Gene
KWMTBOMO11304
Pre Gene Modal
BGIBMGA001988
Annotation
PREDICTED:_transient_receptor_potential_cation_channel_trpm_isoform_X1_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 2.635
Sequence
CDS
ATGTTGTACGGTGAAGTATTCGCTGGCGATATAGATCCTCCTTGCGGGAAGGAGATCGGCGACCGGAAATGCGTCACCGGAAGATGGATCACGCCCATCGCCATGACAGTGTACCTGCTCATCGCCAATATATTACTGATCAATCTCCTAATTGCCGTTTTTAATAATATATTCAACGAAGTCAATGAAGTCTCGCATCAGGTGTGGATGTTCCAAAGGTTCACAGTGGTCATGGAGTATGAACAGAAGCCAGTTCTGCCGCCACCGCTGATCATCTTCTGCCACATCTACGCCGTTGGCAAGTGGGTGACCAGAAACGTCTCGCACAAGAGATTCCAATACGACAACGGACTGAAGCTGTTCTTGGAGAAAGAAGACATGGAGAGGCTTTATGACTTCGAAGAGGAATGCGTGGAGGGATACTTTAGGGAACAAGAAATAAAATTATCGAATTCAATAGAGGAGAGAGTTAGGAATACAACAGATAGGGTAGAACATATAACGACCAAAATGGAAGACTTGAATCAAAAGGCAAATTGCCAAATACAATCGATGCAGAGCGTTGAATTTAGGTTAAGAAAGCTCGAAGATATAGCGGAGCAGACGATGAGCCACCTGGCGGTGATACACAGGTTCATGGCGACTCATCCTTCGCTGGTCAACCGGGGGTCCACAGACACTCTCGGACCGCCGCCCCCAAGCTTCCAGGCCGGGGGGAATAGACTTAGAACTGCCAGTGATCGCTCCGAGGTACTATCAGACAGCGACACTGGTGCTCGCTTCGCCGTGCGCCCGGTACAAGAGGAGGACGAACAACCCTCCTGCCACTCTGACGTAAGCCACTCCGACACGTTAAAACTCCGCGACCGTTGTCTAAACAATCGTTCCTTTAAATCGCGTCTATGGAAGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGAGAGAGAGAGAGCAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAGAGAGCGAGAGAGAGAGAGAGAATACACTTTATTGCACACCAAAACACAATTTACATTCAAAAAACAATCAAAAAAGCAGATACATTTGTTAGGTGAGGACGAGCTGCCGCAGTCGGGAACAGAGGCTATATGTGGAGAAGCCAAGGAGCCGGAAGCCGACTCGATATCCACTTCGTCCGGAGGGAGCGCAGCCGGTCCTGACGTCACGGTGGTACGGCAGCAACCAGCCATAATGCCGGCCAGACAACGATCATCGGATAGCAACAGAACCGAACCCAGTGATGAGAAGAAAGTGGCATCATCGACAGACGACGCTTTCCTCCCAGAAGTCCGCGAAATGCCTACAGACCTGCACAGCTTGAGACCTCGGACCACATCGGGCGGGACAAGACGTACAGCTAACACCAGAAGGCGCTCCGAAGGAGGCAACGACTGCGGCATGGGCCCCTACTCCTCACAACACTTGGCTCCGAGGAGACAGCACAGTCAAACGCATTCTGAACCCGACAACTCTGCAGTAGACGCTGGTTCTGTGCACAATTCCGTAACAGGTCCTCAATCGGGGTGGGAGGGCGCGGGGGCCACGGGGGCCGCGGGGGGGACAGGGAGGGCTCGCGTGGGGGGCGCGCCCCGCTCCCTGCTGCTCGCCATGCACGAGTACACGAGCATCACGGACGAACTGGAAACTGTTTATGGACTCTTCTCACCGCCGCACACGCCGAGAACACCGATAACTTCACGTCTGCTGTCCCCCGCCCGCGCGGCCTCTCCCTCCGTCCGTCGACGTCACGCCAGCGAGATGTCCAACCCCGAGTTCGCACTCTTCATGGAGAAGGAGCACCTTCGAGGAGCCGAAGAAGACGACTACATCATCATGGAGAACCTGATCCAAAGGCGCCTAGAAATGGGGTCTGTACTGGACAGATACGAGGAGGAGGAGGGCGGGGAGGGGGCGGGCGCGGGGGGAGCGATCAGCATCAGCGTCTGCGTCAACACCAGCGAGGAGCAACCACAGAGTTCGAGTCTCCTCACCGTGACAGACAGACGTCTACCGCATCAGAGGCACTCGCTAAGGAGATCTTCAGCTGTCGACAGCCGGGAGCTTCCGTCGCCCCTGCAGCCTCTGCTCGGAGATGATGCTTGTTGCGAAGTCTTACTGCCGGTTCCGAGACAGCCTTCGGGAGATACGAACGCATCTAGCGACCTCTCGCTCTCACACATGGTGAGCGAGAATCTCCCGCCGCGGCCCTCCATAGTGCTCGACTCGTCTCAGCTCCCGCGTCAAAGGTCCGCGGAGCAACAGCAGCCGCAGCCTCCGGGACCGTCCAGACCCGAGACAATGTGCTAA
Protein
MLYGEVFAGDIDPPCGKEIGDRKCVTGRWITPIAMTVYLLIANILLINLLIAVFNNIFNEVNEVSHQVWMFQRFTVVMEYEQKPVLPPPLIIFCHIYAVGKWVTRNVSHKRFQYDNGLKLFLEKEDMERLYDFEEECVEGYFREQEIKLSNSIEERVRNTTDRVEHITTKMEDLNQKANCQIQSMQSVEFRLRKLEDIAEQTMSHLAVIHRFMATHPSLVNRGSTDTLGPPPPSFQAGGNRLRTASDRSEVLSDSDTGARFAVRPVQEEDEQPSCHSDVSHSDTLKLRDRCLNNRSFKSRLWKRERERERERERERERERESERERARERERERERERESERERARERERERESERERERERARERERERESERERARERERERERERERESERERARERGERESERERARERERERESERERARERERESERERERERESERERERERESERERARERERERESERERARERERESERERERERERERERESERERERERESERERESERERARERERERERERERERERERERERARERERERERERERARERERERERERERERARERERERESERERERERESERERARERERARERERERERERERERAERERERARERERERERERERERARERERERERERERERESEREREREREREREYTLLHTKTQFTFKKQSKKQIHLLGEDELPQSGTEAICGEAKEPEADSISTSSGGSAAGPDVTVVRQQPAIMPARQRSSDSNRTEPSDEKKVASSTDDAFLPEVREMPTDLHSLRPRTTSGGTRRTANTRRRSEGGNDCGMGPYSSQHLAPRRQHSQTHSEPDNSAVDAGSVHNSVTGPQSGWEGAGATGAAGGTGRARVGGAPRSLLLAMHEYTSITDELETVYGLFSPPHTPRTPITSRLLSPARAASPSVRRRHASEMSNPEFALFMEKEHLRGAEEDDYIIMENLIQRRLEMGSVLDRYEEEEGGEGAGAGGAISISVCVNTSEEQPQSSSLLTVTDRRLPHQRHSLRRSSAVDSRELPSPLQPLLGDDACCEVLLPVPRQPSGDTNASSDLSLSHMVSENLPPRPSIVLDSSQLPRQRSAEQQQPQPPGPSRPETMC
Summary
Uniprot
ProteinModelPortal
PDB
6BWF
E-value=2.34989e-30,
Score=334
Ontologies
GO
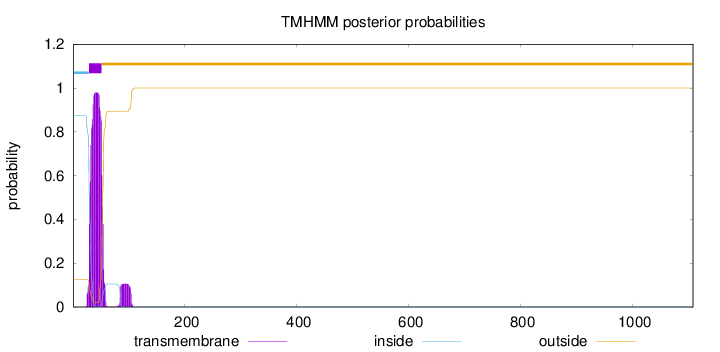
Topology
Length:
1108
Number of predicted TMHs:
1
Exp number of AAs in TMHs:
24.28702
Exp number, first 60 AAs:
22.16046
Total prob of N-in:
0.87363
POSSIBLE N-term signal
sequence
inside
1 - 28
TMhelix
29 - 51
outside
52 - 1108
Population Genetic Test Statistics
Pi
210.471855
Theta
171.070008
Tajima's D
0.717171
CLR
0.412679
CSRT
0.574821258937053
Interpretation
Uncertain
