Gene
KWMTBOMO12169
Annotation
PREDICTED:_uncharacterized_protein_LOC106126637_[Papilio_xuthus]
Location in the cell
Nuclear Reliability : 3.456
Sequence
CDS
ATGTTCGGCATCAGAACACCAGTTAAAAAACCAAACGGCAGTGAGGACAAGTCTGAATTCGAAGGACAGCCAGGTCCGTCTGTAAGGCGCAGCATTGGTGATTGGCCCCCCATAGTCGAAAAAGAGCAGCAAAGGCCAAAAACACCATCTCCACCAAAAGCTGTGCCTCAGCCGCTCCCTCCTAGGCCTAAGACCAAAGCGTTGATCGCCCAGGAAGCAAAAGCTGACCCACGGAGACAGGCGGCAAGAGTCAGTGACAGCCCGCCAATTACAGTCCCGTCGAGATTTCCGAGCAAAACCGCAGAGGCTAAAGCGTGCCTTATGAAAAGCAAGGCGGAGCTCGAAATCTCCAAAAACCTAAAACGGGAGATAAAAGAAGAAGTCTTTAGGTCAATCGAACGGCTCTACCAGCTGGTGAAAGAAGCCGAAGGCGAAAAGGCTTCCCAAGCCACCAACAAGTGGCCGACAGAAGCCTGTAGCATGGTCACTCGGACTGGTAACGAGGGTGACATATCACCGGGCCCAGAAGCGCCGACAGAGCTCATGAAACAGCTAATAGGCGGCCTCAAACAGCACCACGTTGCGTTAGAGGAATGCAAATGGCATACTCGAGCTCTGTCGGAACAGCTGGAGAAAACCCCTCTCTACGCCGCAGGACATGGGTCATTAGATGAACACAGCCAACTCTTAAAGGAAAACAAAGAAGAAATAGAAAAACTAAGTGAAGAAATAAAAAAGCAAAGGACAGAATGGAAAGAAAATAGGAGCTACGCAAGCGTTGTCAGTGGCACAAAGGAGAAAGAACGCGGAAATGCGGAGATGCATTCCGTCGTGATCTCCTCAGACGACCAGAATGCCACAGGGGAAGACGTATTTAATAAGATCAGGTCAACCCTCAACGCTAAAGAGGAAGGGATAAAAATAGACAGGATAAGAAAGGTGAAAGATAGAAAAGTTATAGTCGGCTGTGAATCGAAAGAAGAATTAGTAAAAATAAGAGAAACAATTAAAACAAAGGGAAAGGACTTACACCTCCAGGAGATAAGAAATAAGGAACCCATGGTACAATTGAAGGATGTCCTGAATTATAATAAGGACGAAGATATAGTGCGAGCACTAATTAACCAGAACAAACATCTTCTGGGGGATGTGAGTGTGGACAGCTCAACTCTGAAGGTAAAGTTCAGAAGAAGAGCTAGAAACCCGCTCACATCCCATGTGGCAATACAAGTGCCACCAAAACTATGGAAGCGGCTCACGGAGGCGGGGCTAGTCCATATAGATGTGCAAAGGGTTAGGGTAGAGGATCAGTCTCCCCTGATTCAGTGCTCGCGATGCTTGGGATATGGCCACGGGAAGCGGTTCTGCCGGGAAAAAATTGACGCCTGCAGTCACTGCGGAGAGGCGCATCATAGGTCGGAATGCGCTGACTGGATGGCGAGGGTGCCTCCGTCGTGCTGTAACTGCTTGAAGGCTAAAATCGGTGAAGCCGAGCATAATGCCTTTAGCCCAGAGTGTCCAGTAAGGAAGAGGTGGGAAACCTTGGCAAGATCTACCGTGACTTATTGCTAA
Protein
MFGIRTPVKKPNGSEDKSEFEGQPGPSVRRSIGDWPPIVEKEQQRPKTPSPPKAVPQPLPPRPKTKALIAQEAKADPRRQAARVSDSPPITVPSRFPSKTAEAKACLMKSKAELEISKNLKREIKEEVFRSIERLYQLVKEAEGEKASQATNKWPTEACSMVTRTGNEGDISPGPEAPTELMKQLIGGLKQHHVALEECKWHTRALSEQLEKTPLYAAGHGSLDEHSQLLKENKEEIEKLSEEIKKQRTEWKENRSYASVVSGTKEKERGNAEMHSVVISSDDQNATGEDVFNKIRSTLNAKEEGIKIDRIRKVKDRKVIVGCESKEELVKIRETIKTKGKDLHLQEIRNKEPMVQLKDVLNYNKDEDIVRALINQNKHLLGDVSVDSSTLKVKFRRRARNPLTSHVAIQVPPKLWKRLTEAGLVHIDVQRVRVEDQSPLIQCSRCLGYGHGKRFCREKIDACSHCGEAHHRSECADWMARVPPSCCNCLKAKIGEAEHNAFSPECPVRKRWETLARSTVTYC
Summary
Uniprot
Pubmed
Proteomes
PRIDE
Interpro
Gene 3D
ProteinModelPortal
Ontologies
KEGG
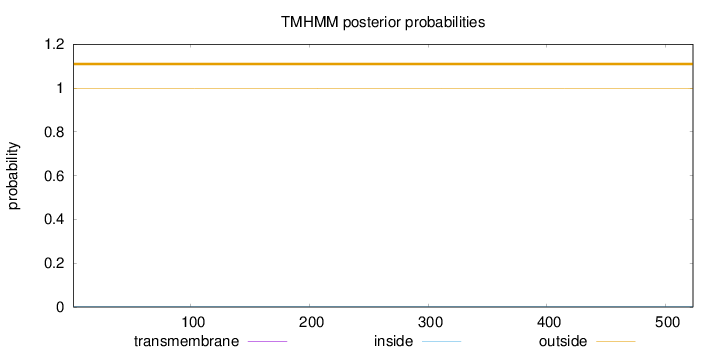
Topology
Length:
523
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00254
outside
1 - 523
Population Genetic Test Statistics
Pi
258.988938
Theta
20.780011
Tajima's D
3.404135
CLR
0.063713
CSRT
0.992450377481126
Interpretation
Uncertain
