Gene
KWMTBOMO10756
Pre Gene Modal
BGIBMGA008418
Annotation
PREDICTED:_indole-3-acetaldehyde_oxidase-like_[Bombyx_mori]
Location in the cell
Cytoplasmic Reliability : 1.5
Sequence
CDS
ATGCCAAGAGCCCAGAATGCTCATGCGCAAGTGAACACCGGTTTTCTTTATGAATTTAATGCAAAAAATAAAGGTGTAATCGATTCGGCTCGAATAGTGATTGGCGGTTTGTCAGCACATTTCGTTCACGCAAAGCAAACTGAGAAGTACCTCAAAGGGAAAGACGTGTTTACCAATGAAGTTCTGCAGCACGCTCTGATGGTGTTGAAAGGGGAATTAATGGTGGATGAAATTCCGGGAGAAATGAGCCCCGAATTCAGAAAGAAGCTGGCCCTGGGTTTGTTTTACAAAGGACTGTTAAATCTTATTCCGAAGAAAACGATAAACCCTCGTTACCGCTCAGGAGCCAGAGATTTTCGGAAAACTCGTCCACTTTCGAAAGGCTCCGAAGTTTACGATACGAATCCCATAATATGGCCACTGACTGAACCAATGCCGAAGGTCGATGCTCTCGTGCAATGTGCGGGAGAAGCTCAATACGTGAATGACTTACCGAAGCAGACGGCTGAAGTATACTGTGCGTTTGTGACGTCTGATGTCGGAACCGGGGAGATAGTAGAAATCGATCCTACACCTGCTCTTAAAATTCCAGGCGTGCTAGCATTCTTTTCAGCGAAAGACATACCGGGCAAGAATAATTTCTTATCCCAACGAGTACCGACTCAACTGTTCCCCGAAGAGATATTCGCAGAAAAAACGATAAAATATTACGATCAAGCCATAGGTCTGATCGTGGCTGAAACTGAACAGCTGGCAAACAGAGCAGCCCTACTCGTACATGTCAAATACAAAGTGGACAGAAAGCCCATACTAACTATTCACGATGCTAGAGCAAAAGACCCAAGTCGAGTCAAACTTTTCATAATAATTCCTCCTAGAGACAGGGGAACGAATGTGCAGAGGGTCCTCAAAGGCTCAGACACTATTTTAGGCCAATACCACTTTCACATGGAGAACCTATCGTGCGTTACGAGACCAAGCGAAGACGGATTGGATGTATTTGTGGCAACGCAATGGACAGATTCAGTTCACATCGGTATTTCGGAGAGCTTAAACATAGAGCAAAACAGGATTAATCTGACAGTACCTAGATGTGGAGGCGGGTACGGGGCTAAGATATCGCGCGCGTCTCACGTGGCCTGCGCATGCGCCTTGGTCACTCACCTCATGGATAGGCCGTGTCGCTTCGTGATGAGCATTCAGGCTAACCTCAGAGTGATCGGGAAACGACTGCCCAGTCAGAGAGATTTTGAGGTTGGACTCAGTGATTCCGGTGAAATCCAGTATATGGAGTACCAGATATACGAAGATAACGGCTACACCGTGTCGGACGCCGTAATGCCTTTCCTTATCGGGGGGGTCAAGAACTGCTACGATAACAGAAGATGGCAGTCCAAGTTGTTCAATGTGGTTACTGATACCGCGTCGAATATTTATGCGAGAGCACCAGGTTCCCTGGAAGCCATATCCATGGCTGAATATTTAATGGAGCGGATATCGTACGAACTGGAACGTGATCCTATCGAAGTTCGTCTCAATAATCTTGATCCTCAGTACACAGAAGTAGCTGAAGTCGTTCAAACACTCCTGCGAGATAGCGAATACCACAAACGCAAAGAAGAAGTCGAGAAATTTAACAAGGAGAATAGATGGAAGAAACGAGGTCTGAGATTAGCTATGATGAACTGGCCGGCTTCTTCGGTCGTAGACTATCACGTTTTGTTGTCTGTGTACCACGGAGACGGCACGGTCGTGGTCACACATGGCGGTATTGAAATAGGACAAGGGATCAATACAAAAGTCATCCAGGCAGTCGCTTACACACTGAAAATTTCCACTGATAAGGTAATATGTAAACCTCCGACTGTCGCTACGAACCCTAACAACTTCACGACTGGAGGCAGCAGAACCACGCAATCCGTCTGCTTCGGTGCTATTAAATGCTGCCAAATTCTTCTCGACCGCTTGTCAGTTGTTCGGGAGGCCCTAAATGACCCGACCTGGGAAATACTTATCGAAGCCGCGTTCAATAGAGGCATCAATCTGCAGACCAGTTACCGCGTGACGCCAAACGATCAAGATCCGTATAGATGCGCTGGAGCAGCCGTTGCTGAGGTAGAGCTGGATGTCATAACGGGAGAACATGAGATACTGAGAGTTGATATCATTGAAGATGTAGGGACCAGTATCAATCCGGAATTAGATGTCGGACAGGTGGAAGGAGCATTCATGATGGGCGTCGGTTATTGGACCCATGAAGAGTTGATATATGATCAGAAAACTGGAGAGCTGCTCACCGACCGAACTTGGTACTACAAGGTGCCTCTGACTAAGGATATCCCGATAGACTTTAGAATTCAATTCAGGAGGAACTCTTACAACCCAATCGGTACCCTTGGGGCAAGAGCTGTTGCTGAACCTCCGACGTGCTTGTCCATTTGCGTCGCACTAGCCCTGCGGGAAGCTATAGTGTCTTCAAGAGAAGAGACTGGATACCCAAGAAACAAATGGTTCAATGTTGGTAAGCGACTTCTTGTATTCTTATATTAG
Protein
MPRAQNAHAQVNTGFLYEFNAKNKGVIDSARIVIGGLSAHFVHAKQTEKYLKGKDVFTNEVLQHALMVLKGELMVDEIPGEMSPEFRKKLALGLFYKGLLNLIPKKTINPRYRSGARDFRKTRPLSKGSEVYDTNPIIWPLTEPMPKVDALVQCAGEAQYVNDLPKQTAEVYCAFVTSDVGTGEIVEIDPTPALKIPGVLAFFSAKDIPGKNNFLSQRVPTQLFPEEIFAEKTIKYYDQAIGLIVAETEQLANRAALLVHVKYKVDRKPILTIHDARAKDPSRVKLFIIIPPRDRGTNVQRVLKGSDTILGQYHFHMENLSCVTRPSEDGLDVFVATQWTDSVHIGISESLNIEQNRINLTVPRCGGGYGAKISRASHVACACALVTHLMDRPCRFVMSIQANLRVIGKRLPSQRDFEVGLSDSGEIQYMEYQIYEDNGYTVSDAVMPFLIGGVKNCYDNRRWQSKLFNVVTDTASNIYARAPGSLEAISMAEYLMERISYELERDPIEVRLNNLDPQYTEVAEVVQTLLRDSEYHKRKEEVEKFNKENRWKKRGLRLAMMNWPASSVVDYHVLLSVYHGDGTVVVTHGGIEIGQGINTKVIQAVAYTLKISTDKVICKPPTVATNPNNFTTGGSRTTQSVCFGAIKCCQILLDRLSVVREALNDPTWEILIEAAFNRGINLQTSYRVTPNDQDPYRCAGAAVAEVELDVITGEHEILRVDIIEDVGTSINPELDVGQVEGAFMMGVGYWTHEELIYDQKTGELLTDRTWYYKVPLTKDIPIDFRIQFRRNSYNPIGTLGARAVAEPPTCLSICVALALREAIVSSREETGYPRNKWFNVGKRLLVFLY
Summary
Cofactor
FAD
Mo-molybdopterin
[2Fe-2S] cluster
Mo-molybdopterin
[2Fe-2S] cluster
Uniprot
H9JFX1
A0A2A4ITG3
A0A0L7L4A2
A0A2H1VUX4
A0A0N0PF58
A0A194QCB6
+ More
A0A194QUG5 A0A194QBY5 A0A2H1W0Z2 A0A2A4JUF7 A0A2A4ITB4 A0A2H1V2I4 A0A1X9ZM11 A0A2W1BJJ3 A0A2W1BIE3 A0A2H1VA65 A0A3S2PBH0 A0A0N0PD45 A0A194PIT3 A0A2A4ISI6 A0A2A4IRV9 A0A3S2NDD1 A0A2A4JU78 A0A194PE80 A0A2H1VNQ0 H9JFE7 A0A2H1V6B6 A0A2W1BG03 A0A0N1IAK3 A0A0F7QJV6 A0A3S2NMY0 A0A2A4JY06 A0A194PDB3 A0A2W1BIK1 A0A2H1VEC0 H9JFH1 A0A194PCQ5 A0A194PDZ5 A0A1X9ZM23 A0A212FKB8 A8TUC0 A0A076FRA1 A0A212EZ25 H9JFZ2 A0A2A4JYR8 A0A2A4JXM9 F4WWU5 A0A195FSQ5 A0A076FX80 A0A2H1VSX5 A0A0L7R0P8 A0A195E465 A0A2H1VCW8 E2BEN6 A0A151I2B1 A0A0N0BES0 A0A2H1VH28 A0A0L7L3S0 A0A151X416 A0A195CT20 A0A2W1BHI1 A0A2A4JX39 A0A0F7QEA3 A0A1X9ZM28 A0A2J7R4N3 A0A154PJ92 A0A1X9ZLZ6 A0A2A4JB30 E2ANE4 A0A023F0C3 A0A195E3I3 A0A0F7QJW1 A0A195CSI2 A0A0F7QEA1 A0A151X3M2 A0A194QMP0 A0A2A3E4V2 V9IH93 A8TUB4 A0A067QY40 A0A1J1J970 A0A310SF38 A0A1B6JH49 A0A087ZYD0 A0A1L8D6P5 A0A0V0G7W2 A0A1B6E3D0 H9JFX4 A0A212F2C6 T1HWZ8 A0A0F7QIE0 A0A3S2NTK8 A0A1U9X1S0 A0A0H4TD12 A0A0F7QHK5 A0A0H4TY60 A0A1Q3FSG4 D6W6X3 A0A2H1V7P8 A0A076FRM9
A0A194QUG5 A0A194QBY5 A0A2H1W0Z2 A0A2A4JUF7 A0A2A4ITB4 A0A2H1V2I4 A0A1X9ZM11 A0A2W1BJJ3 A0A2W1BIE3 A0A2H1VA65 A0A3S2PBH0 A0A0N0PD45 A0A194PIT3 A0A2A4ISI6 A0A2A4IRV9 A0A3S2NDD1 A0A2A4JU78 A0A194PE80 A0A2H1VNQ0 H9JFE7 A0A2H1V6B6 A0A2W1BG03 A0A0N1IAK3 A0A0F7QJV6 A0A3S2NMY0 A0A2A4JY06 A0A194PDB3 A0A2W1BIK1 A0A2H1VEC0 H9JFH1 A0A194PCQ5 A0A194PDZ5 A0A1X9ZM23 A0A212FKB8 A8TUC0 A0A076FRA1 A0A212EZ25 H9JFZ2 A0A2A4JYR8 A0A2A4JXM9 F4WWU5 A0A195FSQ5 A0A076FX80 A0A2H1VSX5 A0A0L7R0P8 A0A195E465 A0A2H1VCW8 E2BEN6 A0A151I2B1 A0A0N0BES0 A0A2H1VH28 A0A0L7L3S0 A0A151X416 A0A195CT20 A0A2W1BHI1 A0A2A4JX39 A0A0F7QEA3 A0A1X9ZM28 A0A2J7R4N3 A0A154PJ92 A0A1X9ZLZ6 A0A2A4JB30 E2ANE4 A0A023F0C3 A0A195E3I3 A0A0F7QJW1 A0A195CSI2 A0A0F7QEA1 A0A151X3M2 A0A194QMP0 A0A2A3E4V2 V9IH93 A8TUB4 A0A067QY40 A0A1J1J970 A0A310SF38 A0A1B6JH49 A0A087ZYD0 A0A1L8D6P5 A0A0V0G7W2 A0A1B6E3D0 H9JFX4 A0A212F2C6 T1HWZ8 A0A0F7QIE0 A0A3S2NTK8 A0A1U9X1S0 A0A0H4TD12 A0A0F7QHK5 A0A0H4TY60 A0A1Q3FSG4 D6W6X3 A0A2H1V7P8 A0A076FRM9
Pubmed
EMBL
BABH01001558
NWSH01007172
PCG63055.1
JTDY01003017
KOB70302.1
ODYU01004588
+ More
SOQ44620.1 KQ459674 KPJ20482.1 KQ459193 KPJ03064.1 KQ461154 KPJ08620.1 KPJ03063.1 ODYU01005636 SOQ46683.1 NWSH01000648 PCG75032.1 NWSH01007964 PCG62736.1 ODYU01000376 SOQ35055.1 KY595781 ARS46835.1 KZ150135 PZC72996.1 PZC72997.1 ODYU01001481 SOQ37735.1 RSAL01000123 RVE46648.1 KQ460326 KPJ15694.1 KQ459606 KPI91000.1 PCG62735.1 PCG62737.1 RSAL01000088 RVE48202.1 PCG75033.1 KPI90999.1 ODYU01003306 SOQ41884.1 BABH01034979 BABH01034980 BABH01034981 ODYU01000882 SOQ36316.1 KZ150165 PZC72655.1 KPJ15693.1 LC017753 BAR64767.1 RSAL01000229 RVE43893.1 NWSH01000413 PCG76558.1 KPI90998.1 PZC72656.1 ODYU01002094 SOQ39169.1 BABH01034958 BABH01034959 BABH01034960 BABH01034961 BABH01034962 BABH01034963 BABH01034964 BABH01034965 KPI90997.1 KPI90909.1 KY595779 ARS46833.1 AGBW02008082 OWR54176.1 EU073424 ABW71272.1 KF960795 AII21997.1 AGBW02011403 OWR46707.1 BABH01001657 BABH01001658 BABH01001659 BABH01001660 PCG76560.1 PCG76559.1 GL888413 EGI61333.1 KQ981281 KYN43332.1 KF960794 AII21996.1 ODYU01004253 SOQ43930.1 KQ414670 KOC64413.1 KQ979701 KYN19697.1 ODYU01001878 SOQ38698.1 GL447829 EFN85870.1 KQ976546 KYM80948.1 KQ435826 KOX72014.1 ODYU01002537 SOQ40155.1 JTDY01003126 KOB70107.1 KQ982557 KYQ55014.1 KQ977306 KYN03667.1 PZC72657.1 PCG76561.1 LC017757 BAR64771.1 KY595780 ARS46834.1 NEVH01007404 PNF35793.1 KQ434912 KZC11368.1 KY595778 ARS46832.1 NWSH01002143 PCG69059.1 GL441193 EFN65044.1 GBBI01004413 JAC14299.1 KYN19698.1 LC017758 BAR64772.1 KYN03668.1 LC017752 BAR64766.1 KYQ55015.1 KQ461196 KPJ06614.1 KZ288369 PBC26783.1 JR045677 AEY60027.1 EU073423 ABW71271.1 KK852888 KDR14320.1 CVRI01000075 CRL08602.1 KQ769812 OAD52839.1 GECU01009219 JAS98487.1 GEYN01000003 JAV02126.1 GECL01002669 JAP03455.1 GEDC01004862 JAS32436.1 BABH01001568 BABH01001569 BABH01001570 BABH01001571 BABH01001572 BABH01001573 AGBW02010767 OWR47879.1 ACPB03000354 ACPB03000355 LC017754 BAR64768.1 RVE48204.1 KY643683 AQY62686.1 KP899546 AKQ06145.1 LC017755 BAR64769.1 KP899547 AKQ06146.1 GFDL01004496 JAV30549.1 KQ971307 EFA11446.2 ODYU01001109 SOQ36868.1 KF960796 AII21998.1
SOQ44620.1 KQ459674 KPJ20482.1 KQ459193 KPJ03064.1 KQ461154 KPJ08620.1 KPJ03063.1 ODYU01005636 SOQ46683.1 NWSH01000648 PCG75032.1 NWSH01007964 PCG62736.1 ODYU01000376 SOQ35055.1 KY595781 ARS46835.1 KZ150135 PZC72996.1 PZC72997.1 ODYU01001481 SOQ37735.1 RSAL01000123 RVE46648.1 KQ460326 KPJ15694.1 KQ459606 KPI91000.1 PCG62735.1 PCG62737.1 RSAL01000088 RVE48202.1 PCG75033.1 KPI90999.1 ODYU01003306 SOQ41884.1 BABH01034979 BABH01034980 BABH01034981 ODYU01000882 SOQ36316.1 KZ150165 PZC72655.1 KPJ15693.1 LC017753 BAR64767.1 RSAL01000229 RVE43893.1 NWSH01000413 PCG76558.1 KPI90998.1 PZC72656.1 ODYU01002094 SOQ39169.1 BABH01034958 BABH01034959 BABH01034960 BABH01034961 BABH01034962 BABH01034963 BABH01034964 BABH01034965 KPI90997.1 KPI90909.1 KY595779 ARS46833.1 AGBW02008082 OWR54176.1 EU073424 ABW71272.1 KF960795 AII21997.1 AGBW02011403 OWR46707.1 BABH01001657 BABH01001658 BABH01001659 BABH01001660 PCG76560.1 PCG76559.1 GL888413 EGI61333.1 KQ981281 KYN43332.1 KF960794 AII21996.1 ODYU01004253 SOQ43930.1 KQ414670 KOC64413.1 KQ979701 KYN19697.1 ODYU01001878 SOQ38698.1 GL447829 EFN85870.1 KQ976546 KYM80948.1 KQ435826 KOX72014.1 ODYU01002537 SOQ40155.1 JTDY01003126 KOB70107.1 KQ982557 KYQ55014.1 KQ977306 KYN03667.1 PZC72657.1 PCG76561.1 LC017757 BAR64771.1 KY595780 ARS46834.1 NEVH01007404 PNF35793.1 KQ434912 KZC11368.1 KY595778 ARS46832.1 NWSH01002143 PCG69059.1 GL441193 EFN65044.1 GBBI01004413 JAC14299.1 KYN19698.1 LC017758 BAR64772.1 KYN03668.1 LC017752 BAR64766.1 KYQ55015.1 KQ461196 KPJ06614.1 KZ288369 PBC26783.1 JR045677 AEY60027.1 EU073423 ABW71271.1 KK852888 KDR14320.1 CVRI01000075 CRL08602.1 KQ769812 OAD52839.1 GECU01009219 JAS98487.1 GEYN01000003 JAV02126.1 GECL01002669 JAP03455.1 GEDC01004862 JAS32436.1 BABH01001568 BABH01001569 BABH01001570 BABH01001571 BABH01001572 BABH01001573 AGBW02010767 OWR47879.1 ACPB03000354 ACPB03000355 LC017754 BAR64768.1 RVE48204.1 KY643683 AQY62686.1 KP899546 AKQ06145.1 LC017755 BAR64769.1 KP899547 AKQ06146.1 GFDL01004496 JAV30549.1 KQ971307 EFA11446.2 ODYU01001109 SOQ36868.1 KF960796 AII21998.1
Proteomes
UP000005204
UP000218220
UP000037510
UP000053240
UP000053268
UP000283053
+ More
UP000007151 UP000007755 UP000078541 UP000053825 UP000078492 UP000008237 UP000078540 UP000053105 UP000075809 UP000078542 UP000235965 UP000076502 UP000000311 UP000242457 UP000027135 UP000183832 UP000005203 UP000015103 UP000007266
UP000007151 UP000007755 UP000078541 UP000053825 UP000078492 UP000008237 UP000078540 UP000053105 UP000075809 UP000078542 UP000235965 UP000076502 UP000000311 UP000242457 UP000027135 UP000183832 UP000005203 UP000015103 UP000007266
Pfam
Interpro
IPR002346
Mopterin_DH_FAD-bd
+ More
IPR037165 AldOxase/xan_DH_Mopterin-bd_sf
IPR016166 FAD-bd_PCMH
IPR036856 Ald_Oxase/Xan_DH_a/b_sf
IPR036683 CO_DH_flav_C_dom_sf
IPR016169 FAD-bd_PCMH_sub2
IPR036318 FAD-bd_PCMH-like_sf
IPR005107 CO_DH_flav_C
IPR008274 AldOxase/xan_DH_Mopterin-bd
IPR036884 2Fe-2S-bd_dom_sf
IPR016208 Ald_Oxase/xanthine_DH
IPR000674 Ald_Oxase/Xan_DH_a/b
IPR002888 2Fe-2S-bd
IPR036010 2Fe-2S_ferredoxin-like_sf
IPR001041 2Fe-2S_ferredoxin-type
IPR006058 2Fe2S_fd_BS
IPR012675 Beta-grasp_dom_sf
IPR002159 CD36_fam
IPR006621 Nose-resist-to-fluoxetine_N
IPR013809 ENTH
IPR008942 ENTH_VHS
IPR011417 ANTH_dom
IPR014712 Clathrin_AP_dom2
IPR037165 AldOxase/xan_DH_Mopterin-bd_sf
IPR016166 FAD-bd_PCMH
IPR036856 Ald_Oxase/Xan_DH_a/b_sf
IPR036683 CO_DH_flav_C_dom_sf
IPR016169 FAD-bd_PCMH_sub2
IPR036318 FAD-bd_PCMH-like_sf
IPR005107 CO_DH_flav_C
IPR008274 AldOxase/xan_DH_Mopterin-bd
IPR036884 2Fe-2S-bd_dom_sf
IPR016208 Ald_Oxase/xanthine_DH
IPR000674 Ald_Oxase/Xan_DH_a/b
IPR002888 2Fe-2S-bd
IPR036010 2Fe-2S_ferredoxin-like_sf
IPR001041 2Fe-2S_ferredoxin-type
IPR006058 2Fe2S_fd_BS
IPR012675 Beta-grasp_dom_sf
IPR002159 CD36_fam
IPR006621 Nose-resist-to-fluoxetine_N
IPR013809 ENTH
IPR008942 ENTH_VHS
IPR011417 ANTH_dom
IPR014712 Clathrin_AP_dom2
SUPFAM
Gene 3D
CDD
ProteinModelPortal
H9JFX1
A0A2A4ITG3
A0A0L7L4A2
A0A2H1VUX4
A0A0N0PF58
A0A194QCB6
+ More
A0A194QUG5 A0A194QBY5 A0A2H1W0Z2 A0A2A4JUF7 A0A2A4ITB4 A0A2H1V2I4 A0A1X9ZM11 A0A2W1BJJ3 A0A2W1BIE3 A0A2H1VA65 A0A3S2PBH0 A0A0N0PD45 A0A194PIT3 A0A2A4ISI6 A0A2A4IRV9 A0A3S2NDD1 A0A2A4JU78 A0A194PE80 A0A2H1VNQ0 H9JFE7 A0A2H1V6B6 A0A2W1BG03 A0A0N1IAK3 A0A0F7QJV6 A0A3S2NMY0 A0A2A4JY06 A0A194PDB3 A0A2W1BIK1 A0A2H1VEC0 H9JFH1 A0A194PCQ5 A0A194PDZ5 A0A1X9ZM23 A0A212FKB8 A8TUC0 A0A076FRA1 A0A212EZ25 H9JFZ2 A0A2A4JYR8 A0A2A4JXM9 F4WWU5 A0A195FSQ5 A0A076FX80 A0A2H1VSX5 A0A0L7R0P8 A0A195E465 A0A2H1VCW8 E2BEN6 A0A151I2B1 A0A0N0BES0 A0A2H1VH28 A0A0L7L3S0 A0A151X416 A0A195CT20 A0A2W1BHI1 A0A2A4JX39 A0A0F7QEA3 A0A1X9ZM28 A0A2J7R4N3 A0A154PJ92 A0A1X9ZLZ6 A0A2A4JB30 E2ANE4 A0A023F0C3 A0A195E3I3 A0A0F7QJW1 A0A195CSI2 A0A0F7QEA1 A0A151X3M2 A0A194QMP0 A0A2A3E4V2 V9IH93 A8TUB4 A0A067QY40 A0A1J1J970 A0A310SF38 A0A1B6JH49 A0A087ZYD0 A0A1L8D6P5 A0A0V0G7W2 A0A1B6E3D0 H9JFX4 A0A212F2C6 T1HWZ8 A0A0F7QIE0 A0A3S2NTK8 A0A1U9X1S0 A0A0H4TD12 A0A0F7QHK5 A0A0H4TY60 A0A1Q3FSG4 D6W6X3 A0A2H1V7P8 A0A076FRM9
A0A194QUG5 A0A194QBY5 A0A2H1W0Z2 A0A2A4JUF7 A0A2A4ITB4 A0A2H1V2I4 A0A1X9ZM11 A0A2W1BJJ3 A0A2W1BIE3 A0A2H1VA65 A0A3S2PBH0 A0A0N0PD45 A0A194PIT3 A0A2A4ISI6 A0A2A4IRV9 A0A3S2NDD1 A0A2A4JU78 A0A194PE80 A0A2H1VNQ0 H9JFE7 A0A2H1V6B6 A0A2W1BG03 A0A0N1IAK3 A0A0F7QJV6 A0A3S2NMY0 A0A2A4JY06 A0A194PDB3 A0A2W1BIK1 A0A2H1VEC0 H9JFH1 A0A194PCQ5 A0A194PDZ5 A0A1X9ZM23 A0A212FKB8 A8TUC0 A0A076FRA1 A0A212EZ25 H9JFZ2 A0A2A4JYR8 A0A2A4JXM9 F4WWU5 A0A195FSQ5 A0A076FX80 A0A2H1VSX5 A0A0L7R0P8 A0A195E465 A0A2H1VCW8 E2BEN6 A0A151I2B1 A0A0N0BES0 A0A2H1VH28 A0A0L7L3S0 A0A151X416 A0A195CT20 A0A2W1BHI1 A0A2A4JX39 A0A0F7QEA3 A0A1X9ZM28 A0A2J7R4N3 A0A154PJ92 A0A1X9ZLZ6 A0A2A4JB30 E2ANE4 A0A023F0C3 A0A195E3I3 A0A0F7QJW1 A0A195CSI2 A0A0F7QEA1 A0A151X3M2 A0A194QMP0 A0A2A3E4V2 V9IH93 A8TUB4 A0A067QY40 A0A1J1J970 A0A310SF38 A0A1B6JH49 A0A087ZYD0 A0A1L8D6P5 A0A0V0G7W2 A0A1B6E3D0 H9JFX4 A0A212F2C6 T1HWZ8 A0A0F7QIE0 A0A3S2NTK8 A0A1U9X1S0 A0A0H4TD12 A0A0F7QHK5 A0A0H4TY60 A0A1Q3FSG4 D6W6X3 A0A2H1V7P8 A0A076FRM9
PDB
3UNI
E-value=4.05682e-96,
Score=900
Ontologies
PATHWAY
GO
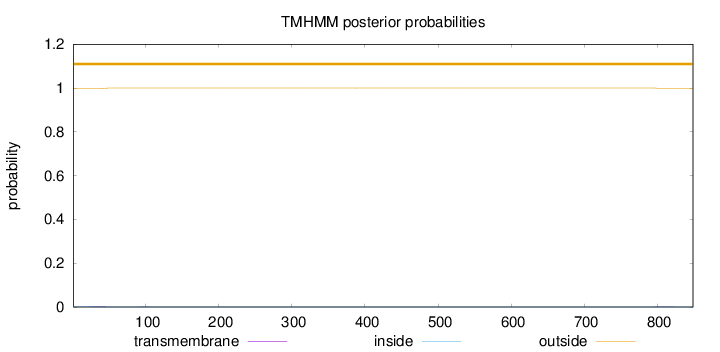
Topology
Length:
849
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.0582600000000003
Exp number, first 60 AAs:
0.0437
Total prob of N-in:
0.00268
outside
1 - 849
Population Genetic Test Statistics
Pi
187.826909
Theta
151.690754
Tajima's D
0.597675
CLR
1.042121
CSRT
0.547722613869307
Interpretation
Uncertain
