Gene
KWMTBOMO10859
Annotation
PREDICTED:_uncharacterized_protein_LOC105396011_[Plutella_xylostella]
Location in the cell
Nuclear Reliability : 3.167
Sequence
CDS
ATGTCCTGTCGCTTATGTAGAAGACGGCATCACACTCTTCTCCACCGCCATCAAGACTCGGTACAAACTGAGGAAAGGAATCAACCGAATGCATCTCACCACGGGGTAGAAGAGAAAGAAGAGGTTATAAATAGCATGGTGGCGTCACATCACAATACTAAACAAAGCACAGCACTACTAGCCACAGCGATGGTTTTAGCGAGGAACGAGCATAACCAAACCGTTGTCCTTCGTGCGCTAGTAGACCAAGGTTCGCAGGCTTCATTTTTAAGCGAGAAGGCGGCTCAAATGCTCAAACTTACCAGGCAATCAGCGAGAGGAAGTATCATCGGGGTGGGATCTACTAGGACTAACGTTGATCACGTGGTTCAACTTAAGATTGGCTCCCACTGGAATCCAAGCTTCGAAATCAACATACAAGCATATGTTATGAGTAAACAGTTGACCACGAAGATCCCCTCCAAAGCTGTAACTGTCACACATTGGCCACATCTAGAAGGACTCAACCTGGCAGATCCAGACTACTTTCAACCAGGTTCTATAGACTTACTTCTTGGTGTTAAAGAGTATGCACAGATTGTGCAACCAACACTTATTAAGGGCCCTCCAGGTTCTCCATGTGCCCAAGAAACCAACCTGGGGTGGATCCTCTTCGGAGAAATCGATAATAACCGTGAAAATGAATCATTTCTGGTCCTTCATCATCAAGTCCATATAGATCATATGTTGAAATCCCTCTGGGAAATTGACAATGATAAAAAGAGAATATTAACAAAGGAAGAAAAGCTATGTGAAACAATATATGAAAGCACACATACGAGAAATAGTGAAGGAAGATATCTAGTAAGGCTGCCATTTAATACAAAGAATCCAATTTCGACTGAAGGAAATACGAAAGAAATAGCTAGAAAGAGGCTACTTCAGCTAGAAAGAAGATTTAAGAAAGAAACTAAACTGAAAGAAGATTATACAAAGGTCATGCAAGAATACATAAAAGCTAACTACATGGAAGAAATATCTAAGAATGAAATAAATGAGAGAAAATCAGTATACTTACCTCATCGCGCCGTGGTACACGCAGACAGAGAAACTTCTAAGACACGCGTAGTATTTGATGCTTCCGCGAAGGGCTCCAACTGCGTTTCTCTAAACGATGAATTGCTTGTAGGTCCGCAACTGCAAGAGGACTTAAGGAACATACTTATGAGATCGAGATTACATCGTGTTTGCTATGCAAGTGATATATAA
Protein
MSCRLCRRRHHTLLHRHQDSVQTEERNQPNASHHGVEEKEEVINSMVASHHNTKQSTALLATAMVLARNEHNQTVVLRALVDQGSQASFLSEKAAQMLKLTRQSARGSIIGVGSTRTNVDHVVQLKIGSHWNPSFEINIQAYVMSKQLTTKIPSKAVTVTHWPHLEGLNLADPDYFQPGSIDLLLGVKEYAQIVQPTLIKGPPGSPCAQETNLGWILFGEIDNNRENESFLVLHHQVHIDHMLKSLWEIDNDKKRILTKEEKLCETIYESTHTRNSEGRYLVRLPFNTKNPISTEGNTKEIARKRLLQLERRFKKETKLKEDYTKVMQEYIKANYMEEISKNEINERKSVYLPHRAVVHADRETSKTRVVFDASAKGSNCVSLNDELLVGPQLQEDLRNILMRSRLHRVCYASDI
Summary
Uniprot
A0A2H1VGJ3
A0A0L7LDY3
A0A3S2L012
A0A0L7LMN3
A0A194QR21
A0A2H1WH28
+ More
A0A0N0PAG9 A0A0N1IHU1 A0A0L7LFG6 W8B1K7 A0A0J7KFC5 A0A1I8M1N7 J9JVD6 A0A226CWA5 A0A226DS81 A0A226DBN6 A0A226E226 W8CCT3 A0A226DMG9 A0A226EAM0 J9LVB4 A0A0J7K9F4 X1WY64 A0A0J7KCX3 A0A2S2PM34 A0A0J7KDE0 A0A1B0C873 A0A151IHU8 A0A0J7N890 A0A0J7N314 A0A034WA88 A0A0J7KDP8 V5GHM7 A0A151IIY3 A0A0J7KNI7 A0A0E4G8R0 A0A2H1WI74 A0A3L8DW93 A0A0J7KGI8 A0A0J7MWB1 A0A151IT82 A0A226DKA3 A0A2A4JL65 A0A0J7N465 A0A3L8DF49 A0A0J7K7B9 A0A1W4VKF9 J9KD60 J9LPA9 A0A226CWQ4 A0A0J7KB34 A0A0J7K7U0 A0A0J7K8M4 A0A226D417 A0A0L7L797 A0A0J7N1U6 A0A1Y1KV73 A0A2S2PWK4 A0A2S2Q0A5 A0A2S2PBM2 A0A2S2PKV3 A0A226D0T3 A0A226E0X4 X1XTX7 V5H006 A0A226EM86 A0A226EUK6 A0A0J7K6B3 X1WY95 A0A1W7R6J0 A0A226EUF5 A0A2A4K6T9 A0A2W1BMV4 A0A182H647 X1X096 A0A0J7K2G5 A0A0J7KCT8 A0A1W7R6N3 A0A0J7KDN6 A0A226E780 A0A0J7K314 A0A1B0D852 A0A0J7K9A7 A0A1Y1LE43 J9K774 A0A226EPI5 A0A1Y1LBZ3 A0A355B931 A0A151IM38 A0A0J7KBS8 A0A2S2NCY5 A0A2S2NCJ9 A0A0J7K907 A0A0J7N2B8 A0A0J7N046 X1XU26 A0A2W1BJC9 A0A0J7KDH1
A0A0N0PAG9 A0A0N1IHU1 A0A0L7LFG6 W8B1K7 A0A0J7KFC5 A0A1I8M1N7 J9JVD6 A0A226CWA5 A0A226DS81 A0A226DBN6 A0A226E226 W8CCT3 A0A226DMG9 A0A226EAM0 J9LVB4 A0A0J7K9F4 X1WY64 A0A0J7KCX3 A0A2S2PM34 A0A0J7KDE0 A0A1B0C873 A0A151IHU8 A0A0J7N890 A0A0J7N314 A0A034WA88 A0A0J7KDP8 V5GHM7 A0A151IIY3 A0A0J7KNI7 A0A0E4G8R0 A0A2H1WI74 A0A3L8DW93 A0A0J7KGI8 A0A0J7MWB1 A0A151IT82 A0A226DKA3 A0A2A4JL65 A0A0J7N465 A0A3L8DF49 A0A0J7K7B9 A0A1W4VKF9 J9KD60 J9LPA9 A0A226CWQ4 A0A0J7KB34 A0A0J7K7U0 A0A0J7K8M4 A0A226D417 A0A0L7L797 A0A0J7N1U6 A0A1Y1KV73 A0A2S2PWK4 A0A2S2Q0A5 A0A2S2PBM2 A0A2S2PKV3 A0A226D0T3 A0A226E0X4 X1XTX7 V5H006 A0A226EM86 A0A226EUK6 A0A0J7K6B3 X1WY95 A0A1W7R6J0 A0A226EUF5 A0A2A4K6T9 A0A2W1BMV4 A0A182H647 X1X096 A0A0J7K2G5 A0A0J7KCT8 A0A1W7R6N3 A0A0J7KDN6 A0A226E780 A0A0J7K314 A0A1B0D852 A0A0J7K9A7 A0A1Y1LE43 J9K774 A0A226EPI5 A0A1Y1LBZ3 A0A355B931 A0A151IM38 A0A0J7KBS8 A0A2S2NCY5 A0A2S2NCJ9 A0A0J7K907 A0A0J7N2B8 A0A0J7N046 X1XU26 A0A2W1BJC9 A0A0J7KDH1
Pubmed
EMBL
ODYU01002449
SOQ39955.1
JTDY01001518
KOB73672.1
RSAL01000327
RVE42512.1
+ More
JTDY01000554 KOB76692.1 KQ461175 KPJ07794.1 ODYU01008608 SOQ52337.1 KQ458473 KPJ05885.1 KQ460036 KPJ18393.1 JTDY01001325 KOB74192.1 GAMC01011599 GAMC01011595 JAB94960.1 LBMM01008324 KMQ88972.1 ABLF02008594 LNIX01000069 OXA36894.1 LNIX01000012 OXA48073.1 LNIX01000026 OXA42254.1 LNIX01000007 OXA51795.1 GAMC01003146 JAC03410.1 LNIX01000016 OXA46198.1 LNIX01000005 OXA54553.1 OXA54559.1 ABLF02004142 LBMM01011423 KMQ86851.1 ABLF02012239 ABLF02012241 ABLF02017372 ABLF02042178 ABLF02066436 LBMM01009210 KMQ88303.1 GGMR01017890 MBY30509.1 LBMM01009331 KMQ88201.1 AJWK01000239 AJWK01000240 KQ977557 KYN02013.1 LBMM01008464 KMQ88855.1 LBMM01011051 KMQ87060.1 GAKP01007887 JAC51065.1 LBMM01008872 KMQ88568.1 GALX01004947 JAB63519.1 KQ977438 KYN02826.1 LBMM01005112 KMQ91804.1 HACL01000129 CFW94423.1 ODYU01008829 SOQ52773.1 QOIP01000004 RLU24048.1 LBMM01007859 KMQ89359.1 LBMM01015575 KMQ84755.1 KQ981026 KYN10209.1 LNIX01000018 OXA45284.1 NWSH01001201 PCG72150.1 LBMM01010417 KMQ87475.1 QOIP01000009 RLU19024.1 LBMM01012568 KMQ86144.1 ABLF02041863 ABLF02035780 LNIX01000063 OXA37200.1 LBMM01010405 KMQ87487.1 LBMM01012238 KMQ86334.1 LBMM01011781 KMQ86634.1 LNIX01000036 OXA39969.1 JTDY01002583 KOB71156.1 LBMM01011810 KMQ86610.1 GEZM01078370 GEZM01078366 GEZM01078363 JAV62757.1 GGMS01000724 MBY69927.1 GGMS01001896 MBY71099.1 GGMR01014230 MBY26849.1 GGMR01017408 MBY30027.1 LNIX01000044 OXA38660.1 OXA51382.1 ABLF02006304 ABLF02036268 GALX01002328 JAB66138.1 LNIX01000003 OXA58579.1 LNIX01000001 OXA61293.1 LBMM01013154 KMQ85784.1 ABLF02030854 ABLF02067347 GEHC01000877 JAV46768.1 LNIX01000002 OXA60848.1 NWSH01000070 PCG79985.1 KZ149989 PZC75621.1 JXUM01113346 KQ565681 KXJ70776.1 ABLF02008032 ABLF02035293 ABLF02060940 LBMM01016259 KMQ84489.1 LBMM01009284 KMQ88248.1 GEHC01000865 JAV46780.1 LBMM01009202 KMQ88311.1 OXA53452.1 LBMM01016077 KMQ84566.1 AJVK01036816 LBMM01011283 KMQ86917.1 GEZM01060253 JAV71158.1 ABLF02023723 OXA59522.1 GEZM01060254 JAV71154.1 DNYX01000039 HBK82233.1 KQ977080 KYN05933.1 LBMM01009888 KMQ87813.1 GGMR01002421 MBY15040.1 GGMR01002219 MBY14838.1 LBMM01011516 KMQ86784.1 LBMM01011492 KMQ86800.1 LBMM01012708 KMQ86040.1 ABLF02001506 ABLF02008029 ABLF02035899 ABLF02037719 ABLF02046327 ABLF02064627 KZ150008 PZC75149.1 LBMM01008962 KMQ88503.1
JTDY01000554 KOB76692.1 KQ461175 KPJ07794.1 ODYU01008608 SOQ52337.1 KQ458473 KPJ05885.1 KQ460036 KPJ18393.1 JTDY01001325 KOB74192.1 GAMC01011599 GAMC01011595 JAB94960.1 LBMM01008324 KMQ88972.1 ABLF02008594 LNIX01000069 OXA36894.1 LNIX01000012 OXA48073.1 LNIX01000026 OXA42254.1 LNIX01000007 OXA51795.1 GAMC01003146 JAC03410.1 LNIX01000016 OXA46198.1 LNIX01000005 OXA54553.1 OXA54559.1 ABLF02004142 LBMM01011423 KMQ86851.1 ABLF02012239 ABLF02012241 ABLF02017372 ABLF02042178 ABLF02066436 LBMM01009210 KMQ88303.1 GGMR01017890 MBY30509.1 LBMM01009331 KMQ88201.1 AJWK01000239 AJWK01000240 KQ977557 KYN02013.1 LBMM01008464 KMQ88855.1 LBMM01011051 KMQ87060.1 GAKP01007887 JAC51065.1 LBMM01008872 KMQ88568.1 GALX01004947 JAB63519.1 KQ977438 KYN02826.1 LBMM01005112 KMQ91804.1 HACL01000129 CFW94423.1 ODYU01008829 SOQ52773.1 QOIP01000004 RLU24048.1 LBMM01007859 KMQ89359.1 LBMM01015575 KMQ84755.1 KQ981026 KYN10209.1 LNIX01000018 OXA45284.1 NWSH01001201 PCG72150.1 LBMM01010417 KMQ87475.1 QOIP01000009 RLU19024.1 LBMM01012568 KMQ86144.1 ABLF02041863 ABLF02035780 LNIX01000063 OXA37200.1 LBMM01010405 KMQ87487.1 LBMM01012238 KMQ86334.1 LBMM01011781 KMQ86634.1 LNIX01000036 OXA39969.1 JTDY01002583 KOB71156.1 LBMM01011810 KMQ86610.1 GEZM01078370 GEZM01078366 GEZM01078363 JAV62757.1 GGMS01000724 MBY69927.1 GGMS01001896 MBY71099.1 GGMR01014230 MBY26849.1 GGMR01017408 MBY30027.1 LNIX01000044 OXA38660.1 OXA51382.1 ABLF02006304 ABLF02036268 GALX01002328 JAB66138.1 LNIX01000003 OXA58579.1 LNIX01000001 OXA61293.1 LBMM01013154 KMQ85784.1 ABLF02030854 ABLF02067347 GEHC01000877 JAV46768.1 LNIX01000002 OXA60848.1 NWSH01000070 PCG79985.1 KZ149989 PZC75621.1 JXUM01113346 KQ565681 KXJ70776.1 ABLF02008032 ABLF02035293 ABLF02060940 LBMM01016259 KMQ84489.1 LBMM01009284 KMQ88248.1 GEHC01000865 JAV46780.1 LBMM01009202 KMQ88311.1 OXA53452.1 LBMM01016077 KMQ84566.1 AJVK01036816 LBMM01011283 KMQ86917.1 GEZM01060253 JAV71158.1 ABLF02023723 OXA59522.1 GEZM01060254 JAV71154.1 DNYX01000039 HBK82233.1 KQ977080 KYN05933.1 LBMM01009888 KMQ87813.1 GGMR01002421 MBY15040.1 GGMR01002219 MBY14838.1 LBMM01011516 KMQ86784.1 LBMM01011492 KMQ86800.1 LBMM01012708 KMQ86040.1 ABLF02001506 ABLF02008029 ABLF02035899 ABLF02037719 ABLF02046327 ABLF02064627 KZ150008 PZC75149.1 LBMM01008962 KMQ88503.1
Proteomes
Pfam
Interpro
IPR021109
Peptidase_aspartic_dom_sf
+ More
IPR008737 Peptidase_asp_put
IPR008042 Retrotrans_Pao
IPR000253 FHA_dom
IPR000742 EGF-like_dom
IPR011042 6-blade_b-propeller_TolB-like
IPR013032 EGF-like_CS
IPR005312 DUF1759
IPR000033 LDLR_classB_rpt
IPR040676 DUF5641
IPR001584 Integrase_cat-core
IPR012337 RNaseH-like_sf
IPR001878 Znf_CCHC
IPR041588 Integrase_H2C2
IPR036397 RNaseH_sf
IPR011993 PH-like_dom_sf
IPR041681 PH_9
IPR002017 Spectrin_repeat
IPR018159 Spectrin/alpha-actinin
IPR001995 Peptidase_A2_cat
IPR000477 RT_dom
IPR036875 Znf_CCHC_sf
IPR000331 Rap_GAP_dom
IPR035974 Rap/Ran-GAP_sf
IPR000425 MIP
IPR023271 Aquaporin-like
IPR022048 Envelope_fusion-like
IPR029526 PGBD
IPR001969 Aspartic_peptidase_AS
IPR008737 Peptidase_asp_put
IPR008042 Retrotrans_Pao
IPR000253 FHA_dom
IPR000742 EGF-like_dom
IPR011042 6-blade_b-propeller_TolB-like
IPR013032 EGF-like_CS
IPR005312 DUF1759
IPR000033 LDLR_classB_rpt
IPR040676 DUF5641
IPR001584 Integrase_cat-core
IPR012337 RNaseH-like_sf
IPR001878 Znf_CCHC
IPR041588 Integrase_H2C2
IPR036397 RNaseH_sf
IPR011993 PH-like_dom_sf
IPR041681 PH_9
IPR002017 Spectrin_repeat
IPR018159 Spectrin/alpha-actinin
IPR001995 Peptidase_A2_cat
IPR000477 RT_dom
IPR036875 Znf_CCHC_sf
IPR000331 Rap_GAP_dom
IPR035974 Rap/Ran-GAP_sf
IPR000425 MIP
IPR023271 Aquaporin-like
IPR022048 Envelope_fusion-like
IPR029526 PGBD
IPR001969 Aspartic_peptidase_AS
SUPFAM
ProteinModelPortal
A0A2H1VGJ3
A0A0L7LDY3
A0A3S2L012
A0A0L7LMN3
A0A194QR21
A0A2H1WH28
+ More
A0A0N0PAG9 A0A0N1IHU1 A0A0L7LFG6 W8B1K7 A0A0J7KFC5 A0A1I8M1N7 J9JVD6 A0A226CWA5 A0A226DS81 A0A226DBN6 A0A226E226 W8CCT3 A0A226DMG9 A0A226EAM0 J9LVB4 A0A0J7K9F4 X1WY64 A0A0J7KCX3 A0A2S2PM34 A0A0J7KDE0 A0A1B0C873 A0A151IHU8 A0A0J7N890 A0A0J7N314 A0A034WA88 A0A0J7KDP8 V5GHM7 A0A151IIY3 A0A0J7KNI7 A0A0E4G8R0 A0A2H1WI74 A0A3L8DW93 A0A0J7KGI8 A0A0J7MWB1 A0A151IT82 A0A226DKA3 A0A2A4JL65 A0A0J7N465 A0A3L8DF49 A0A0J7K7B9 A0A1W4VKF9 J9KD60 J9LPA9 A0A226CWQ4 A0A0J7KB34 A0A0J7K7U0 A0A0J7K8M4 A0A226D417 A0A0L7L797 A0A0J7N1U6 A0A1Y1KV73 A0A2S2PWK4 A0A2S2Q0A5 A0A2S2PBM2 A0A2S2PKV3 A0A226D0T3 A0A226E0X4 X1XTX7 V5H006 A0A226EM86 A0A226EUK6 A0A0J7K6B3 X1WY95 A0A1W7R6J0 A0A226EUF5 A0A2A4K6T9 A0A2W1BMV4 A0A182H647 X1X096 A0A0J7K2G5 A0A0J7KCT8 A0A1W7R6N3 A0A0J7KDN6 A0A226E780 A0A0J7K314 A0A1B0D852 A0A0J7K9A7 A0A1Y1LE43 J9K774 A0A226EPI5 A0A1Y1LBZ3 A0A355B931 A0A151IM38 A0A0J7KBS8 A0A2S2NCY5 A0A2S2NCJ9 A0A0J7K907 A0A0J7N2B8 A0A0J7N046 X1XU26 A0A2W1BJC9 A0A0J7KDH1
A0A0N0PAG9 A0A0N1IHU1 A0A0L7LFG6 W8B1K7 A0A0J7KFC5 A0A1I8M1N7 J9JVD6 A0A226CWA5 A0A226DS81 A0A226DBN6 A0A226E226 W8CCT3 A0A226DMG9 A0A226EAM0 J9LVB4 A0A0J7K9F4 X1WY64 A0A0J7KCX3 A0A2S2PM34 A0A0J7KDE0 A0A1B0C873 A0A151IHU8 A0A0J7N890 A0A0J7N314 A0A034WA88 A0A0J7KDP8 V5GHM7 A0A151IIY3 A0A0J7KNI7 A0A0E4G8R0 A0A2H1WI74 A0A3L8DW93 A0A0J7KGI8 A0A0J7MWB1 A0A151IT82 A0A226DKA3 A0A2A4JL65 A0A0J7N465 A0A3L8DF49 A0A0J7K7B9 A0A1W4VKF9 J9KD60 J9LPA9 A0A226CWQ4 A0A0J7KB34 A0A0J7K7U0 A0A0J7K8M4 A0A226D417 A0A0L7L797 A0A0J7N1U6 A0A1Y1KV73 A0A2S2PWK4 A0A2S2Q0A5 A0A2S2PBM2 A0A2S2PKV3 A0A226D0T3 A0A226E0X4 X1XTX7 V5H006 A0A226EM86 A0A226EUK6 A0A0J7K6B3 X1WY95 A0A1W7R6J0 A0A226EUF5 A0A2A4K6T9 A0A2W1BMV4 A0A182H647 X1X096 A0A0J7K2G5 A0A0J7KCT8 A0A1W7R6N3 A0A0J7KDN6 A0A226E780 A0A0J7K314 A0A1B0D852 A0A0J7K9A7 A0A1Y1LE43 J9K774 A0A226EPI5 A0A1Y1LBZ3 A0A355B931 A0A151IM38 A0A0J7KBS8 A0A2S2NCY5 A0A2S2NCJ9 A0A0J7K907 A0A0J7N2B8 A0A0J7N046 X1XU26 A0A2W1BJC9 A0A0J7KDH1
Ontologies
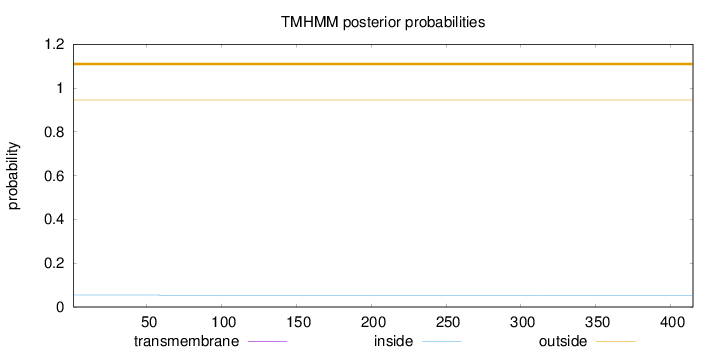
Topology
Length:
415
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.00151
Exp number, first 60 AAs:
0.00022
Total prob of N-in:
0.05426
outside
1 - 415
Population Genetic Test Statistics
Pi
415.575072
Theta
232.698516
Tajima's D
2.437217
CLR
0.001778
CSRT
0.938203089845508
Interpretation
Uncertain
