Gene
KWMTBOMO12069
Pre Gene Modal
BGIBMGA004292
Annotation
reverse_transcriptase_[Bombyx_mori]
Full name
Probable RNA-directed DNA polymerase from transposon BS
+ More
Probable RNA-directed DNA polymerase from transposon X-element
Probable RNA-directed DNA polymerase from transposon X-element
Alternative Name
Reverse transcriptase
Location in the cell
Nuclear Reliability : 2.505
Sequence
CDS
ATGAAGTTCGCTCTATTGTTCGTATTGACCGCGATCGTCGCGGTGTCGACTAATGAAGTGGCTGACCCCAGATCCACTGACAGAGCTAACGCTTTGAGTATCGGAAGTATTACTTCAAGTGATAGACTCTTACGAAGCTTTGTTGTGAGCCGTGCTGCTACAATTAACTGGCGGGTTCTCAACGTGAGATTCACGGCGCCTGCCGGTAGACGCGTCAGGGCTGTCCGCGTCCGAGGATCGACACAGTTCACAAGGGTCCGCCGCACCGGAGGCGGCGTCGGCTTCACTTTCGTGGAACTCACACACCACAACCCGAATGGCAGGCGGCTTGACGCGTTAGTCGATGATCTCGCCTTCGATATCGTCGCTCCGCTAACACCGACTCACTACCCGCTAAATATCGCGCATCGCCCGGATATACTGGACATAGCGTTATTAAAAAACGTAACTCTGAGCTTACACTCGATCGAAGTAGTTTCAGAGTTAGATTCAGACCACCGTCCCGTCGTTATGAAGCTCGGTCGCGCTCCCGACTCCGTTCCCGTCACGAGGACTGTGGTGGATTGGCACACGCTGGGCATCAGCCTGGCTGAATCTGATCCACCATCGCTCCCGTTTAACTCGGACTCTACCCCGTCTCCTCAGGATACCGCTGAAGCCATAGACATCTTAACGTCACACATCACCTCAACATTAGATAGATCATCGAAACAAGTTGTAGCGGAGGACTTCCTTCACCGCTTCAAATTGTCCGATAATATTAGGGAACTCTTTAGAGCTAAGAACGCCTCGATACGCGCCTACGACAGGTCTCCTACCGCGGAAAATCGTATTCGAATGCGTGCCCTACAACGCGACGTAAAGTCTCGCATCGCCGAAGTCCGTGATGCCAGATGGTCTGATTTCTTAGAAGGACTCGCGCCCTCCCAAAGGTCTTACCGCTTAGCTTGTACTCTCAAATCGGATACGGTAGTAACTATGCCCTCCCTCGTAGGCCCCTCAGGCCGACTCGCGGCGTTTGATGATGACGAAAAAGCAGAGTTGCTGGCCGATACATTGCAAACCCAGTGCACGCCCAGCACTCAATCCGTGGACCCTGTTCATGTAGAATTAGTAGACAGTGAGGTAGAACGCAGAGTCTCCTTGCCCCCCTCGGATGCGTTACCACCCGTCACCCCGATGGAAGTTAAAGACTTGATCAAAGACCTACGTCCTCGCAAGGCTCCCGGTTCCGACGGTATATCCAACCGCGTTATTAAACTTCTACCCGTCCAACTCATCGTGATGTTGGCATCTATTTTCAATGCCGCTATGGTGAACTGTAACACCACACCGCGGGAAAGATCCCGCGGTGTGGAAAGAAGCGGACGCAAACTGTATGAGCGTCTGCTCTACAAACGCCTCAGAGACTTCGTCTCATCCAAGGGCATTCTCATCGATGAACAATTCGGATTCCGTACAAATCACTCATGCGTTCAACAGGCGCACCGCCTCACGGAGCACATTCTTGTGGGGCTTAATCGACCAAAACCGTTATACACGGGAGCTCTCTTCTTCGACGTCGCAAAAGCGTTCGACAAAGTCTGGCACAACGGTTTGATTTTCAAACTATTCAACATGGGTGTGCCGGATAGTCTCGTGCTCATCATACGGGACTTCTTGTCGAACCGCTCTTTTCGATATCGAGTCGAGGGAACCCGCTCCTTCCCACGACCTCTCACAGCTGGAGTCCCGCAAGGCTCTGTCCTCTCACCCCTCCTATTTAGCTTATTCGTTAACGATATTCCCCGGTCGCCGCCGACCCATTTAGCTTTATTCGCCGACGACACGACTGTTTACTATTCCAGTAGAAACAAGTCCCTAATCGCGAAGAAGCTTCAGAGCGCAGCCCTAGCCCTAGGACAGTGGTTCCGAAAATGGCGCATAGACATCAACCCAGCGAAAAGTACTGCGGTGCTATTTCAGAGGGGAAGCTCCACACGGATTTCCTCCCGTATTAGGAGGAGGAATCTCACACCCCCGATTACTCTCTTTAGTCAATCCATACCCTGGGCCAGGAAGGTCAAGTACCTGGGCGTTACCCTGGATGCATCGATGACATTCCGCCCGCATATAAAATCAGTCCGTGACCGTGCCGCGTTTATTCTCGGTAGACTCTACCCCATGATCTGTAAGCGGAGTAAAATGTCCCTTCGGAACAAGGTGACACTTTACAAAACTTGCATAAGGCCCGTCATGACTTACGCGAGTGTGGTGTTCGCTCACGCGGCCCGCACACACATAGACACCCTCCAATCTTTACAATCCCGCTTTTGCAGATTAGCCGTCGGAGCTCCGTGGTTCGTGAGGAACGTTGACCTACACGACGACCTGGGCCTCGAATCTATACAGAAATACATGAAGTCAGCGTCGGAACGGTACTTCGATAAGGCTATGCGTCATGATAATCGCCTTATCGTAGCCGCCGCTGACTACTCCCCGAATCCTGATCATGCAGGAGCCAGTCACCGTCGACGCCCTAGACACGTCCTTACGGATCCATCAGATCCAATAACCTTTGCACTAGACGCCTTCAGCTCTAGGAGCAGGCTTAGGGACCTCGTAAAGTTTGCGGTTTGTGTGCGTCAGATGGAACAATGGGTTATGAAAGGTGGCCCTCAGGAAGCTGATCAGGAAAACAGAAAAACAGAGATGAAGTTCGCTCTATTGTTCGTATTGACCGCGATCGTCGCGGTGTCGACTAATGAAGTGGCTGACCCCAGATCCACTGATAGAGCTAACGCTTTGAGTATCGGAAGTATTACTTCAAGTGATAGACTCTTACGGAGCTTTGTTGTGAGCCGTGCTGCTACAATTAACTCGCGGGTTGTCAACGTGAGGTTCACGGCGCCTGCCGGTAGACGCGTCAGGGCTGTCCGCGTCCGAGGATCGACACAGTTCGCGAGGGTCCGCCGCACCGGAGGCGGCGTCGGCTTCACTTTCGTGAATCTTCAGTTCGTGAACGCGCCGCGAAGAGGATTTCGCTTTACCGTACAAATTTGGGGACGCTAA
Protein
MKFALLFVLTAIVAVSTNEVADPRSTDRANALSIGSITSSDRLLRSFVVSRAATINWRVLNVRFTAPAGRRVRAVRVRGSTQFTRVRRTGGGVGFTFVELTHHNPNGRRLDALVDDLAFDIVAPLTPTHYPLNIAHRPDILDIALLKNVTLSLHSIEVVSELDSDHRPVVMKLGRAPDSVPVTRTVVDWHTLGISLAESDPPSLPFNSDSTPSPQDTAEAIDILTSHITSTLDRSSKQVVAEDFLHRFKLSDNIRELFRAKNASIRAYDRSPTAENRIRMRALQRDVKSRIAEVRDARWSDFLEGLAPSQRSYRLACTLKSDTVVTMPSLVGPSGRLAAFDDDEKAELLADTLQTQCTPSTQSVDPVHVELVDSEVERRVSLPPSDALPPVTPMEVKDLIKDLRPRKAPGSDGISNRVIKLLPVQLIVMLASIFNAAMVNCNTTPRERSRGVERSGRKLYERLLYKRLRDFVSSKGILIDEQFGFRTNHSCVQQAHRLTEHILVGLNRPKPLYTGALFFDVAKAFDKVWHNGLIFKLFNMGVPDSLVLIIRDFLSNRSFRYRVEGTRSFPRPLTAGVPQGSVLSPLLFSLFVNDIPRSPPTHLALFADDTTVYYSSRNKSLIAKKLQSAALALGQWFRKWRIDINPAKSTAVLFQRGSSTRISSRIRRRNLTPPITLFSQSIPWARKVKYLGVTLDASMTFRPHIKSVRDRAAFILGRLYPMICKRSKMSLRNKVTLYKTCIRPVMTYASVVFAHAARTHIDTLQSLQSRFCRLAVGAPWFVRNVDLHDDLGLESIQKYMKSASERYFDKAMRHDNRLIVAAADYSPNPDHAGASHRRRPRHVLTDPSDPITFALDAFSSRSRLRDLVKFAVCVRQMEQWVMKGGPQEADQENRKTEMKFALLFVLTAIVAVSTNEVADPRSTDRANALSIGSITSSDRLLRSFVVSRAATINSRVVNVRFTAPAGRRVRAVRVRGSTQFARVRRTGGGVGFTFVNLQFVNAPRRGFRFTVQIWGR
Summary
Catalytic Activity
a 2'-deoxyribonucleoside 5'-triphosphate + DNA(n) = diphosphate + DNA(n+1)
Cofactor
Mg(2+)
Mn(2+)
Mn(2+)
Miscellaneous
At least 30 copies of X-element are present in the genome and some elements are polymorphic between different strains.
Keywords
Nucleotidyltransferase
RNA-directed DNA polymerase
Transferase
Transposable element
Feature
chain Probable RNA-directed DNA polymerase from transposon BS
Uniprot
Q93137
Q9XXW0
Q9BPQ0
A0A0N1PFV1
O96546
A0A1Y1NA23
+ More
V5GNJ2 A0A139WA11 V5GNF3 A0A139W8G9 A0A2S2QEE0 A0A023EYS8 X1WLB2 X1WNI2 J9LC21 A0A224XB44 A0A2S2PPK8 A0A224XF24 X1WZH0 A0A2J7R7D0 A0A1Y1LKN6 A0A0J7MYU2 A0A2S2PX53 A0A224X6U5 A0A2J7PQB5 Q95SX7 A0A224XIB0 A0A2S2NGV8 A0A2H8TF37 J9KPF1 A0A224X6T4 A0A023EZE6 A0A2J7PU96 A0A224X6S1 A0A2J7QR37 A0A2J7PBY3 A0A2J7QY69 Q07995 A0A2J7QFL0 A0A224X746 A0A1Y1KFR3 A0A2J7R2E1 A0A2J7RKP3 J9JZG6 A0A0V0G5B6 A0A2J7Q0Q6 A0A2S2R9D4 A0A2J7R6U1 A0A2A4IUM2 A0A2J7PVX3 J9LFC9 J9KSK5 A0A0A9YJ00 Q9NBX4 A0A2S2PBB4 A0A2J7R8R8 A0A2S2N6Q7 A0A2J7Q6J2 A0A2M4BD67 A0A2A4K918 A0A2M4BDV6 A0A0A9YU04 A0A2S2PJG2 A0A069DZL8 A0A142LX26 A0A087UGF5 A0A1B6EGD8 A0A142LX49 A0A2M4BEF4 A0A2M4BEZ9
V5GNJ2 A0A139WA11 V5GNF3 A0A139W8G9 A0A2S2QEE0 A0A023EYS8 X1WLB2 X1WNI2 J9LC21 A0A224XB44 A0A2S2PPK8 A0A224XF24 X1WZH0 A0A2J7R7D0 A0A1Y1LKN6 A0A0J7MYU2 A0A2S2PX53 A0A224X6U5 A0A2J7PQB5 Q95SX7 A0A224XIB0 A0A2S2NGV8 A0A2H8TF37 J9KPF1 A0A224X6T4 A0A023EZE6 A0A2J7PU96 A0A224X6S1 A0A2J7QR37 A0A2J7PBY3 A0A2J7QY69 Q07995 A0A2J7QFL0 A0A224X746 A0A1Y1KFR3 A0A2J7R2E1 A0A2J7RKP3 J9JZG6 A0A0V0G5B6 A0A2J7Q0Q6 A0A2S2R9D4 A0A2J7R6U1 A0A2A4IUM2 A0A2J7PVX3 J9LFC9 J9KSK5 A0A0A9YJ00 Q9NBX4 A0A2S2PBB4 A0A2J7R8R8 A0A2S2N6Q7 A0A2J7Q6J2 A0A2M4BD67 A0A2A4K918 A0A2M4BDV6 A0A0A9YU04 A0A2S2PJG2 A0A069DZL8 A0A142LX26 A0A087UGF5 A0A1B6EGD8 A0A142LX49 A0A2M4BEF4 A0A2M4BEZ9
EC Number
2.7.7.49
Pubmed
EMBL
U07847
AAA17752.1
AB018558
BAA76304.1
AB055391
BAB21761.1
+ More
KQ459265 KPJ02064.1 AF081103 AAC72921.1 GEZM01008457 JAV94772.1 GALX01005299 JAB63167.1 KQ971729 KYB24751.1 GALX01005359 JAB63107.1 KQ973342 KXZ75569.1 GGMS01006359 MBY75562.1 GBBI01004395 JAC14317.1 ABLF02042518 ABLF02057746 ABLF02008153 ABLF02008462 ABLF02008464 ABLF02036316 ABLF02036319 ABLF02048663 ABLF02066977 GFTR01008262 JAW08164.1 GGMR01018716 MBY31335.1 GFTR01008038 JAW08388.1 ABLF02034467 NEVH01006736 PNF36716.1 GEZM01053022 JAV74219.1 LBMM01013518 KMQ85605.1 GGMS01000737 MBY69940.1 GFTR01008251 JAW08175.1 NEVH01022639 PNF18527.1 AY060438 AAL25477.1 GFTR01008246 JAW08180.1 GGMR01003407 MBY16026.1 GFXV01000908 MBW12713.1 ABLF02041767 GFTR01008261 JAW08165.1 GBBI01004571 JAC14141.1 NEVH01021205 PNF19912.1 GFTR01008271 JAW08155.1 NEVH01011985 PNF31054.1 NEVH01027074 PNF13841.1 NEVH01009134 PNF33528.1 S59870 AAB26437.2 NEVH01014858 PNF27387.1 GFTR01008151 JAW08275.1 GEZM01084969 JAV60363.1 NEVH01007838 PNF34986.1 NEVH01002716 PNF41415.1 ABLF02037436 GECL01003538 JAP02586.1 NEVH01019964 PNF22161.1 GGMS01017365 MBY86568.1 NEVH01006754 PNF36554.1 NWSH01007753 PCG62823.1 NEVH01020940 PNF20481.1 ABLF02023911 ABLF02014118 ABLF02014121 ABLF02054790 GBHO01010547 GBHO01010546 JAG33057.1 JAG33058.1 AF237761 AAF81411.1 GGMR01014053 MBY26672.1 NEVH01006721 PNF37225.1 GGMR01000226 MBY12845.1 NEVH01017470 PNF24199.1 GGFJ01001858 MBW50999.1 NWSH01000032 PCG80496.1 GGFJ01001857 MBW50998.1 GBHO01021814 GBHO01010549 JAG21790.1 JAG33055.1 GGMR01016905 MBY29524.1 GBGD01000415 JAC88474.1 KU543673 AMS38352.1 KK119692 KFM76444.1 GEDC01000337 JAS36961.1 KU543685 AMS38375.1 GGFJ01002256 MBW51397.1 GGFJ01002257 MBW51398.1
KQ459265 KPJ02064.1 AF081103 AAC72921.1 GEZM01008457 JAV94772.1 GALX01005299 JAB63167.1 KQ971729 KYB24751.1 GALX01005359 JAB63107.1 KQ973342 KXZ75569.1 GGMS01006359 MBY75562.1 GBBI01004395 JAC14317.1 ABLF02042518 ABLF02057746 ABLF02008153 ABLF02008462 ABLF02008464 ABLF02036316 ABLF02036319 ABLF02048663 ABLF02066977 GFTR01008262 JAW08164.1 GGMR01018716 MBY31335.1 GFTR01008038 JAW08388.1 ABLF02034467 NEVH01006736 PNF36716.1 GEZM01053022 JAV74219.1 LBMM01013518 KMQ85605.1 GGMS01000737 MBY69940.1 GFTR01008251 JAW08175.1 NEVH01022639 PNF18527.1 AY060438 AAL25477.1 GFTR01008246 JAW08180.1 GGMR01003407 MBY16026.1 GFXV01000908 MBW12713.1 ABLF02041767 GFTR01008261 JAW08165.1 GBBI01004571 JAC14141.1 NEVH01021205 PNF19912.1 GFTR01008271 JAW08155.1 NEVH01011985 PNF31054.1 NEVH01027074 PNF13841.1 NEVH01009134 PNF33528.1 S59870 AAB26437.2 NEVH01014858 PNF27387.1 GFTR01008151 JAW08275.1 GEZM01084969 JAV60363.1 NEVH01007838 PNF34986.1 NEVH01002716 PNF41415.1 ABLF02037436 GECL01003538 JAP02586.1 NEVH01019964 PNF22161.1 GGMS01017365 MBY86568.1 NEVH01006754 PNF36554.1 NWSH01007753 PCG62823.1 NEVH01020940 PNF20481.1 ABLF02023911 ABLF02014118 ABLF02014121 ABLF02054790 GBHO01010547 GBHO01010546 JAG33057.1 JAG33058.1 AF237761 AAF81411.1 GGMR01014053 MBY26672.1 NEVH01006721 PNF37225.1 GGMR01000226 MBY12845.1 NEVH01017470 PNF24199.1 GGFJ01001858 MBW50999.1 NWSH01000032 PCG80496.1 GGFJ01001857 MBW50998.1 GBHO01021814 GBHO01010549 JAG21790.1 JAG33055.1 GGMR01016905 MBY29524.1 GBGD01000415 JAC88474.1 KU543673 AMS38352.1 KK119692 KFM76444.1 GEDC01000337 JAS36961.1 KU543685 AMS38375.1 GGFJ01002256 MBW51397.1 GGFJ01002257 MBW51398.1
Proteomes
Interpro
SUPFAM
SSF56219
SSF56219
Gene 3D
ProteinModelPortal
Q93137
Q9XXW0
Q9BPQ0
A0A0N1PFV1
O96546
A0A1Y1NA23
+ More
V5GNJ2 A0A139WA11 V5GNF3 A0A139W8G9 A0A2S2QEE0 A0A023EYS8 X1WLB2 X1WNI2 J9LC21 A0A224XB44 A0A2S2PPK8 A0A224XF24 X1WZH0 A0A2J7R7D0 A0A1Y1LKN6 A0A0J7MYU2 A0A2S2PX53 A0A224X6U5 A0A2J7PQB5 Q95SX7 A0A224XIB0 A0A2S2NGV8 A0A2H8TF37 J9KPF1 A0A224X6T4 A0A023EZE6 A0A2J7PU96 A0A224X6S1 A0A2J7QR37 A0A2J7PBY3 A0A2J7QY69 Q07995 A0A2J7QFL0 A0A224X746 A0A1Y1KFR3 A0A2J7R2E1 A0A2J7RKP3 J9JZG6 A0A0V0G5B6 A0A2J7Q0Q6 A0A2S2R9D4 A0A2J7R6U1 A0A2A4IUM2 A0A2J7PVX3 J9LFC9 J9KSK5 A0A0A9YJ00 Q9NBX4 A0A2S2PBB4 A0A2J7R8R8 A0A2S2N6Q7 A0A2J7Q6J2 A0A2M4BD67 A0A2A4K918 A0A2M4BDV6 A0A0A9YU04 A0A2S2PJG2 A0A069DZL8 A0A142LX26 A0A087UGF5 A0A1B6EGD8 A0A142LX49 A0A2M4BEF4 A0A2M4BEZ9
V5GNJ2 A0A139WA11 V5GNF3 A0A139W8G9 A0A2S2QEE0 A0A023EYS8 X1WLB2 X1WNI2 J9LC21 A0A224XB44 A0A2S2PPK8 A0A224XF24 X1WZH0 A0A2J7R7D0 A0A1Y1LKN6 A0A0J7MYU2 A0A2S2PX53 A0A224X6U5 A0A2J7PQB5 Q95SX7 A0A224XIB0 A0A2S2NGV8 A0A2H8TF37 J9KPF1 A0A224X6T4 A0A023EZE6 A0A2J7PU96 A0A224X6S1 A0A2J7QR37 A0A2J7PBY3 A0A2J7QY69 Q07995 A0A2J7QFL0 A0A224X746 A0A1Y1KFR3 A0A2J7R2E1 A0A2J7RKP3 J9JZG6 A0A0V0G5B6 A0A2J7Q0Q6 A0A2S2R9D4 A0A2J7R6U1 A0A2A4IUM2 A0A2J7PVX3 J9LFC9 J9KSK5 A0A0A9YJ00 Q9NBX4 A0A2S2PBB4 A0A2J7R8R8 A0A2S2N6Q7 A0A2J7Q6J2 A0A2M4BD67 A0A2A4K918 A0A2M4BDV6 A0A0A9YU04 A0A2S2PJG2 A0A069DZL8 A0A142LX26 A0A087UGF5 A0A1B6EGD8 A0A142LX49 A0A2M4BEF4 A0A2M4BEZ9
Ontologies
Topology
SignalP
Position: 1 - 17,
Likelihood: 0.994537
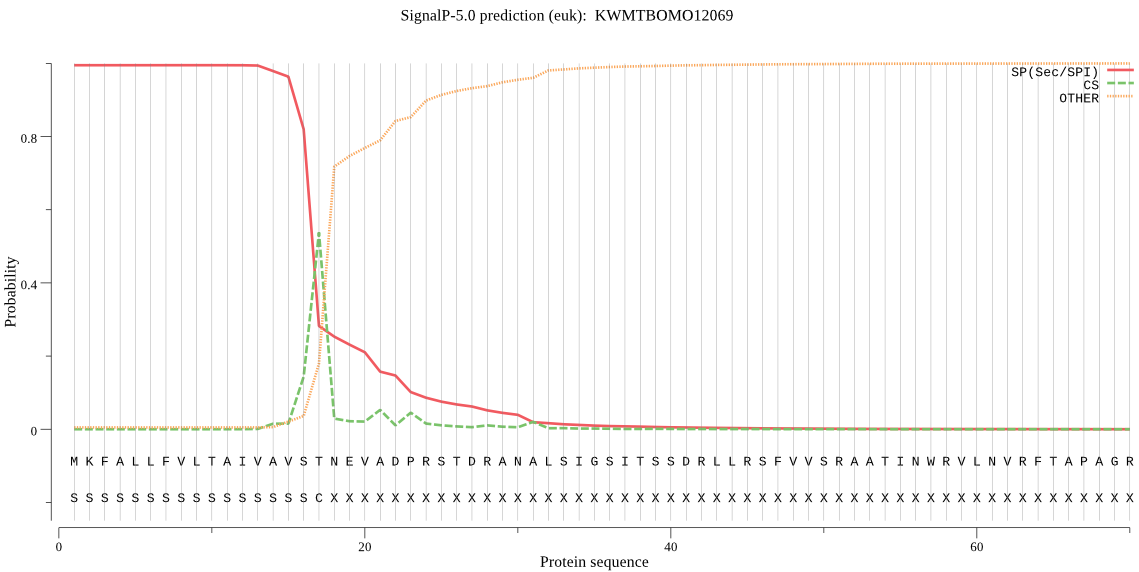
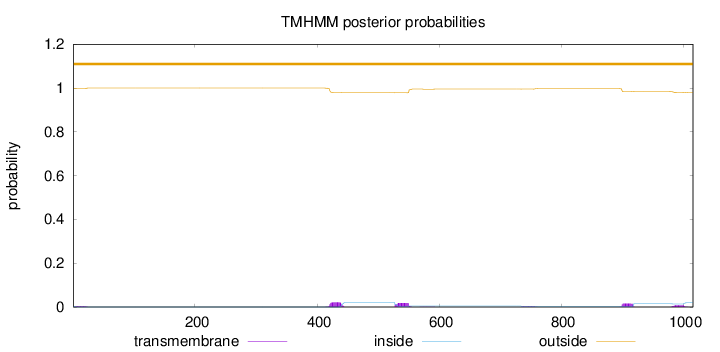
Length:
1016
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
1.40069
Exp number, first 60 AAs:
0.06195
Total prob of N-in:
0.00343
outside
1 - 1016
Population Genetic Test Statistics
Pi
19.34702
Theta
195.457492
Tajima's D
-1.772264
CLR
1087.934635
CSRT
0.031698415079246
Interpretation
Possibly Positive selection
