Pre Gene Modal
BGIBMGA008581
Annotation
PREDICTED:_fatty_acid_synthase-like_[Bombyx_mori]
Location in the cell
Nuclear Reliability : 2.09 PlasmaMembrane Reliability : 1.498
Sequence
CDS
ATGTTGACAACGTACACTCGAGGCTTAGTGTGTACCCAATCGCCGCTCATCGATGGATCTATGGCCGCCGTTGGACTTGGATATCAAAGCATTGCCAGACAATGTCCACCAGAAATAGATATTGCCTGTCACAACAGTTTCGAATCTACTACAATATCAGGACCGGCGGAAGTTACAAAATTATTCGTTGATGATCTAAAAAAACGTGGTATTTTTGCAAAAGAAGTCCCATGTGGAAACGTTGCCTTCCATTCCAGATATATTGCCTTGGCAGGACAAGCACTTTTAGATGCCTTGAAGAAAATTATTAAAGAACCTAAGCCTCGTAACGAGAAATGGTTGTCCACGTCCGTGCCACAACACAAATGGCACGAATTATCGGCAAAGTATTCTTCTCCTGAATACCATACCAATAACATGCTAAGTCCAGTTCTTTTCGAAGAAGTCACAAGGTTAATACCAGACAATGCTATTGTAATAGAAATAGCGCCTCACGGACTTCTCCAAGCCATTCTTAAGAGATCTTTGCAGAACTGTTTGCATATTCCACTGACAAAACGCGGAACGAATGACAGTGTTCAATTTTTGTTGGATGCGTTTGGCAAGATGTTTGAAGCAGGTTTAAATCCGAAAATTGAGAAGTTGTATCCTAAGATAGAGTTTCCAGTTTCGACTCAAACTCCTCTTTTGTCCAATTTAATTGAATGGGAACACAGTGAGGATTGGCTCCATTTGTTTTCCGGTTCCGGAGTAAAATGCACGGCAATCGCTCGAGATGTTGTCATTTCTTTTTACAACGATGATTACAACTTTTTGAAAGGACATTTGAGGAATGGACTAAACATATTACCTGAAGCTGAAATATTAGTTCTGGTTTGGGAAACAATGGCCATGTACAAGAATATGGATTACAGATACCTTTCTGTGATGTTTAAAGACGTACATTTTCATAGGGAAGTCGAGATTAAGCGTGATATCCCGTTAAGATTATCTATAAGTATAACGAAAGGAAACAATAGATTTGAGGTCATTCACGAAGACGCGATAGTGGTATGTGGAATTTTTGATGAGAGCGTCATTATGAAATCCGACCATGGTATAGATGAAACCAAATTTAACAAAGACGAAATTTTACTTAATACTGAAGATATTTACCAGATTCTAGAGCTACGAGGACACTGCTATACAGGTGAACACAAATCTATAGCATCAACGAACTTAGAACGGAATAAGGCGATAGTGCGGTGGAACAATAATTGGGCCTTATATCTGGATTCACTCATACAACTGAACATATTGGCTAATGAACACGAGGGCTTGTCAGTACCGAAATTTATTGGAAAAATAGCAATCAGTGTGTCTGAACATGAAAAAGCGATGCAAAAAGTCAATAATTTAACTGATGGTCATGTAGCCAACATTATCGATACATTTGATGTGACAAGGTGTGCAGGAGTTGAGATAGACCGTTTAATTTTCACCGAGAAAGCAGTCATTAACAAAGAGGCAGACGGCCTACAAATATTATCATTCTGTCCACAATATCCAACCGAGGAAATTAGTTTACAGCGAGCTATCAATGTGGTTTTAAAGATAGTAGCAGAAAATTCGTCTTCAAAAAAAATTAAAATAATTAAATCAGCCTCCTTAAGTATCGATGCATCTGATACTATTGAGAAAATAATTCAGACTGAACCGATGGAGGTAGAAGTTGTTACAGCAAACATCGACAATTTTCACAAAAAGAGTAATGGCATTAATGATGAACTAGCTGAAGTCAACTTAGTGATTATTGATAATATTTTGAATGATAAACAGAAGTTAGCAAAAATACACGAAACAATCTCAAAGGATACATTCATATTGACATTCGAAGCGAATGTACAAAAAAATAATCTAATTAAGAATTATTTTAACATAATAACGTGCATGTCAAGTGAAAATGGAATATGTATACTTCTGAAGAGAATTCCTCTAGAACGAGTTCAGCGTACATGTATACCAATTAACTATAAAACATATAAAGAACATTTGCTGTACGTGTATGAAGAATTGAAAAGGACAAGCGTAGTCTTGGTATCAGAGGGACAAAGCCTTTGCGGTATTTCAGGACTGGTAAAAAAATTAAGAAGAGATGGGCATAAAAATATATCTGCCATGATTATAGAGGATACTGATGCACCAAAGTTTAACCCTGACCATCCATTTTACAAAAGTCAGTTGGATTTGGATTTAGCTGCCAATGTTTTAAGAAAGCGGCAATGGGGAGGCTACTACTACATACCAAGTCAGATGTCAGCAAGCGTTCGTAACATGACCCTAATGAGCAAACAGCCTGGCAGCATTGATGGGTTGACGTGGGTTGAAGCTCCAGCTACAACAGATGGTGCTGAACATGTTCAGGTTTGGTACGCAGGGCTGTCACTAAAAGATTGCCAGAAGGCTATTAGTTGTGAGAATAACAACAGATCGTATTTCGGAATGGACTTCAGTGGCTATAATTCAAAAGGGGAGAAAATAATGGGTTTGGTACGCGAAGGGTCGTTGTCTAGTTCAGTGGAAGCTGATCCAGATCTAACGTGGTCAGTACCTGAACACTGGACGCTTGAAGAGGCAGCCACCGTTCCCCTAGCGTACCTTCATGCCTTCTATTGCTTGGAACTAAGGTGCGATATTTCTAGTGGAAAAACAGTATTCATCACAGGAGGGGCTGGGGCTCTAGGACAAGCAGTCATCTCCATATGCCTGGCTATGAACTGTCCTATTTATACAACCGTGAGAGATATTCATAAGAAAAACTTTTTATTAAAGCTGTTCCCTGAACTTCCAGAAAGTCACATTGGAATTTCCCACCTTGAAGCTTTATATAGCATGCTCATATTTAATTCTGAGAATAAACTTTGCGATATAATTATAAATTCAGACAGTGGTCATCTACGAGAGTGCTTGTTGAAATGTGTCAAGGCGAGTGGCACATTTTTGGACATAAGCAAATCAGACATGAACGAAAACAAAGATTTCTGCATGGGGTTTATTCGAGGCAACAGATCGTATAAGGCTGCTGATTTATCTAACATTTTTAAGCCCGAATTCATTGAAGACAAAAAGATTTTACGAAACATGATAGCTCAAGGTATTGCAAAAGGTATTGTTAAGCCCTTAACTAGAGTTGTATATCCAACTATAGCAATTACGAAAGCCTTTAAACTTTTATCATCGAGTAAACACAAAGGAAAGGTTTTAGTTCAAATGAAGGACCCCAGCTTCAAAATTGAACACAACATCCCTCCAAGAATGAAATATTTACCAAATGCAGTTTATATAGTCGTCTGTGATGATAGCTATTTGGGTATTGAGTTTACTGATGATATTATCAAAAAGGGAGCCAGAAAAGTCACGGTCAATATGAATAATGAAATATCTGGATATGTTTATTCAAAATTAATGTCTTGGAAGAAGCATGCTGCAGCAGTCAACATTACTTCATATTGCTTAAATACCAAAGAAGGATGTACTCAACTGCTTAAGGATAGTGCCAAAATCGGACAAGTAAAAGGAGTATACATAGTTAGTGATTCAGAAAGAAGGGAAAAAGAAAACAATATTGAATTACTCGTGTCTAATATGGACGTTGTTTCCAGAAGTATTTGTAATGGACTAAGTAATTTCGTGTTGATCTCGAAGTATTCCCAAAGTGACACTTACGAATATCAAATGGGAACCGTTGAGAAAATATTTGAAGCGCGAAATGAAATGGCCCTAACAAGTCTAGTTGTACGCTACGACGGACTGGACACATTCACAGATGTTTCCCAAAACACGACTGAGATACACAGGTCGACATTATTTGATGCAGTGGAAAAGAGCATCAGATTAAACCATCAAAACGTATACGTCTACAAATTGAGAAACAAAAAACCTTCGGACTACCATCAGATGATCTCTAAAATACTTGGCATAAAAACTTTGGATGGTGCTGATGATACACACTTGATAAGTGACTTCAAATTGAATGAAATCAGTCTGACGGAGATGAAAAATTTAATAATGAATTATTTTAAAGTTGACTTTCCTGTTGAACAAATACTGTCAATGAGTCTTGAAAGTTTAAAAGATCTTGGTAAAAACAAAAAAAAACCGGACATCAATTTCAATAGTGGATTAGGAGCTTTCATTGCATGTGTCGACGATGAAGAGACACATGTTACTATGCCACATAATATTGTTTCTATGAAGACATTGCTTAATAAGAGTATTGAGAGTGATATCGACCCCGAAAATACCTTCATAACCTTGATCCCAGGGTTCGAAGGTCGTCACCAAATATTCGAAACTCTAGCTGAAACATTAAAAGTTAAAGCAGTTGCCGTGCATTTGAGTTCAGTCATAACTATGGATAGTATACCACAAATGGCTGTAGAAATTCGTAAGCTGATGAAAATGAAATTCAAAGTGAAACCAAAGTTCCACCTACTTGGGTATTCGTTTGGAGTTAATATTACTTTGGAATTAGCTGCAATTTTGGAAAAAGAAGGCCACATCGGAACTGTTTACTGTTTAGATAGCAGCCCGGATGCATTACGAGATCAATTAGAAGCCTACCTGGGCAATCCAGACGAAAAGGAACTCCAAAACTACATAATAAACTACATATACCAACTACTTACTAACCAGAATAGCCAATTAGAAAATACCCTATCAACACTAGATACTTGGGAAGAAAAACTCCATATCAGCACAAAAGAAATCAAAAAATATTCGAGCTTTTCTTACGAATACATAAGGGCTATTGTTGATGGTGCGTACAAAAGAATTGAACTTGCTAAAAATTACAAGCCTGATATAATTTTGGAATCCGAATTGGTTCTCATTAAAGGAGTACCTCATCCATCTTCAAGATTATTACCCGATGATTACAATTTGTCGAAGTACAGCAGTAAGCCTGTGAAAGTATACCAAATCGATACTGACCATTCGCTGGCCACTTATGATTGTAGAGTTTCTAATATTATCAATTCTCTTCTTGAAAATACTCTACTAGAAGATTACAAGAAAAATGATCTTTGCGATGTATACATGACTTGA
Protein
MLTTYTRGLVCTQSPLIDGSMAAVGLGYQSIARQCPPEIDIACHNSFESTTISGPAEVTKLFVDDLKKRGIFAKEVPCGNVAFHSRYIALAGQALLDALKKIIKEPKPRNEKWLSTSVPQHKWHELSAKYSSPEYHTNNMLSPVLFEEVTRLIPDNAIVIEIAPHGLLQAILKRSLQNCLHIPLTKRGTNDSVQFLLDAFGKMFEAGLNPKIEKLYPKIEFPVSTQTPLLSNLIEWEHSEDWLHLFSGSGVKCTAIARDVVISFYNDDYNFLKGHLRNGLNILPEAEILVLVWETMAMYKNMDYRYLSVMFKDVHFHREVEIKRDIPLRLSISITKGNNRFEVIHEDAIVVCGIFDESVIMKSDHGIDETKFNKDEILLNTEDIYQILELRGHCYTGEHKSIASTNLERNKAIVRWNNNWALYLDSLIQLNILANEHEGLSVPKFIGKIAISVSEHEKAMQKVNNLTDGHVANIIDTFDVTRCAGVEIDRLIFTEKAVINKEADGLQILSFCPQYPTEEISLQRAINVVLKIVAENSSSKKIKIIKSASLSIDASDTIEKIIQTEPMEVEVVTANIDNFHKKSNGINDELAEVNLVIIDNILNDKQKLAKIHETISKDTFILTFEANVQKNNLIKNYFNIITCMSSENGICILLKRIPLERVQRTCIPINYKTYKEHLLYVYEELKRTSVVLVSEGQSLCGISGLVKKLRRDGHKNISAMIIEDTDAPKFNPDHPFYKSQLDLDLAANVLRKRQWGGYYYIPSQMSASVRNMTLMSKQPGSIDGLTWVEAPATTDGAEHVQVWYAGLSLKDCQKAISCENNNRSYFGMDFSGYNSKGEKIMGLVREGSLSSSVEADPDLTWSVPEHWTLEEAATVPLAYLHAFYCLELRCDISSGKTVFITGGAGALGQAVISICLAMNCPIYTTVRDIHKKNFLLKLFPELPESHIGISHLEALYSMLIFNSENKLCDIIINSDSGHLRECLLKCVKASGTFLDISKSDMNENKDFCMGFIRGNRSYKAADLSNIFKPEFIEDKKILRNMIAQGIAKGIVKPLTRVVYPTIAITKAFKLLSSSKHKGKVLVQMKDPSFKIEHNIPPRMKYLPNAVYIVVCDDSYLGIEFTDDIIKKGARKVTVNMNNEISGYVYSKLMSWKKHAAAVNITSYCLNTKEGCTQLLKDSAKIGQVKGVYIVSDSERREKENNIELLVSNMDVVSRSICNGLSNFVLISKYSQSDTYEYQMGTVEKIFEARNEMALTSLVVRYDGLDTFTDVSQNTTEIHRSTLFDAVEKSIRLNHQNVYVYKLRNKKPSDYHQMISKILGIKTLDGADDTHLISDFKLNEISLTEMKNLIMNYFKVDFPVEQILSMSLESLKDLGKNKKKPDINFNSGLGAFIACVDDEETHVTMPHNIVSMKTLLNKSIESDIDPENTFITLIPGFEGRHQIFETLAETLKVKAVAVHLSSVITMDSIPQMAVEIRKLMKMKFKVKPKFHLLGYSFGVNITLELAAILEKEGHIGTVYCLDSSPDALRDQLEAYLGNPDEKELQNYIINYIYQLLTNQNSQLENTLSTLDTWEEKLHISTKEIKKYSSFSYEYIRAIVDGAYKRIELAKNYKPDIILESELVLIKGVPHPSSRLLPDDYNLSKYSSKPVKVYQIDTDHSLATYDCRVSNIINSLLENTLLEDYKKNDLCDVYMT
Summary
Uniprot
O01677
A0A3S2NJM7
H9JGF5
A0A2A4JVR3
A0A2H1V933
A0A2A4JTZ4
+ More
A0A2H1WZR1 A0A2A4K3Y5 A0A194PKD7 A0A2A4K4K9 A0A0N1I4Z6 A0A2H1VBX5 A0A2H1VPQ1 A0A3S2M8J0 A0A212FMU5 O01678 A0A2A4K5B4 A0A2A4J169 A0A194Q6Q7 A0A1B6JP24 A0A026WNV9 A0A1B6CGM1 A0A2H1W5A9 A0A232EXC2 Q16ZI9 E2ADN8 A0A1S4FIT0 B0WMC9 A0A1Q3FS82 A0A182GNP7 K7IZC8 A0A0J7NXT7 A0A195BMY6 K7IZC9 A0A0L7R328 A0A151X6P9 A0A158NZW8 A0A336L600 A0A182M3X3 A0A195F360 A0A084VRX8 A0A182VQY2 A0A087ZKN3 A0A182PC27 A0A182JQR8 A0A195CP13 A0A067RHL3 A0A2A4J2W6 A0A310SRI2 A0A182YJT7 F4WIF1 E2B583 A0A1J1IGX7 B4NIW4 B4LXF7 A0A182R1J1 A0A182QDS2 A0A182XBL3 A0A182FQK1 A0A182J9B8 A0A182V5F6 A0A182N5X9 A0A1J1IKA4 A0A182I0F0 Q7PYE4 A0A1W4VE27 A0A182TPE9 A0A0M4EN44 A0A3Q0IWX7 A0A0N1IP09 V5GRN6 A0A1B6IHH2 A0A1J1I2M2 A0A3B0KR31 A0A154PMH1 A0A336MXS7 B3N4Q4 A0A0R1E948 A0A0R1E8I8 A0A232EXB1 A0A1B2M4Y5 A0A1W5YLL8 A0A0G3YL52 J9K3D3 A0A034VP61 A0A034VKR9 A0A1W4XJ56 A0A1A9UN51 A0A0K8UFW4 A0A0A1XSP3 A0A0A1XIY0 T1P9U1 A0A2J7PQX7
A0A2H1WZR1 A0A2A4K3Y5 A0A194PKD7 A0A2A4K4K9 A0A0N1I4Z6 A0A2H1VBX5 A0A2H1VPQ1 A0A3S2M8J0 A0A212FMU5 O01678 A0A2A4K5B4 A0A2A4J169 A0A194Q6Q7 A0A1B6JP24 A0A026WNV9 A0A1B6CGM1 A0A2H1W5A9 A0A232EXC2 Q16ZI9 E2ADN8 A0A1S4FIT0 B0WMC9 A0A1Q3FS82 A0A182GNP7 K7IZC8 A0A0J7NXT7 A0A195BMY6 K7IZC9 A0A0L7R328 A0A151X6P9 A0A158NZW8 A0A336L600 A0A182M3X3 A0A195F360 A0A084VRX8 A0A182VQY2 A0A087ZKN3 A0A182PC27 A0A182JQR8 A0A195CP13 A0A067RHL3 A0A2A4J2W6 A0A310SRI2 A0A182YJT7 F4WIF1 E2B583 A0A1J1IGX7 B4NIW4 B4LXF7 A0A182R1J1 A0A182QDS2 A0A182XBL3 A0A182FQK1 A0A182J9B8 A0A182V5F6 A0A182N5X9 A0A1J1IKA4 A0A182I0F0 Q7PYE4 A0A1W4VE27 A0A182TPE9 A0A0M4EN44 A0A3Q0IWX7 A0A0N1IP09 V5GRN6 A0A1B6IHH2 A0A1J1I2M2 A0A3B0KR31 A0A154PMH1 A0A336MXS7 B3N4Q4 A0A0R1E948 A0A0R1E8I8 A0A232EXB1 A0A1B2M4Y5 A0A1W5YLL8 A0A0G3YL52 J9K3D3 A0A034VP61 A0A034VKR9 A0A1W4XJ56 A0A1A9UN51 A0A0K8UFW4 A0A0A1XSP3 A0A0A1XIY0 T1P9U1 A0A2J7PQX7
Pubmed
EMBL
U67866
AAB53257.1
RSAL01000068
RVE49259.1
BABH01033637
NWSH01000591
+ More
PCG75472.1 ODYU01001294 SOQ37311.1 PCG75471.1 ODYU01012325 SOQ58579.1 NWSH01000156 PCG78967.1 KQ459602 KPI93194.1 PCG78966.1 KQ461178 KPJ07769.1 ODYU01001484 SOQ37744.1 ODYU01003708 SOQ42815.1 RSAL01000013 RVE53445.1 AGBW02007651 OWR55047.1 U67867 AAB53258.1 PCG78963.1 NWSH01003873 PCG65731.1 KQ459582 KPI99075.1 GECU01006695 JAT01012.1 KK107139 EZA57700.1 GEDC01024813 JAS12485.1 ODYU01006431 SOQ48269.1 NNAY01001760 OXU23005.1 CH477489 EAT40085.1 GL438820 EFN68400.1 DS231997 EDS30986.1 GFDL01004709 JAV30336.1 JXUM01076840 KQ562972 KXJ74732.1 LBMM01000911 KMQ97230.1 KQ976440 KYM86527.1 KQ414663 KOC65238.1 KQ982480 KYQ55999.1 ADTU01005072 ADTU01005073 ADTU01005074 ADTU01005075 UFQS01001117 UFQT01001117 SSX09070.1 SSX28981.1 AXCM01001692 KQ981856 KYN34617.1 ATLV01015783 ATLV01015784 KE525036 KFB40722.1 KQ977574 KYN01849.1 KK852680 KDR18643.1 PCG65733.1 KQ761246 OAD57941.1 GL888175 EGI66082.1 GL445739 EFN89136.1 CVRI01000051 CRK99509.1 CH964272 EDW84866.1 CH940650 EDW67835.2 AXCN02000191 CRK99510.1 APCN01002072 AAAB01008987 EAA01044.4 CP012526 ALC46026.1 KQ460651 KPJ13003.1 GALX01004174 JAB64292.1 GECU01021338 JAS86368.1 CVRI01000038 CRK94451.1 OUUW01000014 SPP88346.1 KQ434960 KZC12654.1 UFQT01003008 SSX34421.1 CH954177 EDV60008.2 CH893166 KRK05689.1 KRK05688.1 OXU23004.1 KX467777 AOA60273.1 KX147671 ARI45075.1 KM507100 AKM28424.1 ABLF02036897 GAKP01015045 JAC43907.1 GAKP01015046 JAC43906.1 GDHF01026866 JAI25448.1 GBXI01000719 JAD13573.1 GBXI01003003 JAD11289.1 KA644718 AFP59347.1 NEVH01022635 PNF18741.1
PCG75472.1 ODYU01001294 SOQ37311.1 PCG75471.1 ODYU01012325 SOQ58579.1 NWSH01000156 PCG78967.1 KQ459602 KPI93194.1 PCG78966.1 KQ461178 KPJ07769.1 ODYU01001484 SOQ37744.1 ODYU01003708 SOQ42815.1 RSAL01000013 RVE53445.1 AGBW02007651 OWR55047.1 U67867 AAB53258.1 PCG78963.1 NWSH01003873 PCG65731.1 KQ459582 KPI99075.1 GECU01006695 JAT01012.1 KK107139 EZA57700.1 GEDC01024813 JAS12485.1 ODYU01006431 SOQ48269.1 NNAY01001760 OXU23005.1 CH477489 EAT40085.1 GL438820 EFN68400.1 DS231997 EDS30986.1 GFDL01004709 JAV30336.1 JXUM01076840 KQ562972 KXJ74732.1 LBMM01000911 KMQ97230.1 KQ976440 KYM86527.1 KQ414663 KOC65238.1 KQ982480 KYQ55999.1 ADTU01005072 ADTU01005073 ADTU01005074 ADTU01005075 UFQS01001117 UFQT01001117 SSX09070.1 SSX28981.1 AXCM01001692 KQ981856 KYN34617.1 ATLV01015783 ATLV01015784 KE525036 KFB40722.1 KQ977574 KYN01849.1 KK852680 KDR18643.1 PCG65733.1 KQ761246 OAD57941.1 GL888175 EGI66082.1 GL445739 EFN89136.1 CVRI01000051 CRK99509.1 CH964272 EDW84866.1 CH940650 EDW67835.2 AXCN02000191 CRK99510.1 APCN01002072 AAAB01008987 EAA01044.4 CP012526 ALC46026.1 KQ460651 KPJ13003.1 GALX01004174 JAB64292.1 GECU01021338 JAS86368.1 CVRI01000038 CRK94451.1 OUUW01000014 SPP88346.1 KQ434960 KZC12654.1 UFQT01003008 SSX34421.1 CH954177 EDV60008.2 CH893166 KRK05689.1 KRK05688.1 OXU23004.1 KX467777 AOA60273.1 KX147671 ARI45075.1 KM507100 AKM28424.1 ABLF02036897 GAKP01015045 JAC43907.1 GAKP01015046 JAC43906.1 GDHF01026866 JAI25448.1 GBXI01000719 JAD13573.1 GBXI01003003 JAD11289.1 KA644718 AFP59347.1 NEVH01022635 PNF18741.1
Proteomes
UP000283053
UP000005204
UP000218220
UP000053268
UP000053240
UP000007151
+ More
UP000053097 UP000215335 UP000008820 UP000000311 UP000002320 UP000069940 UP000249989 UP000002358 UP000036403 UP000078540 UP000053825 UP000075809 UP000005205 UP000075883 UP000078541 UP000030765 UP000075920 UP000075885 UP000075881 UP000078542 UP000027135 UP000076408 UP000007755 UP000008237 UP000183832 UP000007798 UP000008792 UP000075900 UP000075886 UP000076407 UP000069272 UP000075880 UP000075903 UP000075884 UP000075840 UP000007062 UP000192221 UP000075902 UP000092553 UP000079169 UP000268350 UP000076502 UP000008711 UP000002282 UP000007819 UP000192223 UP000078200 UP000235965
UP000053097 UP000215335 UP000008820 UP000000311 UP000002320 UP000069940 UP000249989 UP000002358 UP000036403 UP000078540 UP000053825 UP000075809 UP000005205 UP000075883 UP000078541 UP000030765 UP000075920 UP000075885 UP000075881 UP000078542 UP000027135 UP000076408 UP000007755 UP000008237 UP000183832 UP000007798 UP000008792 UP000075900 UP000075886 UP000076407 UP000069272 UP000075880 UP000075903 UP000075884 UP000075840 UP000007062 UP000192221 UP000075902 UP000092553 UP000079169 UP000268350 UP000076502 UP000008711 UP000002282 UP000007819 UP000192223 UP000078200 UP000235965
Pfam
Interpro
IPR018201
Ketoacyl_synth_AS
+ More
IPR001031 Thioesterase
IPR014043 Acyl_transferase
IPR036291 NAD(P)-bd_dom_sf
IPR032821 KAsynt_C_assoc
IPR042104 PKS_dehydratase_sf
IPR011032 GroES-like_sf
IPR014031 Ketoacyl_synth_C
IPR023102 Fatty_acid_synthase_dom_2
IPR016035 Acyl_Trfase/lysoPLipase
IPR016036 Malonyl_transacylase_ACP-bd
IPR020807 PKS_dehydratase
IPR001227 Ac_transferase_dom_sf
IPR029058 AB_hydrolase
IPR020841 PKS_Beta-ketoAc_synthase_dom
IPR020801 PKS_acyl_transferase
IPR016039 Thiolase-like
IPR014030 Ketoacyl_synth_N
IPR020843 PKS_ER
IPR013149 ADH_C
IPR013968 PKS_KR
IPR036736 ACP-like_sf
IPR020806 PKS_PP-bd
IPR009081 PP-bd_ACP
IPR006162 Ppantetheine_attach_site
IPR001031 Thioesterase
IPR014043 Acyl_transferase
IPR036291 NAD(P)-bd_dom_sf
IPR032821 KAsynt_C_assoc
IPR042104 PKS_dehydratase_sf
IPR011032 GroES-like_sf
IPR014031 Ketoacyl_synth_C
IPR023102 Fatty_acid_synthase_dom_2
IPR016035 Acyl_Trfase/lysoPLipase
IPR016036 Malonyl_transacylase_ACP-bd
IPR020807 PKS_dehydratase
IPR001227 Ac_transferase_dom_sf
IPR029058 AB_hydrolase
IPR020841 PKS_Beta-ketoAc_synthase_dom
IPR020801 PKS_acyl_transferase
IPR016039 Thiolase-like
IPR014030 Ketoacyl_synth_N
IPR020843 PKS_ER
IPR013149 ADH_C
IPR013968 PKS_KR
IPR036736 ACP-like_sf
IPR020806 PKS_PP-bd
IPR009081 PP-bd_ACP
IPR006162 Ppantetheine_attach_site
SUPFAM
ProteinModelPortal
O01677
A0A3S2NJM7
H9JGF5
A0A2A4JVR3
A0A2H1V933
A0A2A4JTZ4
+ More
A0A2H1WZR1 A0A2A4K3Y5 A0A194PKD7 A0A2A4K4K9 A0A0N1I4Z6 A0A2H1VBX5 A0A2H1VPQ1 A0A3S2M8J0 A0A212FMU5 O01678 A0A2A4K5B4 A0A2A4J169 A0A194Q6Q7 A0A1B6JP24 A0A026WNV9 A0A1B6CGM1 A0A2H1W5A9 A0A232EXC2 Q16ZI9 E2ADN8 A0A1S4FIT0 B0WMC9 A0A1Q3FS82 A0A182GNP7 K7IZC8 A0A0J7NXT7 A0A195BMY6 K7IZC9 A0A0L7R328 A0A151X6P9 A0A158NZW8 A0A336L600 A0A182M3X3 A0A195F360 A0A084VRX8 A0A182VQY2 A0A087ZKN3 A0A182PC27 A0A182JQR8 A0A195CP13 A0A067RHL3 A0A2A4J2W6 A0A310SRI2 A0A182YJT7 F4WIF1 E2B583 A0A1J1IGX7 B4NIW4 B4LXF7 A0A182R1J1 A0A182QDS2 A0A182XBL3 A0A182FQK1 A0A182J9B8 A0A182V5F6 A0A182N5X9 A0A1J1IKA4 A0A182I0F0 Q7PYE4 A0A1W4VE27 A0A182TPE9 A0A0M4EN44 A0A3Q0IWX7 A0A0N1IP09 V5GRN6 A0A1B6IHH2 A0A1J1I2M2 A0A3B0KR31 A0A154PMH1 A0A336MXS7 B3N4Q4 A0A0R1E948 A0A0R1E8I8 A0A232EXB1 A0A1B2M4Y5 A0A1W5YLL8 A0A0G3YL52 J9K3D3 A0A034VP61 A0A034VKR9 A0A1W4XJ56 A0A1A9UN51 A0A0K8UFW4 A0A0A1XSP3 A0A0A1XIY0 T1P9U1 A0A2J7PQX7
A0A2H1WZR1 A0A2A4K3Y5 A0A194PKD7 A0A2A4K4K9 A0A0N1I4Z6 A0A2H1VBX5 A0A2H1VPQ1 A0A3S2M8J0 A0A212FMU5 O01678 A0A2A4K5B4 A0A2A4J169 A0A194Q6Q7 A0A1B6JP24 A0A026WNV9 A0A1B6CGM1 A0A2H1W5A9 A0A232EXC2 Q16ZI9 E2ADN8 A0A1S4FIT0 B0WMC9 A0A1Q3FS82 A0A182GNP7 K7IZC8 A0A0J7NXT7 A0A195BMY6 K7IZC9 A0A0L7R328 A0A151X6P9 A0A158NZW8 A0A336L600 A0A182M3X3 A0A195F360 A0A084VRX8 A0A182VQY2 A0A087ZKN3 A0A182PC27 A0A182JQR8 A0A195CP13 A0A067RHL3 A0A2A4J2W6 A0A310SRI2 A0A182YJT7 F4WIF1 E2B583 A0A1J1IGX7 B4NIW4 B4LXF7 A0A182R1J1 A0A182QDS2 A0A182XBL3 A0A182FQK1 A0A182J9B8 A0A182V5F6 A0A182N5X9 A0A1J1IKA4 A0A182I0F0 Q7PYE4 A0A1W4VE27 A0A182TPE9 A0A0M4EN44 A0A3Q0IWX7 A0A0N1IP09 V5GRN6 A0A1B6IHH2 A0A1J1I2M2 A0A3B0KR31 A0A154PMH1 A0A336MXS7 B3N4Q4 A0A0R1E948 A0A0R1E8I8 A0A232EXB1 A0A1B2M4Y5 A0A1W5YLL8 A0A0G3YL52 J9K3D3 A0A034VP61 A0A034VKR9 A0A1W4XJ56 A0A1A9UN51 A0A0K8UFW4 A0A0A1XSP3 A0A0A1XIY0 T1P9U1 A0A2J7PQX7
PDB
6FN6
E-value=1.37823e-80,
Score=769
Ontologies
GO
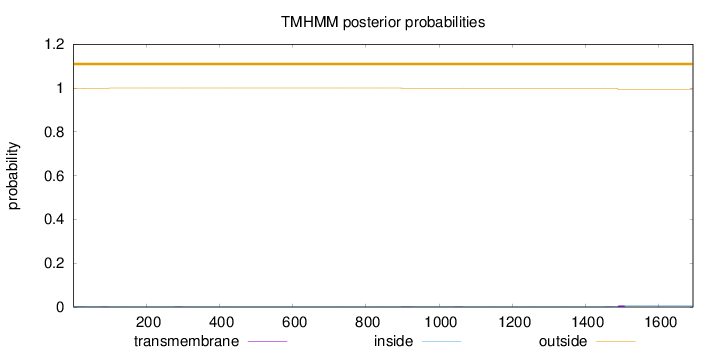
Topology
Length:
1694
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.20284
Exp number, first 60 AAs:
0.02346
Total prob of N-in:
0.00175
outside
1 - 1694
Population Genetic Test Statistics
Pi
19.556474
Theta
20.967769
Tajima's D
-0.918602
CLR
14.452658
CSRT
0.150192490375481
Interpretation
Uncertain
