Gene
KWMTBOMO11598
Annotation
endonuclease-reverse_transcriptase_[Bombyx_mori]
Location in the cell
Mitochondrial Reliability : 1.944
Sequence
CDS
ATGTATGGTAGCGAAACATGGGCGGCAACCAAAAGGAACGTGCATACCATTCAAGTTGCTGAAATGAAAATGCTGAGGTCAATGTGCGGCGTGACTATGCTCAATAAAATCTGTAACGAGTACGTGCGCGGAAGCCTCGGTGTACGGGACATCACCGACAAGATGCCGGAGAGTAGCCTGAGATGGTATGGTCAAGTAAAAAATAAGCCACCTGACTACGTCAAAAATTTAGCGCTACTGCTCGACCTCCCCGGTCGGAGACCTAGAGATAGACCCAAGACAAGATGGAAGGATATAGTGCTGAAGGACAAAAGTGAATGCAATGTCGCTGACGAGGATGTCGAAGACAGAGCGAAGTGGAGGAGTAAAGTGAGGAAAGCTGACCCCACCACCATGTGGGATAAATAG
Protein
MYGSETWAATKRNVHTIQVAEMKMLRSMCGVTMLNKICNEYVRGSLGVRDITDKMPESSLRWYGQVKNKPPDYVKNLALLLDLPGRRPRDRPKTRWKDIVLKDKSECNVADEDVEDRAKWRSKVRKADPTTMWDK
Summary
Uniprot
D7F169
C6Y4D5
A0A016VS97
A0A183FXH7
A0A3P7Z2B4
A0A016WFX2
+ More
A0A183G637 A0A3P8A2N8 A0A016UE98 A0A0N4WFX2 A0A016VNT7 A0A2G9U5Q2 A0A183GA74 A0A3P8F0W9 K1KXQ8 A0A2I0B7E5 A0A016SIC4 A0A016S9A4 A0A016VTA3 A0A016S5Y5 A0A016SNX4 A0A016UBL7 A0A2I0A7U6 A0A2H9ZV89 A0A016VFS4 A0A016VD78 A0A016SDN2 A0A016VLS9 A0A016TQG5 A0A183F2R5 A0A3P7U320 A0A2I0A8C5 A0A016VQL7 A0A2I0BFB2 A0A2I0BDY2 A0A2I0A7B2 A0A183F3S2 A0A3P7UA05 A0A016WHD5 A0A016SAQ3 W6NHZ4 W6NPY2 W6NF39 A0A016VGS9 W6NHB7 A0A2H9ZWL9 W6NDS5 A0A183G016 A0A3P8DTJ1 A0A016VG09 A0A016TKX6 A0A016TSY7 A0A2I0A4R0 A0A016T1D4 A0A016TK27 A0A016T3U9 A0A2I0A8G9 A0A016TQJ7 W6NJ26 A0A183GHA4 A0A3P8C8L5 A0A016VT97 A0A3B3HQB6 A0A016V8D6 A0A183GPW8 A0A3P8ETQ0 A0A016UUZ0 A0A016SDV3 A0A183FT77 A0A3P8AAR6 A0A016SKB8 A0A183GGD8 A0A3P8D1Q5 A0A016V9U6 A0A2G9U478 A0A016WBG8 A0A016WAY5 A0A016U5W8 A0A016VCI1 A0A183G762 A0A3P8A609 A0A183F4B8 A0A3P7UF10 A0A016W910 W6NRE4 I7K8L3 A0A016SG79 A0A183GB20 A0A3P8BZ43 A0A016VMH1 A0A016VLP4 A0A016VVS4 A0A016TLV5 A0A016T5F5 A0A3P9IMA9
A0A183G637 A0A3P8A2N8 A0A016UE98 A0A0N4WFX2 A0A016VNT7 A0A2G9U5Q2 A0A183GA74 A0A3P8F0W9 K1KXQ8 A0A2I0B7E5 A0A016SIC4 A0A016S9A4 A0A016VTA3 A0A016S5Y5 A0A016SNX4 A0A016UBL7 A0A2I0A7U6 A0A2H9ZV89 A0A016VFS4 A0A016VD78 A0A016SDN2 A0A016VLS9 A0A016TQG5 A0A183F2R5 A0A3P7U320 A0A2I0A8C5 A0A016VQL7 A0A2I0BFB2 A0A2I0BDY2 A0A2I0A7B2 A0A183F3S2 A0A3P7UA05 A0A016WHD5 A0A016SAQ3 W6NHZ4 W6NPY2 W6NF39 A0A016VGS9 W6NHB7 A0A2H9ZWL9 W6NDS5 A0A183G016 A0A3P8DTJ1 A0A016VG09 A0A016TKX6 A0A016TSY7 A0A2I0A4R0 A0A016T1D4 A0A016TK27 A0A016T3U9 A0A2I0A8G9 A0A016TQJ7 W6NJ26 A0A183GHA4 A0A3P8C8L5 A0A016VT97 A0A3B3HQB6 A0A016V8D6 A0A183GPW8 A0A3P8ETQ0 A0A016UUZ0 A0A016SDV3 A0A183FT77 A0A3P8AAR6 A0A016SKB8 A0A183GGD8 A0A3P8D1Q5 A0A016V9U6 A0A2G9U478 A0A016WBG8 A0A016WAY5 A0A016U5W8 A0A016VCI1 A0A183G762 A0A3P8A609 A0A183F4B8 A0A3P7UF10 A0A016W910 W6NRE4 I7K8L3 A0A016SG79 A0A183GB20 A0A3P8BZ43 A0A016VMH1 A0A016VLP4 A0A016VVS4 A0A016TLV5 A0A016T5F5 A0A3P9IMA9
EMBL
FJ265554
ADI61822.1
CU462842
CBA11992.1
JARK01001341
EYC29922.1
+ More
UZAH01027842 VDO95476.1 JARK01000336 EYC38182.1 UZAH01029822 VDP08057.1 JARK01001380 EYC13251.1 UZAF01001415 UZAF01017114 VDO08263.1 VDO38063.1 JARK01001343 EYC28702.1 KZ348915 PIO65553.1 UZAH01030983 VDP13266.1 AMGM01000184 EKB47241.1 KZ451907 PKA63681.1 JARK01001560 EYB90061.1 JARK01001602 EYB87210.1 EYC29983.1 JARK01001630 EYB85649.1 JARK01001534 EYB92051.1 JARK01001384 EYC12232.1 KZ452013 PKA51585.1 KZ453539 PKA47214.1 JARK01001347 EYC25573.1 EYC25574.1 JARK01001582 EYB88451.1 EYC28385.1 JARK01001481 JARK01001420 EYB96960.1 EYC04966.1 UZAH01000306 VDO18824.1 KZ452012 PKA51786.1 JARK01001342 EYC29596.1 KZ451885 PKA66495.1 KZ451888 PKA66013.1 PKA51430.1 UZAH01000708 VDO19197.1 JARK01000281 EYC39011.1 JARK01001594 EYB87753.1 CAVP010062000 CDL96883.1 CAVP010057772 CDL94282.1 CAVP010059598 CDL95931.1 JARK01001346 EYC26232.1 CAVP010061241 CDL96703.1 KZ453102 PKA47709.1 CAVP010058665 CDL95025.1 UZAH01028345 VDO99512.1 EYC26231.1 JARK01001430 EYC03360.1 JARK01001415 EYC05905.1 KZ452023 PKA50536.1 JARK01001486 EYB96487.1 EYC03359.1 JARK01001478 EYB97301.1 PKA51832.1 JARK01001422 EYC04723.1 CAVP010061812 CAVP010061814 CAVP010061816 CDL96846.1 CDL96848.1 CDL96850.1 UZAH01033489 VDP29161.1 EYC30277.1 JARK01001351 EYC23521.1 UZAH01036764 VDP46995.1 JARK01001362 EYC18796.1 EYB88469.1 UZAH01027034 VDO87933.1 JARK01001546 EYB91138.1 UZAH01033088 VDP26174.1 JARK01001350 EYC24439.1 KZ350014 PIO64310.1 JARK01000474 EYC36617.1 JARK01000458 EYC36771.1 JARK01001392 EYC10247.1 EYC24438.1 UZAH01030113 VDP09327.1 UZAH01000951 VDO19393.1 JARK01000539 EYC36061.1 CAVP010058483 CDL94742.1 HE805490 CCH50976.1 JARK01001564 EYB89718.1 UZAH01031239 VDP14558.1 JARK01001344 EYC28222.1 EYC28221.1 JARK01001340 EYC31495.1 JARK01001428 EYC03715.1 JARK01001472 EYB97834.1
UZAH01027842 VDO95476.1 JARK01000336 EYC38182.1 UZAH01029822 VDP08057.1 JARK01001380 EYC13251.1 UZAF01001415 UZAF01017114 VDO08263.1 VDO38063.1 JARK01001343 EYC28702.1 KZ348915 PIO65553.1 UZAH01030983 VDP13266.1 AMGM01000184 EKB47241.1 KZ451907 PKA63681.1 JARK01001560 EYB90061.1 JARK01001602 EYB87210.1 EYC29983.1 JARK01001630 EYB85649.1 JARK01001534 EYB92051.1 JARK01001384 EYC12232.1 KZ452013 PKA51585.1 KZ453539 PKA47214.1 JARK01001347 EYC25573.1 EYC25574.1 JARK01001582 EYB88451.1 EYC28385.1 JARK01001481 JARK01001420 EYB96960.1 EYC04966.1 UZAH01000306 VDO18824.1 KZ452012 PKA51786.1 JARK01001342 EYC29596.1 KZ451885 PKA66495.1 KZ451888 PKA66013.1 PKA51430.1 UZAH01000708 VDO19197.1 JARK01000281 EYC39011.1 JARK01001594 EYB87753.1 CAVP010062000 CDL96883.1 CAVP010057772 CDL94282.1 CAVP010059598 CDL95931.1 JARK01001346 EYC26232.1 CAVP010061241 CDL96703.1 KZ453102 PKA47709.1 CAVP010058665 CDL95025.1 UZAH01028345 VDO99512.1 EYC26231.1 JARK01001430 EYC03360.1 JARK01001415 EYC05905.1 KZ452023 PKA50536.1 JARK01001486 EYB96487.1 EYC03359.1 JARK01001478 EYB97301.1 PKA51832.1 JARK01001422 EYC04723.1 CAVP010061812 CAVP010061814 CAVP010061816 CDL96846.1 CDL96848.1 CDL96850.1 UZAH01033489 VDP29161.1 EYC30277.1 JARK01001351 EYC23521.1 UZAH01036764 VDP46995.1 JARK01001362 EYC18796.1 EYB88469.1 UZAH01027034 VDO87933.1 JARK01001546 EYB91138.1 UZAH01033088 VDP26174.1 JARK01001350 EYC24439.1 KZ350014 PIO64310.1 JARK01000474 EYC36617.1 JARK01000458 EYC36771.1 JARK01001392 EYC10247.1 EYC24438.1 UZAH01030113 VDP09327.1 UZAH01000951 VDO19393.1 JARK01000539 EYC36061.1 CAVP010058483 CDL94742.1 HE805490 CCH50976.1 JARK01001564 EYB89718.1 UZAH01031239 VDP14558.1 JARK01001344 EYC28222.1 EYC28221.1 JARK01001340 EYC31495.1 JARK01001428 EYC03715.1 JARK01001472 EYB97834.1
Proteomes
PRIDE
Interpro
SUPFAM
SSF56219
SSF56219
Gene 3D
ProteinModelPortal
D7F169
C6Y4D5
A0A016VS97
A0A183FXH7
A0A3P7Z2B4
A0A016WFX2
+ More
A0A183G637 A0A3P8A2N8 A0A016UE98 A0A0N4WFX2 A0A016VNT7 A0A2G9U5Q2 A0A183GA74 A0A3P8F0W9 K1KXQ8 A0A2I0B7E5 A0A016SIC4 A0A016S9A4 A0A016VTA3 A0A016S5Y5 A0A016SNX4 A0A016UBL7 A0A2I0A7U6 A0A2H9ZV89 A0A016VFS4 A0A016VD78 A0A016SDN2 A0A016VLS9 A0A016TQG5 A0A183F2R5 A0A3P7U320 A0A2I0A8C5 A0A016VQL7 A0A2I0BFB2 A0A2I0BDY2 A0A2I0A7B2 A0A183F3S2 A0A3P7UA05 A0A016WHD5 A0A016SAQ3 W6NHZ4 W6NPY2 W6NF39 A0A016VGS9 W6NHB7 A0A2H9ZWL9 W6NDS5 A0A183G016 A0A3P8DTJ1 A0A016VG09 A0A016TKX6 A0A016TSY7 A0A2I0A4R0 A0A016T1D4 A0A016TK27 A0A016T3U9 A0A2I0A8G9 A0A016TQJ7 W6NJ26 A0A183GHA4 A0A3P8C8L5 A0A016VT97 A0A3B3HQB6 A0A016V8D6 A0A183GPW8 A0A3P8ETQ0 A0A016UUZ0 A0A016SDV3 A0A183FT77 A0A3P8AAR6 A0A016SKB8 A0A183GGD8 A0A3P8D1Q5 A0A016V9U6 A0A2G9U478 A0A016WBG8 A0A016WAY5 A0A016U5W8 A0A016VCI1 A0A183G762 A0A3P8A609 A0A183F4B8 A0A3P7UF10 A0A016W910 W6NRE4 I7K8L3 A0A016SG79 A0A183GB20 A0A3P8BZ43 A0A016VMH1 A0A016VLP4 A0A016VVS4 A0A016TLV5 A0A016T5F5 A0A3P9IMA9
A0A183G637 A0A3P8A2N8 A0A016UE98 A0A0N4WFX2 A0A016VNT7 A0A2G9U5Q2 A0A183GA74 A0A3P8F0W9 K1KXQ8 A0A2I0B7E5 A0A016SIC4 A0A016S9A4 A0A016VTA3 A0A016S5Y5 A0A016SNX4 A0A016UBL7 A0A2I0A7U6 A0A2H9ZV89 A0A016VFS4 A0A016VD78 A0A016SDN2 A0A016VLS9 A0A016TQG5 A0A183F2R5 A0A3P7U320 A0A2I0A8C5 A0A016VQL7 A0A2I0BFB2 A0A2I0BDY2 A0A2I0A7B2 A0A183F3S2 A0A3P7UA05 A0A016WHD5 A0A016SAQ3 W6NHZ4 W6NPY2 W6NF39 A0A016VGS9 W6NHB7 A0A2H9ZWL9 W6NDS5 A0A183G016 A0A3P8DTJ1 A0A016VG09 A0A016TKX6 A0A016TSY7 A0A2I0A4R0 A0A016T1D4 A0A016TK27 A0A016T3U9 A0A2I0A8G9 A0A016TQJ7 W6NJ26 A0A183GHA4 A0A3P8C8L5 A0A016VT97 A0A3B3HQB6 A0A016V8D6 A0A183GPW8 A0A3P8ETQ0 A0A016UUZ0 A0A016SDV3 A0A183FT77 A0A3P8AAR6 A0A016SKB8 A0A183GGD8 A0A3P8D1Q5 A0A016V9U6 A0A2G9U478 A0A016WBG8 A0A016WAY5 A0A016U5W8 A0A016VCI1 A0A183G762 A0A3P8A609 A0A183F4B8 A0A3P7UF10 A0A016W910 W6NRE4 I7K8L3 A0A016SG79 A0A183GB20 A0A3P8BZ43 A0A016VMH1 A0A016VLP4 A0A016VVS4 A0A016TLV5 A0A016T5F5 A0A3P9IMA9
Ontologies
PANTHER
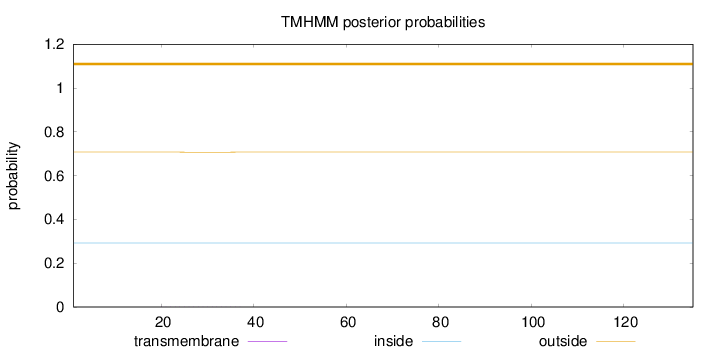
Topology
Length:
135
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00444
Exp number, first 60 AAs:
0.00444
Total prob of N-in:
0.29253
outside
1 - 135
Population Genetic Test Statistics
Pi
131.775598
Theta
111.237912
Tajima's D
-0.259133
CLR
0
CSRT
0.302034898255087
Interpretation
Uncertain
