Gene
KWMTBOMO10647 Validated by peptides from experiments
Pre Gene Modal
BGIBMGA014429
Annotation
serine_protease_[Bombyx_mandarina]
Location in the cell
Extracellular Reliability : 2.753
Sequence
CDS
ATGAAGGTCTTCGCAGCAGTACTGATGGCGTTGGCGGCCGTGGTCGTGGCAGAAGAGGCCATCGAACTTGACTACCACAACAAGATCGGTATCCCCCGGGCTGAGAGTCTTAGACGCGCCGAGGAAGCCGCTGACTTCGACGGTACCAGGATTGTGGGTGGTTCTGCCGCCAACGCTGGTGCTCACCCCCATCTTGCTGGACTTGTGATCGCACTCACGAATGGCAGAACTTCCATCTGCGGAGCTTCCTTACTGACCAACACCCGCTCCGTGACCGCGGCTCACTGCTGGAGAACCAGGAACGCCCAGGCTCGTCAGTTCACCCTCGCTTTTGGTACAGCTAACATCTTCTCCGGCGGCACCAGGGTCACCACCTCCAGTGTCCACCTGCACGGCAGCTACAACATGAACAACCTCAACAACGACGTCGCCATCATCAACCACAACCATGTTGGCTTCAACAACAACATCCAGCGCATCAACCTAGCCAGTGGAAACAACAACTTTGCTGGTACTTGGGCCTGGGCTGCCGGCTTCGGGAGGACCTCCGATGCTGCTTCGGGAGCCAACAACCAACAAAAACGCCAAGTGAGCCTCCAGGTCATCACCAACGCCGTCTGCGCCCGCACGTACGGAAACTCTGTGATCATTGGCTCCACCCTCTGTGTTTCCGGCGCTAACGGTCGCAGCACCTGCAGCGGAGACTCCGGCGGCCCTCTCTCCATCGGCAGCGGCGGAGGCCGTCAGCTGATCGGTATCACGTCGTTCGGATCAGACAGAGGCTGCCAGAGAGGCTACCCTGCCGGCTTCGCTAGAGTCACATCTTTCAACTCCTGGATCCGGGCTAGAATTTAA
Protein
MKVFAAVLMALAAVVVAEEAIELDYHNKIGIPRAESLRRAEEAADFDGTRIVGGSAANAGAHPHLAGLVIALTNGRTSICGASLLTNTRSVTAAHCWRTRNAQARQFTLAFGTANIFSGGTRVTTSSVHLHGSYNMNNLNNDVAIINHNHVGFNNNIQRINLASGNNNFAGTWAWAAGFGRTSDAASGANNQQKRQVSLQVITNAVCARTYGNSVIIGSTLCVSGANGRSTCSGDSGGPLSIGSGGGRQLIGITSFGSDRGCQRGYPAGFARVTSFNSWIRARI
Summary
Similarity
Belongs to the peptidase S1 family.
Belongs to the ETS family.
Belongs to the ETS family.
Uniprot
H9JY11
Q58I80
Q762I9
H9JY13
H9JY12
Q58I79
+ More
H9JY10 O01953 A0A194Q3W8 A0A194Q399 H9JY09 A0A3S2LL18 A0A0N0PAA7 T1WJA5 A0A0N0PCX7 A0A194Q0R8 A0A0N0PA14 A0A0N1PGY7 A0A2W1BAG4 A0A0N1IFV7 A0A0N0PAA9 A0A2H1WCW9 A0A089QFN1 A0A194R2V7 A0A0N1IAY5 A0A212EYA6 A0A2A4JJ77 Q56IC0 C9W8H4 A0A0M4LSN9 A0A0N1PIX2 I6TRU6 A0A0M3T9F4 I6SW50 A0A0N1I5V5 H9JCY0 A0A0N0PEU9 A0A0N1PHR5 A0A1E1WV21 B2ZJV9 H9JJ25 Q9NH07 A0A0N1ICZ9 A0A194QCZ1 A0A1B0RHP3 O18450 E7D003 O18445 A0A2A4K228 C9W8G8 O18444 A0PGD6 C9W8F9 Q9N6C6 A0A142I706 A0A2A4K0D3 A0PGD7 B4X987 A0A2A4J8H7 Q56IB9 A0A2A4J6X3 B4X988 A0A2A4K1C2 A0A194PUF7 A0A0L7L962 E7D008 A0A2A4K1Q8 O18443 A0A0L7KVY7 J7FDV3 A0A2A4JSJ6 A0A089RSZ3 B4X990 A0A194QF72 I6TEX2 C9W8H8 B4X989 S5QJ14 E7D013 C7FF06 D7RZZ6 D2Y4U4 D7RZZ8 A9P3S5 B6CMF0 Q25503 D7RZZ7 Q9NH10 J7FCZ4 D7RZZ9 A0A2A4JFK9 Q86M89 A0A2H1V6V4 A0A194QK37 O18438 A0A212F4C3 A0PGE0 B6D1R1 C9W8G7 A0PGD5
H9JY10 O01953 A0A194Q3W8 A0A194Q399 H9JY09 A0A3S2LL18 A0A0N0PAA7 T1WJA5 A0A0N0PCX7 A0A194Q0R8 A0A0N0PA14 A0A0N1PGY7 A0A2W1BAG4 A0A0N1IFV7 A0A0N0PAA9 A0A2H1WCW9 A0A089QFN1 A0A194R2V7 A0A0N1IAY5 A0A212EYA6 A0A2A4JJ77 Q56IC0 C9W8H4 A0A0M4LSN9 A0A0N1PIX2 I6TRU6 A0A0M3T9F4 I6SW50 A0A0N1I5V5 H9JCY0 A0A0N0PEU9 A0A0N1PHR5 A0A1E1WV21 B2ZJV9 H9JJ25 Q9NH07 A0A0N1ICZ9 A0A194QCZ1 A0A1B0RHP3 O18450 E7D003 O18445 A0A2A4K228 C9W8G8 O18444 A0PGD6 C9W8F9 Q9N6C6 A0A142I706 A0A2A4K0D3 A0PGD7 B4X987 A0A2A4J8H7 Q56IB9 A0A2A4J6X3 B4X988 A0A2A4K1C2 A0A194PUF7 A0A0L7L962 E7D008 A0A2A4K1Q8 O18443 A0A0L7KVY7 J7FDV3 A0A2A4JSJ6 A0A089RSZ3 B4X990 A0A194QF72 I6TEX2 C9W8H8 B4X989 S5QJ14 E7D013 C7FF06 D7RZZ6 D2Y4U4 D7RZZ8 A9P3S5 B6CMF0 Q25503 D7RZZ7 Q9NH10 J7FCZ4 D7RZZ9 A0A2A4JFK9 Q86M89 A0A2H1V6V4 A0A194QK37 O18438 A0A212F4C3 A0PGE0 B6D1R1 C9W8G7 A0PGD5
Pubmed
EMBL
BABH01041853
AY945210
EU338400
AAX39408.1
ABY58153.1
AB117641
+ More
BAD13342.1 BABH01041861 BABH01041859 BABH01041860 AY945211 DQ310733 AAX39409.1 ABC26003.1 BABH01041852 BABH01042790 AB003670 BAA20136.1 KQ459472 KPJ00237.1 KQ459586 KPI97880.1 BABH01041868 RSAL01000073 RVE48982.1 KQ458725 KPJ05397.1 KF039687 AGU27161.1 KQ460401 KPJ14971.1 KQ459581 KPI99177.1 KQ458978 KPJ04519.1 KQ461033 KPJ09997.1 KZ150191 PZC72392.1 KQ460140 KPJ17495.1 KPJ05400.1 ODYU01007826 SOQ50918.1 KM083780 AIR09761.1 KQ460845 KPJ12032.1 KPJ05401.1 AGBW02011570 OWR46476.1 NWSH01001385 PCG71452.1 AY953056 AY953057 AAX62029.1 AAX62030.1 FJ205420 ACR15987.3 KR024679 ALE15221.1 KPJ17496.1 JQ904141 AFM77771.1 KR024680 ALE15222.1 JQ904140 AFM77770.1 KPJ05396.1 BABH01027594 KQ459620 KPJ20544.1 KPJ17494.1 GDQN01000181 JAT90873.1 EU672968 ACD44927.1 BABH01023267 AF233732 AAF71519.1 KQ459388 KPJ01122.1 KQ459185 KPJ03332.1 KM360186 AKH49602.1 Y12287 CAA72966.1 HM990173 ADT80821.1 Y12281 CAA72960.1 NWSH01000291 PCG77692.1 FJ205413 ACR15981.1 Y12280 CAA72959.1 AY618891 AAV33654.1 FJ205404 ACR15972.2 AF233731 AF233734 AAF71518.1 AAF71716.1 KT907053 AMR44225.1 PCG77697.1 AY618892 AAV33655.1 EF531623 ABR88234.1 PCG77696.1 NWSH01002619 PCG67834.1 AY953058 AAX62031.1 PCG67835.1 EF531624 ABR88235.1 PCG77694.1 KQ459591 KPI97041.1 JTDY01002254 KOB71786.1 HM990178 ADT80826.1 PCG77693.1 Y12279 CAA72958.1 JTDY01005262 KOB67184.1 JN252035 AFM28248.1 NWSH01000651 PCG75015.1 KM083791 ODYU01000971 AIR09772.1 SOQ36530.1 EF531626 ABR88237.1 KQ459252 KPJ02116.1 JQ904139 AFM77769.1 FJ205424 ACR15991.1 EF531625 ABR88236.1 KC175563 AGR92346.1 HM990183 ADT80831.1 GQ354838 ACU00133.1 HM209419 ADI32880.1 GU323796 ADA83701.1 HM209421 ADI32882.1 EF104914 ABP96916.1 EU325550 ACB54940.1 L34168 AAA58743.1 HM209420 ADI32881.1 AF233728 AAF71515.1 JN252036 AFM28249.1 HM209422 ADI32883.1 NWSH01001552 PCG70875.1 AY251276 AAO75039.1 SOQ36529.1 KQ461198 KPJ05938.1 Y12273 CAA72952.1 AGBW02010392 OWR48581.1 AY618895 AAV33658.1 EU673454 ACI45417.1 FJ205412 ACR15980.1 AY618890 AAV33653.1
BAD13342.1 BABH01041861 BABH01041859 BABH01041860 AY945211 DQ310733 AAX39409.1 ABC26003.1 BABH01041852 BABH01042790 AB003670 BAA20136.1 KQ459472 KPJ00237.1 KQ459586 KPI97880.1 BABH01041868 RSAL01000073 RVE48982.1 KQ458725 KPJ05397.1 KF039687 AGU27161.1 KQ460401 KPJ14971.1 KQ459581 KPI99177.1 KQ458978 KPJ04519.1 KQ461033 KPJ09997.1 KZ150191 PZC72392.1 KQ460140 KPJ17495.1 KPJ05400.1 ODYU01007826 SOQ50918.1 KM083780 AIR09761.1 KQ460845 KPJ12032.1 KPJ05401.1 AGBW02011570 OWR46476.1 NWSH01001385 PCG71452.1 AY953056 AY953057 AAX62029.1 AAX62030.1 FJ205420 ACR15987.3 KR024679 ALE15221.1 KPJ17496.1 JQ904141 AFM77771.1 KR024680 ALE15222.1 JQ904140 AFM77770.1 KPJ05396.1 BABH01027594 KQ459620 KPJ20544.1 KPJ17494.1 GDQN01000181 JAT90873.1 EU672968 ACD44927.1 BABH01023267 AF233732 AAF71519.1 KQ459388 KPJ01122.1 KQ459185 KPJ03332.1 KM360186 AKH49602.1 Y12287 CAA72966.1 HM990173 ADT80821.1 Y12281 CAA72960.1 NWSH01000291 PCG77692.1 FJ205413 ACR15981.1 Y12280 CAA72959.1 AY618891 AAV33654.1 FJ205404 ACR15972.2 AF233731 AF233734 AAF71518.1 AAF71716.1 KT907053 AMR44225.1 PCG77697.1 AY618892 AAV33655.1 EF531623 ABR88234.1 PCG77696.1 NWSH01002619 PCG67834.1 AY953058 AAX62031.1 PCG67835.1 EF531624 ABR88235.1 PCG77694.1 KQ459591 KPI97041.1 JTDY01002254 KOB71786.1 HM990178 ADT80826.1 PCG77693.1 Y12279 CAA72958.1 JTDY01005262 KOB67184.1 JN252035 AFM28248.1 NWSH01000651 PCG75015.1 KM083791 ODYU01000971 AIR09772.1 SOQ36530.1 EF531626 ABR88237.1 KQ459252 KPJ02116.1 JQ904139 AFM77769.1 FJ205424 ACR15991.1 EF531625 ABR88236.1 KC175563 AGR92346.1 HM990183 ADT80831.1 GQ354838 ACU00133.1 HM209419 ADI32880.1 GU323796 ADA83701.1 HM209421 ADI32882.1 EF104914 ABP96916.1 EU325550 ACB54940.1 L34168 AAA58743.1 HM209420 ADI32881.1 AF233728 AAF71515.1 JN252036 AFM28249.1 HM209422 ADI32883.1 NWSH01001552 PCG70875.1 AY251276 AAO75039.1 SOQ36529.1 KQ461198 KPJ05938.1 Y12273 CAA72952.1 AGBW02010392 OWR48581.1 AY618895 AAV33658.1 EU673454 ACI45417.1 FJ205412 ACR15980.1 AY618890 AAV33653.1
Proteomes
Interpro
Gene 3D
CDD
ProteinModelPortal
H9JY11
Q58I80
Q762I9
H9JY13
H9JY12
Q58I79
+ More
H9JY10 O01953 A0A194Q3W8 A0A194Q399 H9JY09 A0A3S2LL18 A0A0N0PAA7 T1WJA5 A0A0N0PCX7 A0A194Q0R8 A0A0N0PA14 A0A0N1PGY7 A0A2W1BAG4 A0A0N1IFV7 A0A0N0PAA9 A0A2H1WCW9 A0A089QFN1 A0A194R2V7 A0A0N1IAY5 A0A212EYA6 A0A2A4JJ77 Q56IC0 C9W8H4 A0A0M4LSN9 A0A0N1PIX2 I6TRU6 A0A0M3T9F4 I6SW50 A0A0N1I5V5 H9JCY0 A0A0N0PEU9 A0A0N1PHR5 A0A1E1WV21 B2ZJV9 H9JJ25 Q9NH07 A0A0N1ICZ9 A0A194QCZ1 A0A1B0RHP3 O18450 E7D003 O18445 A0A2A4K228 C9W8G8 O18444 A0PGD6 C9W8F9 Q9N6C6 A0A142I706 A0A2A4K0D3 A0PGD7 B4X987 A0A2A4J8H7 Q56IB9 A0A2A4J6X3 B4X988 A0A2A4K1C2 A0A194PUF7 A0A0L7L962 E7D008 A0A2A4K1Q8 O18443 A0A0L7KVY7 J7FDV3 A0A2A4JSJ6 A0A089RSZ3 B4X990 A0A194QF72 I6TEX2 C9W8H8 B4X989 S5QJ14 E7D013 C7FF06 D7RZZ6 D2Y4U4 D7RZZ8 A9P3S5 B6CMF0 Q25503 D7RZZ7 Q9NH10 J7FCZ4 D7RZZ9 A0A2A4JFK9 Q86M89 A0A2H1V6V4 A0A194QK37 O18438 A0A212F4C3 A0PGE0 B6D1R1 C9W8G7 A0PGD5
H9JY10 O01953 A0A194Q3W8 A0A194Q399 H9JY09 A0A3S2LL18 A0A0N0PAA7 T1WJA5 A0A0N0PCX7 A0A194Q0R8 A0A0N0PA14 A0A0N1PGY7 A0A2W1BAG4 A0A0N1IFV7 A0A0N0PAA9 A0A2H1WCW9 A0A089QFN1 A0A194R2V7 A0A0N1IAY5 A0A212EYA6 A0A2A4JJ77 Q56IC0 C9W8H4 A0A0M4LSN9 A0A0N1PIX2 I6TRU6 A0A0M3T9F4 I6SW50 A0A0N1I5V5 H9JCY0 A0A0N0PEU9 A0A0N1PHR5 A0A1E1WV21 B2ZJV9 H9JJ25 Q9NH07 A0A0N1ICZ9 A0A194QCZ1 A0A1B0RHP3 O18450 E7D003 O18445 A0A2A4K228 C9W8G8 O18444 A0PGD6 C9W8F9 Q9N6C6 A0A142I706 A0A2A4K0D3 A0PGD7 B4X987 A0A2A4J8H7 Q56IB9 A0A2A4J6X3 B4X988 A0A2A4K1C2 A0A194PUF7 A0A0L7L962 E7D008 A0A2A4K1Q8 O18443 A0A0L7KVY7 J7FDV3 A0A2A4JSJ6 A0A089RSZ3 B4X990 A0A194QF72 I6TEX2 C9W8H8 B4X989 S5QJ14 E7D013 C7FF06 D7RZZ6 D2Y4U4 D7RZZ8 A9P3S5 B6CMF0 Q25503 D7RZZ7 Q9NH10 J7FCZ4 D7RZZ9 A0A2A4JFK9 Q86M89 A0A2H1V6V4 A0A194QK37 O18438 A0A212F4C3 A0PGE0 B6D1R1 C9W8G7 A0PGD5
PDB
2HLC
E-value=1.54167e-22,
Score=261
Ontologies
GO
Topology
Subcellular location
Nucleus
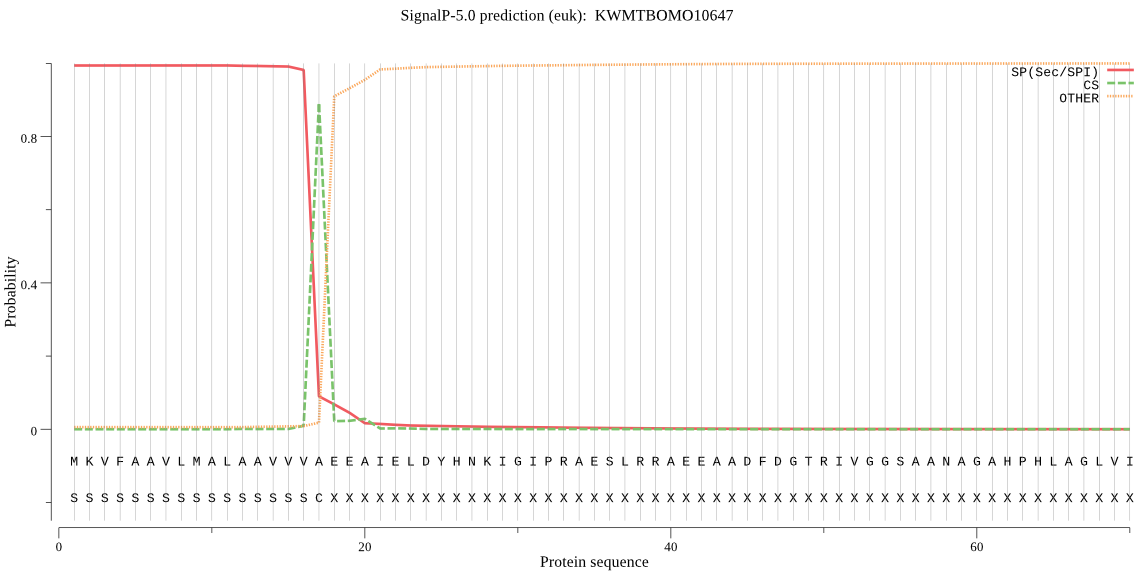
SignalP
Position: 1 - 17,
Likelihood: 0.993425
Length:
284
Number of predicted TMHs:
0
Exp number of AAs in TMHs:
0.43393
Exp number, first 60 AAs:
0.24718
Total prob of N-in:
0.03903
outside
1 - 284

Population Genetic Test Statistics
Pi
373.811449
Theta
217.687539
Tajima's D
2.278943
CLR
0.000005
CSRT
0.92070396480176
Interpretation
Uncertain
