Gene
KWMTBOMO11702
Annotation
PREDICTED:_uncharacterized_protein_LOC105842184_[Bombyx_mori]
Location in the cell
Mitochondrial Reliability : 1.918
Sequence
CDS
ATGGATGATTTAAAGTTATATGCCAACAATAAGAACAGCCTCAAAGTTTTACTTGATAGCACAGAGATTTTTACTAGAGACATAGGAATGGAATTTGGCTTAAATAAATGCAATGTGCTTCATTTAACATCTGGAAGGAGAGCTGAGCACCAGAGCGAAGGTCACATACTGTTGAGTGGCTTAAACTTCGAACATTTGAAGGAAGACGAAAATTATAAATACTTAGGTATAGATGAAGCAGGCAAAATAGATCATAAAAGAATTCAGGATCAGGTGAAAAGAGAGTACTTCAAAAGAGTAAAATGCATCACTAACACACAACTTAGCGCGAGAAACCTCATAACGGCCATCAACACTTATGCCACACCAGTTCTTTTATACACGTTTGGAATAATAAACTACAAAAATAGTGACTTAAACAAAATAGATATAAAGACAAGGAAGATACTCACTTTGAATAAAGCCCAACAACAAAAAGCGGATATCGAAAGACTCTATTTACCCATAGAAGAAGGAGGCAGAGGACTTATAAATATAGAAATGTTACACAAAACTCAGATTATTAAATTTAGGCAATATTTAAACAATGAACAGGACCAAATCATAAAAGCAATAGTTAAACACGATAAACATAGAGATAAACATTCAATACACAAAGATGGTAAAAATAATGCAGTAAGAATAGAATTAACAGAAAACAGAAACTACACCGATACTGAAATTAAGAAAGCAGTAATAAAGACTGGCATCAGTAAATGGAAAGAGAAACCGCTACATGGACAATTCCGTTGGTCCGTTATAGAGAAGGGTAATATAGATCGAGAGTATTCGTTCCGCTGGCTTAAGAAACAAATCGTGTCCCCCACGTTAGAATCTTCGGTATTCGCTATACAAGATCAGGCAGTAATTACGAGACAAATAGAAAGAGATATTCTAAAGAAATCTGTCGATGGTAAGTGTCGCCTATGCCTAGCAAAAGACGAGTCCATAGAGCATATAATCTCGGGATGTGAAAAACTGGCCGGCACTTACTATGTGAACAGACATAACAACATAGTAAAGTATATTTGGTGGGCAATAGGAAGAAAACACAATTTTGAATTAGACTCGCTATGGTGGAAGGTGAATTTGACTCAACCTCAGATAAAAGAAAACAGCTCCGCGAAGCTAATGTGGGAGATACCAGTGCAGACGGACGTTACAATAAAGCACAATAGACCAGACATAATATACATAGACAAGACAGAAAATAAAACCTACTTAATAGATATAACTGTACCATCTGACTACAATATAGGACCAAAAGAACTGGAAAAACTTAGTAAATATCACCCCTTAAAAGTAGAACTAACACGACTCTGGAATACACACACAGAAACCATCCCCATAGTAATAGGCGCAACGGGCGTGGTGGCAAAAAGTATACATAGATACATAGACAAACTGGAGGCAAATATAAACATACACATAATACAAAAACAAGCAGCCATACACACATGCACAATAATAAGTAAAGTTTTGGGTAATACAGTTTTCATAAATTCAAGGGAGTCCGAACCTCCTCTTCCCTGA
Protein
MDDLKLYANNKNSLKVLLDSTEIFTRDIGMEFGLNKCNVLHLTSGRRAEHQSEGHILLSGLNFEHLKEDENYKYLGIDEAGKIDHKRIQDQVKREYFKRVKCITNTQLSARNLITAINTYATPVLLYTFGIINYKNSDLNKIDIKTRKILTLNKAQQQKADIERLYLPIEEGGRGLINIEMLHKTQIIKFRQYLNNEQDQIIKAIVKHDKHRDKHSIHKDGKNNAVRIELTENRNYTDTEIKKAVIKTGISKWKEKPLHGQFRWSVIEKGNIDREYSFRWLKKQIVSPTLESSVFAIQDQAVITRQIERDILKKSVDGKCRLCLAKDESIEHIISGCEKLAGTYYVNRHNNIVKYIWWAIGRKHNFELDSLWWKVNLTQPQIKENSSAKLMWEIPVQTDVTIKHNRPDIIYIDKTENKTYLIDITVPSDYNIGPKELEKLSKYHPLKVELTRLWNTHTETIPIVIGATGVVAKSIHRYIDKLEANINIHIIQKQAAIHTCTIISKVLGNTVFINSRESEPPLP
Summary
Uniprot
H3AYE9
A0A2J7PDQ6
E0W0M4
E0VPJ4
E0VSD9
E0VA19
+ More
E0W0H3 A0A2J7PIT9 E0VPK6 A0A1S3DB34 A0A1S4EQ35 A0A3B3I0I7 A0A3P9MNJ5 W8BF34 A0A3P8PU69 H3A710 A0A3B5PPD0 A0A3P8NFL6 A0A3B3HW39 A0A3B3I5C9 A0A3B3HU40 A0A3P8QRC6 A0A3P8QAR5 A0A3B3H8G1 A0A3P8R5E8 A0A3P8PU63 A0A3P8NVY4 A0A3P8QC13 A0A3P8NT65 A0A2P2HZD8 A0A2H1VUG2 A0A3P9LVD0 A0A3P8P1I6 A0A3P9QFG6 A0A3P8PST2 A0A3P8P9Q9 A0A3P8QRQ3 A0A3P9MMW4 A0A3P8NRB8 A0A3P8Q3U1 A0A3P8QPZ9 A0A3P8Q8N4 A0A3P8PUT7 A0A3P8QTM2 A0A3P8PSV9 A0A3P8NLS5 A0A3P9M4C8 A0A3P9DUW1 A0A3B3H9G8 A0A3P8NKM6 A0A3P8P8J8 A0A3P8PIV2 A0A3B3HQX9 A0A0S7ESG3 A0A3P8QK92 A0A3S2LN57 A0A3P8QJR1 A0A3P8PQZ9 A0A3P8P1Q8 A0A3P8Q816 A0A3B3ILW9 A0A3P9K6Z6 H3B0S5 A0A3P8NH43 D7GYL6 A0A3B3HNT5 A0A3B3HQ99 H2ZWF0 Q76IN5 A0A1Y1MT60 H2ZUK9 H3AA24 H3AB60 H3A541 W8AXW6 A0A336M9S3 H2ZXX1 R4GD69 H3BBL0 A0A1S3DFN1 A0A2I4AQW5 A0A3B3H9C1 R4GBE7 D7GYE3 R4GAG7 A0A1S3CYA3 H3B0R3 A0A3S2ND61 A0A3B3HDB2 H3AWP0 H3A107 H3A543
E0W0H3 A0A2J7PIT9 E0VPK6 A0A1S3DB34 A0A1S4EQ35 A0A3B3I0I7 A0A3P9MNJ5 W8BF34 A0A3P8PU69 H3A710 A0A3B5PPD0 A0A3P8NFL6 A0A3B3HW39 A0A3B3I5C9 A0A3B3HU40 A0A3P8QRC6 A0A3P8QAR5 A0A3B3H8G1 A0A3P8R5E8 A0A3P8PU63 A0A3P8NVY4 A0A3P8QC13 A0A3P8NT65 A0A2P2HZD8 A0A2H1VUG2 A0A3P9LVD0 A0A3P8P1I6 A0A3P9QFG6 A0A3P8PST2 A0A3P8P9Q9 A0A3P8QRQ3 A0A3P9MMW4 A0A3P8NRB8 A0A3P8Q3U1 A0A3P8QPZ9 A0A3P8Q8N4 A0A3P8PUT7 A0A3P8QTM2 A0A3P8PSV9 A0A3P8NLS5 A0A3P9M4C8 A0A3P9DUW1 A0A3B3H9G8 A0A3P8NKM6 A0A3P8P8J8 A0A3P8PIV2 A0A3B3HQX9 A0A0S7ESG3 A0A3P8QK92 A0A3S2LN57 A0A3P8QJR1 A0A3P8PQZ9 A0A3P8P1Q8 A0A3P8Q816 A0A3B3ILW9 A0A3P9K6Z6 H3B0S5 A0A3P8NH43 D7GYL6 A0A3B3HNT5 A0A3B3HQ99 H2ZWF0 Q76IN5 A0A1Y1MT60 H2ZUK9 H3AA24 H3AB60 H3A541 W8AXW6 A0A336M9S3 H2ZXX1 R4GD69 H3BBL0 A0A1S3DFN1 A0A2I4AQW5 A0A3B3H9C1 R4GBE7 D7GYE3 R4GAG7 A0A1S3CYA3 H3B0R3 A0A3S2ND61 A0A3B3HDB2 H3AWP0 H3A107 H3A543
Pubmed
EMBL
AFYH01132769
NEVH01026385
PNF14463.1
DS235861
EEB19180.1
DS235366
+ More
EEB15300.1 DS235750 EEB16295.1 DS235004 EEB10225.1 DS235859 EEB19129.1 NEVH01024955 PNF16260.1 EEB15312.1 GAMC01018221 JAB88334.1 AFYH01245796 IACF01001441 LAB67141.1 ODYU01004516 SOQ44481.1 GBYX01473740 JAO07919.1 RSAL01001347 RVE40825.1 AFYH01092381 AFYH01092382 GG695857 EFA13462.1 AFYH01259886 AB097127 BAC82595.1 GEZM01022180 JAV88769.1 AFYH01268965 AFYH01207636 AFYH01227415 AFYH01248595 GAMC01016897 JAB89658.1 UFQT01000728 SSX26770.1 AFYH01264165 AFYH01052121 AAWZ02019863 GG695465 EFA13373.1 AFYH01017561 RSAL01000200 RVE44451.1 AFYH01142702 AFYH01142703 AFYH01248007
EEB15300.1 DS235750 EEB16295.1 DS235004 EEB10225.1 DS235859 EEB19129.1 NEVH01024955 PNF16260.1 EEB15312.1 GAMC01018221 JAB88334.1 AFYH01245796 IACF01001441 LAB67141.1 ODYU01004516 SOQ44481.1 GBYX01473740 JAO07919.1 RSAL01001347 RVE40825.1 AFYH01092381 AFYH01092382 GG695857 EFA13462.1 AFYH01259886 AB097127 BAC82595.1 GEZM01022180 JAV88769.1 AFYH01268965 AFYH01207636 AFYH01227415 AFYH01248595 GAMC01016897 JAB89658.1 UFQT01000728 SSX26770.1 AFYH01264165 AFYH01052121 AAWZ02019863 GG695465 EFA13373.1 AFYH01017561 RSAL01000200 RVE44451.1 AFYH01142702 AFYH01142703 AFYH01248007
Proteomes
Pfam
PF00078 RVT_1
SUPFAM
SSF53098
SSF53098
ProteinModelPortal
H3AYE9
A0A2J7PDQ6
E0W0M4
E0VPJ4
E0VSD9
E0VA19
+ More
E0W0H3 A0A2J7PIT9 E0VPK6 A0A1S3DB34 A0A1S4EQ35 A0A3B3I0I7 A0A3P9MNJ5 W8BF34 A0A3P8PU69 H3A710 A0A3B5PPD0 A0A3P8NFL6 A0A3B3HW39 A0A3B3I5C9 A0A3B3HU40 A0A3P8QRC6 A0A3P8QAR5 A0A3B3H8G1 A0A3P8R5E8 A0A3P8PU63 A0A3P8NVY4 A0A3P8QC13 A0A3P8NT65 A0A2P2HZD8 A0A2H1VUG2 A0A3P9LVD0 A0A3P8P1I6 A0A3P9QFG6 A0A3P8PST2 A0A3P8P9Q9 A0A3P8QRQ3 A0A3P9MMW4 A0A3P8NRB8 A0A3P8Q3U1 A0A3P8QPZ9 A0A3P8Q8N4 A0A3P8PUT7 A0A3P8QTM2 A0A3P8PSV9 A0A3P8NLS5 A0A3P9M4C8 A0A3P9DUW1 A0A3B3H9G8 A0A3P8NKM6 A0A3P8P8J8 A0A3P8PIV2 A0A3B3HQX9 A0A0S7ESG3 A0A3P8QK92 A0A3S2LN57 A0A3P8QJR1 A0A3P8PQZ9 A0A3P8P1Q8 A0A3P8Q816 A0A3B3ILW9 A0A3P9K6Z6 H3B0S5 A0A3P8NH43 D7GYL6 A0A3B3HNT5 A0A3B3HQ99 H2ZWF0 Q76IN5 A0A1Y1MT60 H2ZUK9 H3AA24 H3AB60 H3A541 W8AXW6 A0A336M9S3 H2ZXX1 R4GD69 H3BBL0 A0A1S3DFN1 A0A2I4AQW5 A0A3B3H9C1 R4GBE7 D7GYE3 R4GAG7 A0A1S3CYA3 H3B0R3 A0A3S2ND61 A0A3B3HDB2 H3AWP0 H3A107 H3A543
E0W0H3 A0A2J7PIT9 E0VPK6 A0A1S3DB34 A0A1S4EQ35 A0A3B3I0I7 A0A3P9MNJ5 W8BF34 A0A3P8PU69 H3A710 A0A3B5PPD0 A0A3P8NFL6 A0A3B3HW39 A0A3B3I5C9 A0A3B3HU40 A0A3P8QRC6 A0A3P8QAR5 A0A3B3H8G1 A0A3P8R5E8 A0A3P8PU63 A0A3P8NVY4 A0A3P8QC13 A0A3P8NT65 A0A2P2HZD8 A0A2H1VUG2 A0A3P9LVD0 A0A3P8P1I6 A0A3P9QFG6 A0A3P8PST2 A0A3P8P9Q9 A0A3P8QRQ3 A0A3P9MMW4 A0A3P8NRB8 A0A3P8Q3U1 A0A3P8QPZ9 A0A3P8Q8N4 A0A3P8PUT7 A0A3P8QTM2 A0A3P8PSV9 A0A3P8NLS5 A0A3P9M4C8 A0A3P9DUW1 A0A3B3H9G8 A0A3P8NKM6 A0A3P8P8J8 A0A3P8PIV2 A0A3B3HQX9 A0A0S7ESG3 A0A3P8QK92 A0A3S2LN57 A0A3P8QJR1 A0A3P8PQZ9 A0A3P8P1Q8 A0A3P8Q816 A0A3B3ILW9 A0A3P9K6Z6 H3B0S5 A0A3P8NH43 D7GYL6 A0A3B3HNT5 A0A3B3HQ99 H2ZWF0 Q76IN5 A0A1Y1MT60 H2ZUK9 H3AA24 H3AB60 H3A541 W8AXW6 A0A336M9S3 H2ZXX1 R4GD69 H3BBL0 A0A1S3DFN1 A0A2I4AQW5 A0A3B3H9C1 R4GBE7 D7GYE3 R4GAG7 A0A1S3CYA3 H3B0R3 A0A3S2ND61 A0A3B3HDB2 H3AWP0 H3A107 H3A543
Ontologies
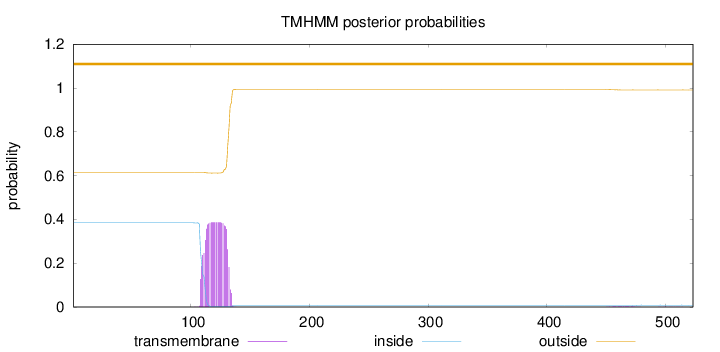
Topology
Length:
523
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
8.62785000000001
Exp number, first 60 AAs:
0.00123
Total prob of N-in:
0.38427
outside
1 - 523
Population Genetic Test Statistics
Pi
6.865318
Theta
5.90465
Tajima's D
-1.071784
CLR
2.709507
CSRT
0.124443777811109
Interpretation
Uncertain
