Gene
KWMTBOMO11835
Pre Gene Modal
BGIBMGA014054
Annotation
PREDICTED:_facilitated_trehalose_transporter_Tret1_[Bombyx_mori]
Location in the cell
PlasmaMembrane Reliability : 4.868
Sequence
CDS
ATGGAAAGATTTAACCTTATTGCAATTGAGGATTTGCCCGCTGGGGAACCGATAGAGCAGGCATTGACGGAGCGAAGTAAGCGTTCATGGAACCCCCGAAATTATGAACCTGACGGAGAGGACATTCCAGTAATTTTGTCTTCACCTGCGATTCCGCCATCACCGGCAATGTACGTTGAATGTGGCGGCAGTGGTACGAGCACGCCATCTTCTAGATATAATATACAGATTTCTGTACATGGCAGCCGTAGTCCTTCGCCGTCACATAAACGTGTGCGTCGATCGCTTTGCCCAGACCAAGAACATGATACTACGAACAGAGATTCTATTTCATCGCTACGTCGACCGCTACGAGACTCTGAAATTTGCTACTTGTTGCACTATGGAAGTGACGACGATGACAGCGATGATGAGAGCCAGAATGATAACGACGAAATCGTTCCCATGATTATCCCCCGGGCAGCTGGAATACTGATGGACACTCCGGACGACGAGGAGATTGAGATTGACACGGCGGCGGCAACTAAACCTCTTGTGACATCTTCCGCGCCTTCCATCATTTCAGCACCTACATCACCAGCGCGTGATACCGGTATGAAGCCTAGTCATTTTTTTGAATTTGACTGGGGAACTTTCCCTGACTCTCCTATACCGCCAACAGAAAGGCGTGAATCTTTTAAGGAACACTCTGGTCCCACTGTTGCTGTAAGTGACCCTTACGAGATTTTTAGACTTATTTGGGACCAGGAGTTCATGGAATTTATTGTGCAGGAAACAAATAGATATGCCCAACAATTAGCTGCTGAAATGTTGGACGGCGGGGAATTACAGGCCTCCAGTAGGATTACAGAATGGAAAGAAACCAATGTAGACGAGTTATTGGTGTTCTTCGGAATATTACTGGCGATGGGAATCGTCATAAAGAACCGGGTTGAAGAGTACTGGAATACGGAACAAAACATCTTCTCCACTCCAGGTTTCAAAGTATATATGTCGCTTAGGAGGTTCCAGTTACTCAGCAGGTGCTTACATTTTAATAATTCCGAGAACTTGAGAAACCTCAACCTGGATCCCTCACAGGCAAAGCTTTTTAAAGTGGAGCCTGTGATTAGCCATTTGAATTCCAAGTTCACGGAATTATATATAATGAAGCAGAACATTGCTTTAGATGAATCGCTGCTGCAGTGGAAAGGTTGGTTGAACATCAACCAATTTATTCCAAACAAAGCTGCAGCAGTGGGCATAAAAACTTACGAAATATGTGAGTCGCAGACGGGGTATTTATGGCGATTTAAAATCCATGCCCATAAGGCTTCGCCTACAGTCTCAGAGGCTGATCCATTCACAGCGAGTACACCGGCGTTGGTTCTTAACCTTATCAAAGGTTTAGAACACAAAGGATACACGTTGTGGATGGATAACTTCTACAATTCCCCAGCACTTGCCCGAAAACTCAAATCGATTGGGTTTGATTGTGTCGGTACGTTGCGGACAAACCGCAAGTATGTCCCGACGGAACTGACCAATCTGAAAAAATCTCAGATGAAGCCCGGGCAAGTAGTAGGCTATACCAGTGGAGATGTCGATTGCATAATATGGAGAGATCAGAACCGCGTAGCGACGATCTCAACGTACCATGGCAATGCCGTCTCCACTAAAAACGGAGTAACAAAACTTATTTTGATACGTGACTACAACATCTGCATGGGTGGCGTGGATAAAAAAGATCAGATGTTGGCTGCATTCCCAATTGAACGCAAGAGGACACAAATTTGGTACAAGAAATTGTTTAAAAGGCCGATGTCTGCGTGGGAGATATCACTACTGGGCAGCTTGCCTAACTTCGGAGCACTGTTGGCTAGTCCCTTCTGTGGACTCGCCTTCAACATTTTAGGAAGAAAATACGCTACGATATTCTTCGGTCTACCTTACGTGATCACATGGACAATAATATCTCTTACGGATAATGTGATCCTAGTACTTGTGGCCATGTCGATCGCGGGTGTGGGCGTTGCGGGGCAAAACGTTGCAATAATTTACGTGTCAGAAATCGCTCATGATTCAATCCGAGGTGGTCTCACGGCATTCTCATCATCGGGGTATATCATAGGGCTTTTGACGTCATACATCTTGGGAGGAAAACTGACGTATCAACAGGTTGTTTATACGCATTTGGGGTTCTCAGTAATGACGATGCTGTTGTTGACCTTACTCCGCGAGAGTCCTGTGTTCCTGGTAATGAAAGGCAGGGAAGAAGAAGCGGAAAAATCTATAGCGTTCTATAAAAGATTGAAAATCGGTTCAAAGGACATAGCATTTGAAATTGACAAAATCAGATTGCAGATTAATCCTAGTTCCGAGAAAACTTATCTACCTGTAGATACTGAAGAAGCAAAAGATTTAGCGAAACATGAAGAGCAACAAGTTGAACAGAAACCGACATCGTCTTGGAAATTTTTGATGAAGTCAAAGTCTTCACAGCGCGCCCTGGCCACAGTGCTAATAATCATGACCCTGATCATATTAATGGGCTGCGTGGTGCTACAAGTGTATGCCGAACCGCTCTTCAAAGAAGCTGTACCTTCGATGCCGCCGAACCAATGCGCCATATTACTCGCTGTGGACTTTTTAATTGCCAGTCTCGTTTGTGTATTTATTATTGATAAGTTTGGCAGAAAGCCCCTGCTAATAGTGACGGCAGCAATTTCGGGCGTCTGCATGTTCCTGCTGGGCTTGCAACTGCAGACACATTTCGCGCCACATTGGTTCACGGTGCTAATCATCTACACGTACAGCTTCGTCTACATGCTGGGCTGCGCTGCTATACCGTTCGTATTGAGCGCCGAACTTTTCTTGCCGGAGGTGAGGAGTCTCTGTAATAGCATCACAATGGTTTGCCTTTGGTCGGTCAATTTTGTGACTCTTCTGGTGTTCAAGCCGCTGGCGGAATGGCTGGGCCTCGGAATAGTGTTCTACTGCTTTTCAATGTTCAGTTTCGTGGGAGTTTTATTTGGCTACGCATACATACCGGAGACGAAAGGTTTATCAGTGGACGCGATACAGTTATTATTTTTGAAGAAAAAATAG
Protein
MERFNLIAIEDLPAGEPIEQALTERSKRSWNPRNYEPDGEDIPVILSSPAIPPSPAMYVECGGSGTSTPSSRYNIQISVHGSRSPSPSHKRVRRSLCPDQEHDTTNRDSISSLRRPLRDSEICYLLHYGSDDDDSDDESQNDNDEIVPMIIPRAAGILMDTPDDEEIEIDTAAATKPLVTSSAPSIISAPTSPARDTGMKPSHFFEFDWGTFPDSPIPPTERRESFKEHSGPTVAVSDPYEIFRLIWDQEFMEFIVQETNRYAQQLAAEMLDGGELQASSRITEWKETNVDELLVFFGILLAMGIVIKNRVEEYWNTEQNIFSTPGFKVYMSLRRFQLLSRCLHFNNSENLRNLNLDPSQAKLFKVEPVISHLNSKFTELYIMKQNIALDESLLQWKGWLNINQFIPNKAAAVGIKTYEICESQTGYLWRFKIHAHKASPTVSEADPFTASTPALVLNLIKGLEHKGYTLWMDNFYNSPALARKLKSIGFDCVGTLRTNRKYVPTELTNLKKSQMKPGQVVGYTSGDVDCIIWRDQNRVATISTYHGNAVSTKNGVTKLILIRDYNICMGGVDKKDQMLAAFPIERKRTQIWYKKLFKRPMSAWEISLLGSLPNFGALLASPFCGLAFNILGRKYATIFFGLPYVITWTIISLTDNVILVLVAMSIAGVGVAGQNVAIIYVSEIAHDSIRGGLTAFSSSGYIIGLLTSYILGGKLTYQQVVYTHLGFSVMTMLLLTLLRESPVFLVMKGREEEAEKSIAFYKRLKIGSKDIAFEIDKIRLQINPSSEKTYLPVDTEEAKDLAKHEEQQVEQKPTSSWKFLMKSKSSQRALATVLIIMTLIILMGCVVLQVYAEPLFKEAVPSMPPNQCAILLAVDFLIASLVCVFIIDKFGRKPLLIVTAAISGVCMFLLGLQLQTHFAPHWFTVLIIYTYSFVYMLGCAAIPFVLSAELFLPEVRSLCNSITMVCLWSVNFVTLLVFKPLAEWLGLGIVFYCFSMFSFVGVLFGYAYIPETKGLSVDAIQLLFLKKK
Summary
Uniprot
ProteinModelPortal
PDB
4ZWC
E-value=7.27122e-17,
Score=217
Ontologies
GO
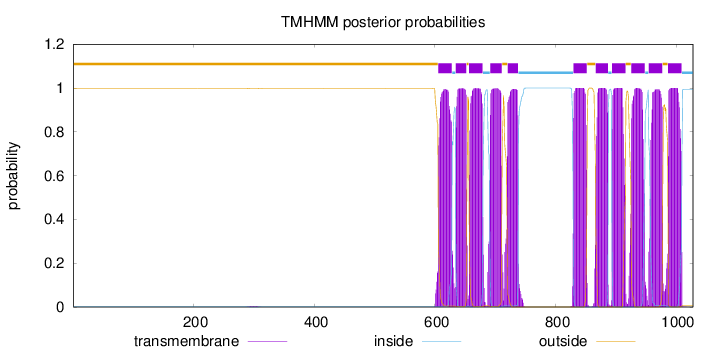
Topology
Length:
1028
Number of predicted TMHs:
11
Exp number of AAs in TMHs:
237.54572
Exp number, first 60 AAs:
0
Total prob of N-in:
0.00196
outside
1 - 605
TMhelix
606 - 628
inside
629 - 634
TMhelix
635 - 652
outside
653 - 656
TMhelix
657 - 679
inside
680 - 691
TMhelix
692 - 711
outside
712 - 720
TMhelix
721 - 738
inside
739 - 829
TMhelix
830 - 852
outside
853 - 866
TMhelix
867 - 887
inside
888 - 893
TMhelix
894 - 916
outside
917 - 925
TMhelix
926 - 948
inside
949 - 954
TMhelix
955 - 977
outside
978 - 986
TMhelix
987 - 1009
inside
1010 - 1028
Population Genetic Test Statistics
Pi
205.085331
Theta
145.845751
Tajima's D
0.396557
CLR
0.532338
CSRT
0.485675716214189
Interpretation
Uncertain
