAAAAGAAACAGACTTTTCTCGCCGCTACTTTAAATGAAAAATCTCCTATATTATCACTAGGAGATCGTATCTCATCTTGGTCTAAACTATTGCGCAGTGTAGTTTTCGTTTGTCGATTTGTAAAATTGTTACCTCACAGATATCGCGTGTCAGCATCGGATTATGAGTTCGCTGAGTTATTAATTCTTCGTGAACTTCAACGGTTTTATTTTCGCGCTGAGATCGCGAAGCTCGAGGAAGGTCAAGTATGTAGCCCTGCATTTCGACACTTAAAACCATTTCTTAAAGATGGAGTAATTAGAGTAGGGGGACGTTTAAGAAACTCTGACTTGAGCTACGACGAGAAATATCCCATTTTGCTTCCGAGAAAAGATCCGATCTTGAATTTACTTGTCGAATATCATCATAAATTAAATTGTCACGCCGGTCCTGACCTGCTGATGGCGCTCTTACGTCAGAAGTATTGGATACTTTCTGCTCGTAATTTAATTAGGAATAGGATACACAAATGCATGCCTTGCTTTCGACTTCGTCCTAAGTAA
Protein
MESLVVDKITERLPTTTVDASLLSNFEDIALADDTYFNPGPIQLLIGASIFPHLLLSNKVQGRPSHVSPIALETILGYVIVGDAPVRVANNESVSYCCTGDPLDSTIRKFWELEEVDVPSVLSPEDKMSEEFYTSTTTRDSDGRYVVALPFKGDVNTLGNSRQASESQFYCLERKMQASPQLKLAYDNIILEYLEKNYLSAVDCDSSDISYFIPHHGVIRQDKVTTKLRVVLDGSCKTTTNVSLNDLLYRGQNFQGNLFNIIINFRLFPVALSADIRLMFLSIGVRESDRRFQRILYRFSPQQPLQVYEFNRVVFGLKSSPFHALRTIKQLAADEGVAYPRAREIIDTSLYMDDFVYSIPTEDVGISTASEMIGLMKAGQFELVKWTSNSQVVLDSIPPSHRLPGVKEFDESNQHKVLGLCWSPTSDNFCIKIDTPIEICTKRTILSCVAKIWDVMGFVAPVVLFVKLIIKQLWLSNCDWDDTPSEDIVKMWSRFKEELPLLSHIKIPRHVGMNVDSIATILGFADASEKAYGGVIYFHVQNNNDSTIRLITAKSKVSPKKVVSLARLELCAILILAQLIKNVVVACQNRIVLRNIFAFSDSTVALCWTHSSPHRWSTFVANRVTKIQSYLSPQNLYHVSGKDNPADCLSRGLTPSQIINHPHWFSGPSWAHLDVAQWPAVPFNPSVEPEAPEEKKQTFLAATLNEKSPILSLGDRISSWSKLLRSVVFVCRFVKLLPHRYRVSASDYEFAELLILRELQRFYFRAEIAKLEEGQVCSPAFRHLKPFLKDGVIRVGGRLRNSDLSYDEKYPILLPRKDPILNLLVEYHHKLNCHAGPDLLMALLRQKYWILSARNLIRNRIHKCMPCFRLRPK
Summary
Uniprot
A0A1S4EIX5
A0A3Q0JJJ7
A0A3Q0JFQ3
A0A1S3DJ34
A0A1S3DJJ0
A0A1S3DJC0
+ More
A0A1S3DIZ2 A0A1S3DMT1 A0A1Y1LIC8 A0A1S3DMK8 A0A3Q0J7N1 A0A3Q0J7K9 S4NIN9 A0A023F096 A0A3Q0JCL2 A0A1S3DH93 A0A1S3DHM0 A0A3Q0JBZ9 A0A1B6MUZ9 A0A1B6KZV2 V5GHM7 J9JVD6 J9JLG6 X1WY95 X1XTX7 A0A1Y1KMX0 A0A146M0N4 A0A0A9VTB0 A0A0A9VUS3 A0A0A9W266 A0A2S2QPS5 A0A2S2NCY5 A0A1Y1LIG5 A0A2S2PM34 J9KD60 J9LPA9 X1XU26 A0A182HA33 X1WZY4 A0A3Q0J0P0 A0A0A9YNG0 A0A224XGH3 A0A1B6KIX6 A0A1B6LLT3 A0A1B6MT91 A0A087T9T6 A0A226CWA5 A0A226DMG9 X1WRA3 W8C1U1 A0A226E226 A0A0A9YQQ8 V5GC07 A0A0J7K7U0 A0A0A9W8L8 A0A0J7KDN6 A0A0A9XKM2 J9JNP4 A0A0A9YU87 A0A146LBU6 A0A1Y1LBZ3 A0A2S2PBK9 A0A0A9WP39 X1WNM7 A0A0J7KCR4 A0A0A9Z001 A0A1U8N896 A0A0K8SX96 A0A226EPI5 A0A1Y1LM33 A0A2S2P1P0 A0A1S3DTS8 A0A226DUQ5 A0A2S2P963 A0A0A9WE49 A0A1S3DBK1 A0A2S2N9D0 A0A2S2PNC2 J9K351 A0A0A9WW84 A0A1S3DM22 A0A023EYR9 A0A023F0C4 A0A1S3DM17 A0A0A9X3X8 A0A224XG39 A0A2S2QLL5 A0A1S3DGE3 A0A0J7N189 A0A0A9VSF7 A0A0A9VYL4 X1XU06 A0A3Q0JI79 A0A1S3DHZ9 A0A3Q0JP94 J9JUK4 A0A182H647 X1X412 J9L9H5 A0A2S2PNZ4
A0A1S3DIZ2 A0A1S3DMT1 A0A1Y1LIC8 A0A1S3DMK8 A0A3Q0J7N1 A0A3Q0J7K9 S4NIN9 A0A023F096 A0A3Q0JCL2 A0A1S3DH93 A0A1S3DHM0 A0A3Q0JBZ9 A0A1B6MUZ9 A0A1B6KZV2 V5GHM7 J9JVD6 J9JLG6 X1WY95 X1XTX7 A0A1Y1KMX0 A0A146M0N4 A0A0A9VTB0 A0A0A9VUS3 A0A0A9W266 A0A2S2QPS5 A0A2S2NCY5 A0A1Y1LIG5 A0A2S2PM34 J9KD60 J9LPA9 X1XU26 A0A182HA33 X1WZY4 A0A3Q0J0P0 A0A0A9YNG0 A0A224XGH3 A0A1B6KIX6 A0A1B6LLT3 A0A1B6MT91 A0A087T9T6 A0A226CWA5 A0A226DMG9 X1WRA3 W8C1U1 A0A226E226 A0A0A9YQQ8 V5GC07 A0A0J7K7U0 A0A0A9W8L8 A0A0J7KDN6 A0A0A9XKM2 J9JNP4 A0A0A9YU87 A0A146LBU6 A0A1Y1LBZ3 A0A2S2PBK9 A0A0A9WP39 X1WNM7 A0A0J7KCR4 A0A0A9Z001 A0A1U8N896 A0A0K8SX96 A0A226EPI5 A0A1Y1LM33 A0A2S2P1P0 A0A1S3DTS8 A0A226DUQ5 A0A2S2P963 A0A0A9WE49 A0A1S3DBK1 A0A2S2N9D0 A0A2S2PNC2 J9K351 A0A0A9WW84 A0A1S3DM22 A0A023EYR9 A0A023F0C4 A0A1S3DM17 A0A0A9X3X8 A0A224XG39 A0A2S2QLL5 A0A1S3DGE3 A0A0J7N189 A0A0A9VSF7 A0A0A9VYL4 X1XU06 A0A3Q0JI79 A0A1S3DHZ9 A0A3Q0JP94 J9JUK4 A0A182H647 X1X412 J9L9H5 A0A2S2PNZ4
EMBL
GEZM01054789
JAV73399.1
GAIX01013989
JAA78571.1
GBBI01004276
JAC14436.1
+ More
GEBQ01007199 GEBQ01000222 JAT32778.1 JAT39755.1 GEBQ01022991 GEBQ01006185 JAT16986.1 JAT33792.1 GALX01004947 JAB63519.1 ABLF02008594 ABLF02030698 ABLF02030705 ABLF02030709 ABLF02030854 ABLF02067347 ABLF02006304 ABLF02036268 GEZM01085145 GEZM01085143 JAV60187.1 GDHC01006469 JAQ12160.1 GBHO01044645 JAF98958.1 GBHO01044643 JAF98960.1 GBHO01044644 JAF98959.1 GGMS01010524 MBY79727.1 GGMR01002421 MBY15040.1 GEZM01054765 JAV73413.1 GGMR01017890 MBY30509.1 ABLF02041863 ABLF02035780 ABLF02001506 ABLF02008029 ABLF02035899 ABLF02037719 ABLF02046327 ABLF02064627 JXUM01030959 KQ560903 KXJ80414.1 ABLF02036131 ABLF02041527 ABLF02064049 GBHO01027036 GBHO01010996 GBHO01010995 GBHO01010993 GBHO01010991 JAG16568.1 JAG32608.1 JAG32609.1 JAG32611.1 JAG32613.1 GFTR01008866 JAW07560.1 GEBQ01028569 JAT11408.1 GEBQ01015458 JAT24519.1 GEBQ01000825 JAT39152.1 KK114194 KFM61875.1 LNIX01000069 OXA36894.1 LNIX01000016 OXA46198.1 ABLF02066034 GAMC01006864 JAB99691.1 LNIX01000007 OXA51795.1 GBHO01009080 JAG34524.1 GALX01000776 JAB67690.1 LBMM01012238 KMQ86334.1 GBHO01039838 GBHO01006034 JAG03766.1 JAG37570.1 LBMM01009202 KMQ88311.1 GBHO01023090 JAG20514.1 ABLF02005684 GBHO01008418 GBHO01008416 JAG35186.1 JAG35188.1 GDHC01014057 JAQ04572.1 GEZM01060254 JAV71154.1 GGMR01014165 MBY26784.1 GBHO01034065 GBHO01034061 JAG09539.1 JAG09543.1 ABLF02042806 LBMM01009551 KMQ88039.1 GBHO01006408 JAG37196.1 GBRD01007977 JAG57844.1 LNIX01000002 OXA59522.1 GEZM01054766 JAV73410.1 GGMR01010750 MBY23369.1 LNIX01000011 OXA48427.1 GGMR01013370 MBY25989.1 GBHO01038805 GBHO01038804 GBHO01038801 JAG04799.1 JAG04800.1 JAG04803.1 GGMR01000737 MBY13356.1 GGMR01018294 MBY30913.1 ABLF02025193 GBHO01031928 GBHO01031926 JAG11676.1 JAG11678.1 GBBI01004410 JAC14302.1 GBBI01004408 JAC14304.1 GBHO01030122 GBHO01030121 GBHO01030120 GBHO01030116 JAG13482.1 JAG13483.1 JAG13484.1 JAG13488.1 GFTR01008874 JAW07552.1 GGMS01009443 MBY78646.1 LBMM01012077 KMQ86420.1 GBHO01044317 GBHO01044315 JAF99286.1 JAF99288.1 GBHO01044316 JAF99287.1 ABLF02025958 ABLF02039606 JXUM01113346 KQ565681 KXJ70776.1 ABLF02008635 ABLF02023121 ABLF02065930 ABLF02065931 ABLF02008636 GGMR01018506 MBY31125.1
GEBQ01007199 GEBQ01000222 JAT32778.1 JAT39755.1 GEBQ01022991 GEBQ01006185 JAT16986.1 JAT33792.1 GALX01004947 JAB63519.1 ABLF02008594 ABLF02030698 ABLF02030705 ABLF02030709 ABLF02030854 ABLF02067347 ABLF02006304 ABLF02036268 GEZM01085145 GEZM01085143 JAV60187.1 GDHC01006469 JAQ12160.1 GBHO01044645 JAF98958.1 GBHO01044643 JAF98960.1 GBHO01044644 JAF98959.1 GGMS01010524 MBY79727.1 GGMR01002421 MBY15040.1 GEZM01054765 JAV73413.1 GGMR01017890 MBY30509.1 ABLF02041863 ABLF02035780 ABLF02001506 ABLF02008029 ABLF02035899 ABLF02037719 ABLF02046327 ABLF02064627 JXUM01030959 KQ560903 KXJ80414.1 ABLF02036131 ABLF02041527 ABLF02064049 GBHO01027036 GBHO01010996 GBHO01010995 GBHO01010993 GBHO01010991 JAG16568.1 JAG32608.1 JAG32609.1 JAG32611.1 JAG32613.1 GFTR01008866 JAW07560.1 GEBQ01028569 JAT11408.1 GEBQ01015458 JAT24519.1 GEBQ01000825 JAT39152.1 KK114194 KFM61875.1 LNIX01000069 OXA36894.1 LNIX01000016 OXA46198.1 ABLF02066034 GAMC01006864 JAB99691.1 LNIX01000007 OXA51795.1 GBHO01009080 JAG34524.1 GALX01000776 JAB67690.1 LBMM01012238 KMQ86334.1 GBHO01039838 GBHO01006034 JAG03766.1 JAG37570.1 LBMM01009202 KMQ88311.1 GBHO01023090 JAG20514.1 ABLF02005684 GBHO01008418 GBHO01008416 JAG35186.1 JAG35188.1 GDHC01014057 JAQ04572.1 GEZM01060254 JAV71154.1 GGMR01014165 MBY26784.1 GBHO01034065 GBHO01034061 JAG09539.1 JAG09543.1 ABLF02042806 LBMM01009551 KMQ88039.1 GBHO01006408 JAG37196.1 GBRD01007977 JAG57844.1 LNIX01000002 OXA59522.1 GEZM01054766 JAV73410.1 GGMR01010750 MBY23369.1 LNIX01000011 OXA48427.1 GGMR01013370 MBY25989.1 GBHO01038805 GBHO01038804 GBHO01038801 JAG04799.1 JAG04800.1 JAG04803.1 GGMR01000737 MBY13356.1 GGMR01018294 MBY30913.1 ABLF02025193 GBHO01031928 GBHO01031926 JAG11676.1 JAG11678.1 GBBI01004410 JAC14302.1 GBBI01004408 JAC14304.1 GBHO01030122 GBHO01030121 GBHO01030120 GBHO01030116 JAG13482.1 JAG13483.1 JAG13484.1 JAG13488.1 GFTR01008874 JAW07552.1 GGMS01009443 MBY78646.1 LBMM01012077 KMQ86420.1 GBHO01044317 GBHO01044315 JAF99286.1 JAF99288.1 GBHO01044316 JAF99287.1 ABLF02025958 ABLF02039606 JXUM01113346 KQ565681 KXJ70776.1 ABLF02008635 ABLF02023121 ABLF02065930 ABLF02065931 ABLF02008636 GGMR01018506 MBY31125.1
Proteomes
Pfam
Interpro
IPR008042
Retrotrans_Pao
+ More
IPR012337 RNaseH-like_sf
IPR001584 Integrase_cat-core
IPR041588 Integrase_H2C2
IPR036397 RNaseH_sf
IPR040676 DUF5641
IPR008737 Peptidase_asp_put
IPR005312 DUF1759
IPR000477 RT_dom
IPR021109 Peptidase_aspartic_dom_sf
IPR001878 Znf_CCHC
IPR041373 RT_RNaseH
IPR001969 Aspartic_peptidase_AS
IPR041577 RT_RNaseH_2
IPR036875 Znf_CCHC_sf
IPR001995 Peptidase_A2_cat
IPR012337 RNaseH-like_sf
IPR001584 Integrase_cat-core
IPR041588 Integrase_H2C2
IPR036397 RNaseH_sf
IPR040676 DUF5641
IPR008737 Peptidase_asp_put
IPR005312 DUF1759
IPR000477 RT_dom
IPR021109 Peptidase_aspartic_dom_sf
IPR001878 Znf_CCHC
IPR041373 RT_RNaseH
IPR001969 Aspartic_peptidase_AS
IPR041577 RT_RNaseH_2
IPR036875 Znf_CCHC_sf
IPR001995 Peptidase_A2_cat
Gene 3D
ProteinModelPortal
A0A1S4EIX5
A0A3Q0JJJ7
A0A3Q0JFQ3
A0A1S3DJ34
A0A1S3DJJ0
A0A1S3DJC0
+ More
A0A1S3DIZ2 A0A1S3DMT1 A0A1Y1LIC8 A0A1S3DMK8 A0A3Q0J7N1 A0A3Q0J7K9 S4NIN9 A0A023F096 A0A3Q0JCL2 A0A1S3DH93 A0A1S3DHM0 A0A3Q0JBZ9 A0A1B6MUZ9 A0A1B6KZV2 V5GHM7 J9JVD6 J9JLG6 X1WY95 X1XTX7 A0A1Y1KMX0 A0A146M0N4 A0A0A9VTB0 A0A0A9VUS3 A0A0A9W266 A0A2S2QPS5 A0A2S2NCY5 A0A1Y1LIG5 A0A2S2PM34 J9KD60 J9LPA9 X1XU26 A0A182HA33 X1WZY4 A0A3Q0J0P0 A0A0A9YNG0 A0A224XGH3 A0A1B6KIX6 A0A1B6LLT3 A0A1B6MT91 A0A087T9T6 A0A226CWA5 A0A226DMG9 X1WRA3 W8C1U1 A0A226E226 A0A0A9YQQ8 V5GC07 A0A0J7K7U0 A0A0A9W8L8 A0A0J7KDN6 A0A0A9XKM2 J9JNP4 A0A0A9YU87 A0A146LBU6 A0A1Y1LBZ3 A0A2S2PBK9 A0A0A9WP39 X1WNM7 A0A0J7KCR4 A0A0A9Z001 A0A1U8N896 A0A0K8SX96 A0A226EPI5 A0A1Y1LM33 A0A2S2P1P0 A0A1S3DTS8 A0A226DUQ5 A0A2S2P963 A0A0A9WE49 A0A1S3DBK1 A0A2S2N9D0 A0A2S2PNC2 J9K351 A0A0A9WW84 A0A1S3DM22 A0A023EYR9 A0A023F0C4 A0A1S3DM17 A0A0A9X3X8 A0A224XG39 A0A2S2QLL5 A0A1S3DGE3 A0A0J7N189 A0A0A9VSF7 A0A0A9VYL4 X1XU06 A0A3Q0JI79 A0A1S3DHZ9 A0A3Q0JP94 J9JUK4 A0A182H647 X1X412 J9L9H5 A0A2S2PNZ4
A0A1S3DIZ2 A0A1S3DMT1 A0A1Y1LIC8 A0A1S3DMK8 A0A3Q0J7N1 A0A3Q0J7K9 S4NIN9 A0A023F096 A0A3Q0JCL2 A0A1S3DH93 A0A1S3DHM0 A0A3Q0JBZ9 A0A1B6MUZ9 A0A1B6KZV2 V5GHM7 J9JVD6 J9JLG6 X1WY95 X1XTX7 A0A1Y1KMX0 A0A146M0N4 A0A0A9VTB0 A0A0A9VUS3 A0A0A9W266 A0A2S2QPS5 A0A2S2NCY5 A0A1Y1LIG5 A0A2S2PM34 J9KD60 J9LPA9 X1XU26 A0A182HA33 X1WZY4 A0A3Q0J0P0 A0A0A9YNG0 A0A224XGH3 A0A1B6KIX6 A0A1B6LLT3 A0A1B6MT91 A0A087T9T6 A0A226CWA5 A0A226DMG9 X1WRA3 W8C1U1 A0A226E226 A0A0A9YQQ8 V5GC07 A0A0J7K7U0 A0A0A9W8L8 A0A0J7KDN6 A0A0A9XKM2 J9JNP4 A0A0A9YU87 A0A146LBU6 A0A1Y1LBZ3 A0A2S2PBK9 A0A0A9WP39 X1WNM7 A0A0J7KCR4 A0A0A9Z001 A0A1U8N896 A0A0K8SX96 A0A226EPI5 A0A1Y1LM33 A0A2S2P1P0 A0A1S3DTS8 A0A226DUQ5 A0A2S2P963 A0A0A9WE49 A0A1S3DBK1 A0A2S2N9D0 A0A2S2PNC2 J9K351 A0A0A9WW84 A0A1S3DM22 A0A023EYR9 A0A023F0C4 A0A1S3DM17 A0A0A9X3X8 A0A224XG39 A0A2S2QLL5 A0A1S3DGE3 A0A0J7N189 A0A0A9VSF7 A0A0A9VYL4 X1XU06 A0A3Q0JI79 A0A1S3DHZ9 A0A3Q0JP94 J9JUK4 A0A182H647 X1X412 J9L9H5 A0A2S2PNZ4
Ontologies
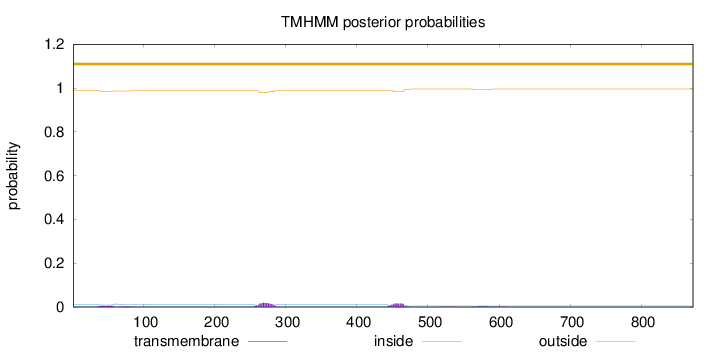
Topology
Length:
873
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
1.00645
Exp number, first 60 AAs:
0.11562
Total prob of N-in:
0.01123
outside
1 - 873
Population Genetic Test Statistics
Pi
11.361299
Theta
23.616085
Tajima's D
-1.605362
CLR
15.671636
CSRT
0.047897605119744
Interpretation
Uncertain
